Abstract
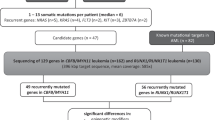
Pediatric acute myeloid leukemia (AML) is a rare disease whose prognosis is highly variable according to factors such as chromosomal abnormalities. Recurrent genomic rearrangements are detected in half of pediatric AML by karyotype. NUcleoPorin 98 (NUP98) gene is rearranged with 31 different fusion partner genes. These rearrangements are frequently undetected by conventional cytogenetics, as the NUP98 gene is located at the end of the chromosome 11 short arm (11p15). By screening a series of 574 pediatric AML, we detected a NUP98 rearrangement in 22 cases (3.8%), a frequency similar to CBFB-MYH11 fusion gene (4.0%). The most frequent NUP98 fusion gene partner is NSD1. These cases are homogeneous regarding their biological and clinical characteristics, and associated with bad prognosis only improved by bone marrow transplantation. We detailed the biological characteristics of these AML by exome sequencing which demonstrated few recurrent mutations (FLT3 ITD, WT1, CEBPA, NBPF14, BCR and ODF1). The analysis of the clonal structure in these cases suggests that the mutation order in the NUP98-rearranged pediatric AML begins with the NUP98 rearrangement leading to epigenetic dysregulations then followed by mutations of critical hematopoietic transcription factors and finally, activation of the FLT3 signaling pathway.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Creutzig U, van den Heuvel-Eibrink MM, Gibson B, Dworzak MN, Adachi S, de Bont E et al. Diagnosis and management of acute myeloid leukemia in children and adolescents: recommendations from an international expert panel. Blood 2012; 120: 3187–3205.
Zwaan CM, Kolb EA, Reinhardt D, Abrahamsson J, Adachi S, Aplenc R et al. Collaborative efforts driving progress in pediatric acute myeloid leukemia. J Clin Oncol 2015; 33: 2949–2962.
Fontoura BM, Blobel G, Matunis MJ . A conserved biogenesis pathway for nucleoporins: proteolytic processing of a 186-kilodalton precursor generates Nup98 and the novel nucleoporin, Nup96. J Cell Biol 1999; 144: 1097–1112.
Borrow J, Shearman AM, Stanton VP Jr., Becher R, Collins T, Williams AJ et al. The t(7;11)(p15;p15) translocation in acute myeloid leukaemia fuses the genes for nucleoporin NUP98 and class I homeoprotein HOXA9. Nat Genet 1996; 12: 159–167.
Nakamura T, Largaespada DA, Lee MP, Johnson LA, Ohyashiki K, Toyama K et al. Fusion of the nucleoporin gene NUP98 to HOXA9 by the chromosome translocation t(7;11)(p15;p15) in human myeloid leukaemia. Nat Genet 1996; 12: 154–158.
Romana SP, Radford-Weiss I, Ben Abdelali R, Schluth C, Petit A, Dastugue N et al. NUP98 rearrangements in hematopoietic malignancies: a study of the Groupe Francophone de Cytogenetique Hematologique. Leukemia 2006; 20: 696–706.
Gough SM, Slape CI, Aplan PD . NUP98 gene fusions and hematopoietic malignancies: common themes and new biologic insights. Blood 2011; 118: 6247–6257.
Lisboa S, Cerveira N, Bizarro S, Correia C, Vieira J, Torres L et al. POU1F1 is a novel fusion partner of NUP98 in acute myeloid leukemia with t(3;11)(p11;p15). Mol Cancer 2013; 12: 5.
Soler G, Kaltenbach S, Dobbelstein S, Broccardo C, Radford I, Mozziconacci MJ et al. Identification of GSX2 and AF10 as NUP98 partner genes in myeloid malignancies. Blood Cancer J 2013; 3: e124.
Togni M, Masetti R, Pigazzi M, Astolfi A, Zama D, Indio V et al. Identification of the NUP98-PHF23 fusion gene in pediatric cytogenetically normal acute myeloid leukemia by whole-transcriptome sequencing. J Hematol Oncol 2015; 8: 69.
Jaju RJ, Fidler C, Haas OA, Strickson AJ, Watkins F, Clark K et al. A novel gene, NSD1, is fused to NUP98 in the t(5;11)(q35;p15.5) in de novo childhood acute myeloid leukemia. Blood 2001; 98: 1264–1267.
Hollink IH, van den Heuvel-Eibrink MM, Arentsen-Peters ST, Pratcorona M, Abbas S, Kuipers JE et al. NUP98/NSD1 characterizes a novel poor prognostic group in acute myeloid leukemia with a distinct HOX gene expression pattern. Blood 2011; 118: 3645–3656.
Cerveira N, Correia C, Doria S, Bizarro S, Rocha P, Gomes P et al. Frequency of NUP98-NSD1 fusion transcript in childhood acute myeloid leukaemia. Leukemia 2003; 17: 2244–2247.
Radtke I, Mullighan CG, Ishii M, Su X, Cheng J, Ma J et al. Genomic analysis reveals few genetic alterations in pediatric acute myeloid leukemia. Proc Natl Acad Sci USA 2009; 106: 12944–12949.
Akiki S, Dyer SA, Grimwade D, Ivey A, Abou-Zeid N, Borrow J et al. NUP98-NSD1 fusion in association with FLT3-ITD mutation identifies a prognostically relevant subgroup of pediatric acute myeloid leukemia patients suitable for monitoring by real time quantitative PCR. Genes Chromosomes Cancer 2013; 52: 1053–1064.
Shiba N, Ichikawa H, Taki T, Park MJ, Jo A, Mitani S et al. NUP98-NSD1 gene fusion and its related gene expression signature are strongly associated with a poor prognosis in pediatric acute myeloid leukemia. Genes Chromosomes Cancer 2013; 52: 683–693.
Ostronoff F, Othus M, Gerbing RB, Loken MR, Raimondi SC, Hirsch BA et al. NUP98/NSD1 and FLT3/ITD coexpression is more prevalent in younger AML patients and leads to induction failure: a COG and SWOG report. Blood 2014; 124: 2400–2407.
Breems DA, Van Putten WL, De Greef GE, Van Zelderen-Bhola SL, Gerssen-Schoorl KB, Mellink CH et al. Monosomal karyotype in acute myeloid leukemia: a better indicator of poor prognosis than a complex karyotype. J Clin Oncol 2008; 26: 4791–4797.
Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL . TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 2013; 14: R36.
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 2012; 7: 562–578.
Nakao M, Yokota S, Iwai T, Kaneko H, Horiike S, Kashima K et al. Internal tandem duplication of the flt3 gene found in acute myeloid leukemia. Leukemia 1996; 10: 1911–1918.
Pabst T, Mueller BU, Zhang P, Radomska HS, Narravula S, Schnittger S et al. Dominant-negative mutations of CEBPA, encoding CCAAT/enhancer binding protein-alpha (C/EBPalpha), in acute myeloid leukemia. Nat Genet 2001; 27: 263–270.
Kaspers GJ, Zimmermann M, Reinhardt D, Gibson BE, Tamminga RY, Aleinikova O et al. Improved outcome in pediatric relapsed acute myeloid leukemia: results of a randomized trial on liposomal daunorubicin by the International BFM Study Group. J Clin Oncol 2013; 31: 599–607.
Chou WC, Chen CY, Hou HA, Lin LI, Tang JL, Yao M et al. Acute myeloid leukemia bearing t(7;11)(p15;p15) is a distinct cytogenetic entity with poor outcome and a distinct mutation profile: comparative analysis of 493 adult patients. Leukemia 2009; 23: 1303–1310.
Taketani T, Taki T, Nakamura T, Kobayashi Y, Ito E, Fukuda S et al. High frequencies of simultaneous FLT3-ITD, WT1 and KIT mutations in hematological malignancies with NUP98-fusion genes. Leukemia 2010; 24: 1975–1977.
Qiao Q, Li Y, Chen Z, Wang M, Reinberg D, Xu RM . The structure of NSD1 reveals an autoregulatory mechanism underlying histone H3K36 methylation. J Biol Chem 2011; 286: 8361–8368.
van Zutven LJ, Onen E, Velthuizen SC, van Drunen E, von Bergh AR, van den Heuvel-Eibrink MM et al. Identification of NUP98 abnormalities in acute leukemia: JARID1A (12p13) as a new partner gene. Genes Chromosomes Cancer 2006; 45: 437–446.
de Rooij JD, Masetti R, van den Heuvel-Eibrink MM, Cayuela JM, Trka J, Reinhardt D et al. Recurrent genetic abnormalities can be used for risk-group stratification in pediatric AMKL: results of a retrospective intergroup study. Blood 2016; 127: 3424–3430.
Ahuja HG, Felix CA, Aplan PD . The t(11;20)(p15;q11) chromosomal translocation associated with therapy-related myelodysplastic syndrome results in an NUP98-TOP1 fusion. Blood 1999; 94: 3258–3261.
Fasan A, Haferlach C, Alpermann T, Kern W, Haferlach T, Schnittger S . A rare but specific subset of adult AML patients can be defined by the cytogenetically cryptic NUP98-NSD1 fusion gene. Leukemia 2013; 27: 245–248.
Thol F, Kolking B, Hollink IH, Damm F, van den Heuvel-Eibrink MM, Michel Zwaan C et al. Analysis of NUP98/NSD1 translocations in adult AML and MDS patients. Leukemia 2013; 27: 750–754.
Lavallee VP, Lemieux S, Boucher G, Gendron P, Boivin I, Girard S et al. Identification of MYC mutations in acute myeloid leukemias with NUP98-NSD1 translocations. Leukemia 2016; 30: 1621–1624.
Wang GG, Cai L, Pasillas MP, Kamps MP . NUP98-NSD1 links H3K36 methylation to Hox-A gene activation and leukaemogenesis. Nat Cell Biol 2007; 9: 804–812.
Wang GG, Song J, Wang Z, Dormann HL, Casadio F, Li H et al. Haematopoietic malignancies caused by dysregulation of a chromatin-binding PHD finger. Nature 2009; 459: 847–851.
Thanasopoulou A, Tzankov A, Schwaller J . Potent co-operation between the NUP98-NSD1 fusion and the FLT3-ITD mutation in acute myeloid leukemia induction. Haematologica 2014; 99: 1465–1471.
Acknowledgements
The study was initiated by a GFCH collaborative study regarding NUP98 rearrangement in pediatric AML. DE thanks the support of the research in his laboratory: 111 des Arts, Association Laurette Fugain, Société Française des Cancers des Enfants, Ligue Régionale contre le Cancer. Patients samples from the HIMIP collection were obtained after informed consent in accordance with the Declaration of Helsinki. According to the French law, HIMIP collection has been declared to the Ministry of Higher Education and Research (DC 2008-307 collection 1) and obtained a transfer agreement (AC 2008-129) after approbation by ethical committees (Comité de Protection des Personnes Sud-Ouest et Outremer II and APHP ethical committee). Clinical and biological annotations of the samples have been declared to the CNIL (Comité National Informatique et Libertés).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Supplementary information
Rights and permissions
About this article
Cite this article
Struski, S., Lagarde, S., Bories, P. et al. NUP98 is rearranged in 3.8% of pediatric AML forming a clinical and molecular homogenous group with a poor prognosis. Leukemia 31, 565–572 (2017). https://doi.org/10.1038/leu.2016.267
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2016.267
This article is cited by
-
Acute myeloid leukemia with rare recurring translocations—an overview of the entities included in the international consensus classification
Annals of Hematology (2024)
-
Phase separations in oncogenesis, tumor progressions and metastasis: a glance from hallmarks of cancer
Journal of Hematology & Oncology (2023)
-
Nuclear transport proteins: structure, function, and disease relevance
Signal Transduction and Targeted Therapy (2023)
-
A novel gene fusion RUNX1/ZNF423 promotes leukemic relapse of NUP98-rearranged AML
Leukemia (2023)
-
Epigenetic regulation in hematopoiesis and its implications in the targeted therapy of hematologic malignancies
Signal Transduction and Targeted Therapy (2023)