Abstract
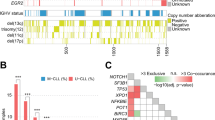
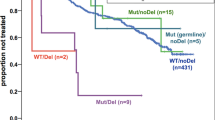
Through the European Research Initiative on chronic lymphocytic leukemia (CLL) (ERIC), we screened 3490 patients with CLL for mutations within the NOTCH1 (n=3334), SF3B1 (n=2322), TP53 (n=2309), MYD88 (n=1080) and BIRC3 (n=919) genes, mainly at diagnosis (75%) and before treatment (>90%). BIRC3 mutations (2.5%) were associated with unmutated IGHV genes (U-CLL), del(11q) and trisomy 12, whereas MYD88 mutations (2.2%) were exclusively found among M-CLL. NOTCH1, SF3B1 and TP53 exhibited variable frequencies and were mostly enriched within clinically aggressive cases. Interestingly, as the timespan between diagnosis and mutational screening increased, so too did the incidence of SF3B1 mutations; no such increase was observed for NOTCH1 mutations. Regarding the clinical impact, NOTCH1 mutations, SF3B1 mutations and TP53 aberrations (deletion/mutation, TP53ab) correlated with shorter time-to-first-treatment (P<0.0001) in 889 treatment-naive Binet stage A cases. In multivariate analysis (n=774), SF3B1 mutations and TP53ab along with del(11q) and U-CLL, but not NOTCH1 mutations, retained independent significance. Importantly, TP53ab and SF3B1 mutations had an adverse impact even in U-CLL. In conclusion, we support the clinical relevance of novel recurrent mutations in CLL, highlighting the adverse impact of SF3B1 and TP53 mutations, even independent of IGHV mutational status, thus underscoring the need for urgent standardization/harmonization of the detection methods.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Fabbri G, Rasi S, Rossi D, Trifonov V, Khiabanian H, Ma J et al. Analysis of the chronic lymphocytic leukemia coding genome: role of NOTCH1 mutational activation. J Exp Med 2011; 208: 1389–1401.
Puente XS, Pinyol M, Quesada V, Conde L, Ordonez GR, Villamor N et al. Whole-genome sequencing identifies recurrent mutations in chronic lymphocytic leukaemia. Nature 2011; 475: 101–105.
Quesada V, Conde L, Villamor N, Ordonez GR, Jares P, Bassaganyas L et al. Exome sequencing identifies recurrent mutations of the splicing factor SF3B1 gene in chronic lymphocytic leukemia. Nat Genet 2012; 44: 47–52.
Wang L, Lawrence MS, Wan Y, Stojanov P, Sougnez C, Stevenson K et al. SF3B1 and other novel cancer genes in chronic lymphocytic leukemia. N Engl J Med 2011; 365: 2497–2506.
Stilgenbauer S, Schnaiter A, Paschka P, Zenz T, Rossi M, Dohner K et al. Gene mutations and treatment outcome in chronic lymphocytic leukemia: results from the CLL8 trial. Blood 2014; 123: 3247–3254.
Schnaiter A, Paschka P, Rossi M, Zenz T, Buhler A, Winkler D et al. NOTCH1, SF3B1, and TP53 mutations in fludarabine-refractory CLL patients treated with alemtuzumab: results from the CLL2H trial of the GCLLSG. Blood 2013; 122: 1266–1270.
Rossi D, Bruscaggin A, Spina V, Rasi S, Khiabanian H, Messina M et al. Mutations of the SF3B1 splicing factor in chronic lymphocytic leukemia: association with progression and fludarabine-refractoriness. Blood 2011; 118: 6904–6908.
Cortese D, Sutton LA, Cahill N, Smedby KE, Geisler C, Gunnarsson R et al. On the way towards a 'CLL prognostic index': focus on TP53, BIRC3, SF3B1, NOTCH1 and MYD88 in a population-based cohort. Leukemia 2013; 28: 710–713.
Del Giudice I, Rossi D, Chiaretti S, Marinelli M, Tavolaro S, Gabrielli S et al. NOTCH1 mutations in +12 chronic lymphocytic leukemia (CLL) confer an unfavorable prognosis, induce a distinctive transcriptional profiling and refine the intermediate prognosis of +12 CLL. Haematologica 2012; 97: 437–441.
Jeromin S, Weissmann S, Haferlach C, Dicker F, Bayer K, Grossmann V et al. SF3B1 mutations correlated to cytogenetics and mutations in NOTCH1, FBXW7, MYD88, XPO1 and TP53 in 1160 untreated CLL patients. Leukemia 2014; 28: 108–117.
Oscier DG, Rose-Zerilli MJ, Winkelmann N, Gonzalez de Castro D, Gomez B, Forster J et al. The clinical significance of NOTCH1 and SF3B1 mutations in the UK LRF CLL4 trial. Blood 2013; 121: 468–475.
Rossi D, Fangazio M, Rasi S, Vaisitti T, Monti S, Cresta S et al. Disruption of BIRC3 associates with fludarabine chemorefractoriness in TP53 wild-type chronic lymphocytic leukemia. Blood 2012; 119: 2854–2862.
Rossi D, Rasi S, Spina V, Bruscaggin A, Monti S, Ciardullo C et al. Integrated mutational and cytogenetic analysis identifies new prognostic subgroups in chronic lymphocytic leukemia. Blood 2013; 121: 1403–1412.
Weissmann S, Roller A, Jeromin S, Hernandez M, Abaigar M, Hernandez-Rivas JM et al. Prognostic impact and landscape of NOTCH1 mutations in chronic lymphocytic leukemia (CLL): a study on 852 patients. Leukemia 2013; 27: 2393–2396.
Villamor N, Conde L, Martinez-Trillos A, Cazorla M, Navarro A, Bea S et al. NOTCH1 mutations identify a genetic subgroup of chronic lymphocytic leukemia patients with high risk of transformation and poor outcome. Leukemia 2013; 27: 1100–1106.
Mansouri L, Cahill N, Gunnarsson R, Smedby KE, Tjonnfjord E, Hjalgrim H et al. NOTCH1 and SF3B1 mutations can be added to the hierarchical prognostic classification in chronic lymphocytic leukemia. Leukemia 2013; 27: 512–514.
Schuh A, Becq J, Humphray S, Alexa A, Burns A, Clifford R et al. Monitoring chronic lymphocytic leukemia progression by whole genome sequencing reveals heterogeneous clonal evolution patterns. Blood 2012; 120: 4191–4196.
Schwaederle M, Ghia E, Rassenti LZ, Obara M, Dell'Aquila ML, Fecteau JF et al. Subclonal evolution involving SF3B1 mutations in chronic lymphocytic leukemia. Leukemia 2013; 27: 1214–1217.
Landau DA, Carter SL, Stojanov P, McKenna A, Stevenson K, Lawrence MS et al. Evolution and impact of subclonal mutations in chronic lymphocytic leukemia. Cell 2013; 152: 714–726.
Trbusek M, Smardova J, Malcikova J, Sebejova L, Dobes P, Svitakova M et al. Missense mutations located in structural p53 DNA-binding motifs are associated with extremely poor survival in chronic lymphocytic leukemia. J Clin Oncol 2011; 29: 2703–2708.
Balatti V, Bottoni A, Palamarchuk A, Alder H, Rassenti LZ, Kipps TJ et al. NOTCH1 mutations in CLL associated with trisomy 12. Blood 2012; 119: 329–331.
Lopez C, Delgado J, Costa D, Conde L, Ghita G, Villamor N et al. Different distribution of NOTCH1 mutations in chronic lymphocytic leukemia with isolated trisomy 12 or associated with other chromosomal alterations. Genes Chromosomes Cancer 2012; 51: 881–889.
Dohner H, Stilgenbauer S, Benner A, Leupolt E, Krober A, Bullinger L et al. Genomic aberrations and survival in chronic lymphocytic leukemia. N Engl J Med 2000; 343: 1910–1916.
Gunnarsson R, Isaksson A, Mansouri M, Goransson H, Jansson M, Cahill N et al. Large but not small copy-number alterations correlate to high-risk genomic aberrations and survival in chronic lymphocytic leukemia: a high-resolution genomic screening of newly diagnosed patients. Leukemia 2010; 24: 211–215.
Agathangelidis A, Darzentas N, Hadzidimitriou A, Brochet X, Murray F, Yan XJ et al. Stereotyped B-cell receptors in one-third of chronic lymphocytic leukemia: a molecular classification with implications for targeted therapies. Blood 2012; 119: 4467–4475.
Murray F, Darzentas N, Hadzidimitriou A, Tobin G, Boudjogra M, Scielzo C et al. Stereotyped patterns of somatic hypermutation in subsets of patients with chronic lymphocytic leukemia: implications for the role of antigen selection in leukemogenesis. Blood 2008; 111: 1524–1533.
Hallek M, Cheson BD, Catovsky D, Caligaris-Cappio F, Dighiero G, Dohner H et al. Guidelines for the diagnosis and treatment of chronic lymphocytic leukemia: a report from the International Workshop on Chronic Lymphocytic Leukemia updating the National Cancer Institute-Working Group 1996 guidelines. Blood 2008; 111: 5446–5456.
Xu L, Hunter ZR, Yang G, Zhou Y, Cao Y, Liu X et al. MYD88 L265P in Waldenstrom macroglobulinemia, immunoglobulin M monoclonal gammopathy, and other B-cell lymphoproliferative disorders using conventional and quantitative allele-specific polymerase chain reaction. Blood 2013; 121: 2051–2058.
Rose-Zerilli MJ, Forster J, Parker H, Parker A, Rodriguez AE, Chaplin T et al. ATM mutation rather than BIRC3 deletion and/or mutation predicts reduced survival in 11q-deleted chronic lymphocytic leukemia: data from the UK LRF CLL4 trial. Haematologica 2014; 99: 736–742.
Shedden K, Li Y, Ouillette P, Malek SN . Characteristics of chronic lymphocytic leukemia with somatically acquired mutations in NOTCH1 exon 34. Leukemia 2012; 26: 1108–1110.
Gentien D, Kosmider O, Nguyen-Khac F, Albaud B, Rapinat A, Dumont AG et al. A common alternative splicing signature is associated with SF3B1 mutations in malignancies from different cell lineages. Leukemia 2014; 28: 1355–1357.
Rahal R, Frick M, Romero R, Korn JM, Kridel R, Chan FC et al. Pharmacological and genomic profiling identifies NF-kappaB-targeted treatment strategies for mantle cell lymphoma. Nat Med 2014; 20: 87–92.
Oscier DG, Gardiner AC, Mould SJ, Glide S, Davis ZA, Ibbotson RE et al. Multivariate analysis of prognostic factors in CLL: clinical stage, IGVH gene mutational status, and loss or mutation of the p53 gene are independent prognostic factors. Blood 2002; 100: 1177–1184.
Krober A, Seiler T, Benner A, Bullinger L, Bruckle E, Lichter P et al. V(H) mutation status, CD38 expression level, genomic aberrations, and survival in chronic lymphocytic leukemia. Blood 2002; 100: 1410–1416.
Acknowledgements
We are grateful to the European Research Initiative in CLL steering board for all their support during this project. We thank Jitka Malcikova and Karla Plevova for their valuable comments and help in sample management and to Jana Smardova for FASAY analysis. This research was funded by the Nordic Cancer Union, the Swedish Cancer Society, the Swedish Research Council and Lion’s Cancer Research Foundation, Uppsala; Cancer Research UK, Leukaemia and Lymphoma Research, Kay Kendall Leukaemia Fund, United Kingdom; Associazione Italiana per la Ricerca sul Cancro (AIRC) (Investigator grant and Molecular Clinical Oncology Program 5xMille no. 9965, no. 10007), Milano, Italy; Ricerca Finalizzata 2010, Ministero della Salute, Roma, Italy; the ENosAI project (code 09SYN-13-880) co-funded by the EU and the Hellenic General Secretariat for Research and Technology, Greece; by project MSMT-CR VaVPI CZ.1.05/1.1.00/02.0068 (CEITEC), research grant IGA-MZ-CR NT13493-4/2012 and 7th FP Health project NGS-PTL/2012-2015/no. 306242 of European Commission and MSMT-CR (2013-2015, no. 7E13008), and the ICGC-CLL Genome Project from the Spanish Ministry of Economy and Competitivity (MINECO) through the Instituto de Salud Carlos III (RTICC RD12/0036/0023 and PIE PI043043), Spain. Andreas Agathangelidis is a recipient of a fellowship by Associazione Italiana per la Ricerca sul Cancro AIRC (Triennial fellowship ‘Guglielmina Lucatello é Gino Mazzega’).
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
KS receives research support from Roche SA. JCS has financial support from Hoffman La Roche.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Supplementary information
Rights and permissions
About this article
Cite this article
Baliakas, P., Hadzidimitriou, A., Sutton, LA. et al. Recurrent mutations refine prognosis in chronic lymphocytic leukemia. Leukemia 29, 329–336 (2015). https://doi.org/10.1038/leu.2014.196
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2014.196
This article is cited by
-
Different prognostic impact of recurrent gene mutations in chronic lymphocytic leukemia depending on IGHV gene somatic hypermutation status: a study by ERIC in HARMONY
Leukemia (2023)
-
Immunoglobulin gene sequence analysis in chronic lymphocytic leukemia: the 2022 update of the recommendations by ERIC, the European Research Initiative on CLL
Leukemia (2022)
-
Identification of CD105 (endoglin) as novel risk marker in CLL
Annals of Hematology (2022)
-
Targeting oncogenic Notch signaling with SERCA inhibitors
Journal of Hematology & Oncology (2021)
-
Biclonal lymphoproliferative disorders: another association with NOTCH1-mutated chronic lymphocytic leukaemias
Irish Journal of Medical Science (1971 -) (2021)