Abstract
Theragnostics represent cutting-edge, multi-disciplinary strategies that combine diagnostics with therapeutics in order to generate personalized therapies that improve patient outcome. In oncology, the approach is aimed at more accurate diagnosis of cancer, optimization of patient selection to identify those most likely to benefit from a specific therapy and to generate effective therapeutics that enhance patient survival. MicroRNAs (miRNAs) are master regulators of the human genome that orchestrate myriad cellular pathways to control growth during physiologic and pathologic conditions. Compelling evidence shows that miRNA deregulation promotes events linked to tumor initiation, metastasis and drug resistance as seen in multiple myeloma (MM), an invariably fatal hematologic malignancy. miRNAs are readily detected in body fluids, for example, serum, plasma, urine, as well as circulating tumor cells to demonstrate their potential as readily accessible, non-invasive diagnostic and prognostic biomarkers and potential therapeutics. Specific miRNAs are aberrantly expressed early in myelomagenesis and may more readily detect high-risk disease than current methods. Although only recently discovered miRNAs have rapidly advanced from preclinical studies to evaluation in human clinical trials. The development of miRNA theragnostics should provide widely applicable tools for the targeted delivery of personalized medicines to improve the outcome of patients with MM.
Similar content being viewed by others
Introduction
Theragnostics (a portmanteau of therapeutics and diagnostics) incorporate multiple disciplines, for example, bioinformatics, pharmacogenomics, proteomics and metabolomics, to design accurate diagnostic assays with a targeted therapy that is selected based upon testing results.1, 2 The strategy offers the opportunity to generate an advanced molecular understanding of the disease, to develop more effective molecular targets and to design therapeutic agents based upon patient-specific biology of disease. As a result, patients should more efficiently receive personalized, targeted therapy tailored to specific biological and molecular features of their tumor.
Accurate diagnostic testing is also designed to avoid the unnecessary treatment of patients that are not likely to demonstrate a significant clinical response or for whom a specific therapy is not appropriate. The results should lead to the earlier treatment of high-risk forms of cancers such as multiple myeloma (MM) that may not respond to standard-of-care therapy or that require earlier treatment at a less advanced stage for significant clinical improvement. More broadly, theragnostics can increase drug efficacy and safety while significantly reducing pharmaceutical and overall health-care costs. Theragnostics require further bioinformatic and genomic advances before wide-scale use as standardized tools in predictive medicine, but are key to the development and implementation of personalized therapies that eventually will improve the overall survival of cancer patients.
MicroRNAs (miRNAs) are an abundant class of naturally occurring, evolutionarily conserved, RNA molecules that are not translated into proteins but serve an important function to negatively regulate gene expression. miRNAs specifically bind to complementary regions within messenger RNAs (mRNAs) to promote their degradation and/or to inhibit translation.3 By degrading or blocking the translation of mRNA targets, miRNAs modulate the expression of more than half of the protein-coding genes within the human genome.4 Genomic and bioinformatic analyses indicate that miRNAs have a key role in gene expression to control diverse biological processes such as cellular growth, differentiation, development and apoptosis.5 Hence, deregulation of a single miRNA has profound consequences because each miRNA can bind and control a number of mRNAs.
A multitude of recent studies link the abnormal expression of miRNAs to the pathogenesis of many solid tumor and hematological malignancies.4, 5, 6, 7, 8 The context-dependent loss of tumor-suppressive miRNAs enhances the expression of target oncogenes, whereas increased expression of oncogenic miRNAs represses target tumor-suppressor genes (Figure 1).7, 8, 9 miRNAs are dispersed throughout the human genome, but nearly half appear to be located within fragile sites or regions with altered expression in cancers.10 Moreover, a significant number of miRNAs are encoded in genomic regions associated with cancer to suggest that altered expression has a causal relationship with tumourigenesis. In summary, the overexpression, amplification and/or deletion of miRNAs and miRNA-mediated modification of epigenetic silencing impact events that contribute to oncogenesis.11
Therapeutic application of miRNAs. In tumor cells, miRNAs can be either increased to promote oncogenic pathways (oncomirs) or tumor-suppressive miRNAs are decreased to confer a loss-of-function phenotype. Antagomirs are synthetically engineered RNA molecules that are complementary to oncomirs and designed to inhibit the tumor-promoting effect of oncomirs. Synthetically engineered miRNA replacements can be transfected into tumor cells to restore the loss-of-function phenotype. miRNA replacement corrects miRNA deficiencies to restore the loss-of-function and provide therapeutic benefit.
MiRNA deregulation in MM
Myeloma is a clonal B-cell malignancy characterized by the aberrant accumulation of plasma cells (PCs) within bone marrow (BM) and extramedullary sites.12, 13 Myeloma arises from the multifocal proliferation of long-lived PCs and, despite all available therapies, remains invariably fatal.14, 15 Autologous stem-cell transplantation and immunomodulatory agents, for example, lenalidomide and pomalidamide, and the proteasome inhibitor bortezomib have improved clinical management to extend overall survival for selected myeloma patients from <3 to >5 years.16, 17 Although the precise molecular events that generate MM remain incompletely understood, our understanding of events that underlie its development have greatly advanced.18 Moreover, patients’ response to bortezomib-based regimens is highly variable, and drug resistance inevitably emerges even in patients who initially responded. The incidence and prevalence of MM are steadily rising in Western countries because of an aging population combined with the increased overall survival of patients with MM. Therefore, there is an urgent, unmet need to more accurately detect high-risk MM at an earlier stage and to clinically advance practice.
Genomic and bioinformatic computational tools now allow researchers to predict and validate the biological role of individual miRNAs during tumorigenesis, to identify miRNA target genes and to develop miRNA expression signatures that distinguish cancerous from non-cancerous tissue.19 Global miRNA profiling has revealed drastic changes in expression of multiple miRNAs in many common human cancers. miRNA expression signatures also provide a more accurate method of classifying cancer subtypes than transcriptome profiling and allow classification of different stages in tumor progression and can predict disease outcome in some cases. Existing algorithms combined with intracellular network analysis can be supported with phylogenetic footprinting and functional studies to better define the role of miRNAs during tumorigenesis. MiRNA regulation is essential to many cellular processes, and escape from this regulatory network seems to be a common characteristic of several disease processes and malignant transformation. For example, the first evidence of aberrant miRNA expression in a human cancer was described in B-cell chronic lymphocytic leukemia, wherein hemizygous and/or homozygous chromosomal deletion at the 13q14 locus resulted in the loss or reduction of miRNA-15 and miRNA-16 expression (Table 1).20 The observation that various miRNAs are specifically overexpressed or lost in tumors when compared with normal tissues supports a direct, causal link between miRNAs and myelomagenesis.21 Archival, unstained BM slides and fresh BM samples are typically the preferred source of miRNA for MM analysis by microarray profiling and PCR amplification. Improved extraction methods are now available to analyze miRNA obtained from freshly obtained BM aspirates, BM biopsy samples and tumor cells circulating in the peripheral blood. These technical advances have improved the quality of miRNA extracted and elevated the reproducibility of such studies. A novel method has been developed using a protocol for isolation of total RNA from archival samples that includes an overnight proteinase K digestion followed by a column based kit extraction. This method appears robust enough to allow the use of such samples for disease classification, diagnosis and to predict prognosis.22 Moreover, extracellular miRNAs are present in the serum, plasma, saliva, urine and other body fluids and have recently been shown to be associated with various pathological conditions including MM. miRNAs circulate within the bloodstream in a highly stable, extracellular form to more readily provide genetic material for relevant studies. Therefore, miRNAs represent readily accessible tools and an unprecedented means to diagnose diseases at an earlier stage than was previously available.
Several studies have identified miRNAs that are deregulated during myelomagenesis, and subsequent studies have explored the role of miRNAs as diagnostics to detect disease or to monitor myeloma progression.23, 24, 25, 26, 27, 28, 29, 30, 31 These studies have found that the vast majority of miRNAs that are aberrantly expressed in MM cells are upregulated compared with their expression in normal PCS.23 A recent study compared miRNA expression profiles (miREP) of 52 newly diagnosed MM patients with that obtained from PCs of two healthy donors. Among 464 miRNAs analyzed, 95 had a higher mean expression in PC samples of MM patients compared with those of healthy donors.24 In patients with high-risk MM, a global increase in specific miRNAs and overexpression of EIF2C2/AGO2 was also observed. In related studies, miRNA-15a was downregulated in relapsed and/or refractory MM and found to regulate tumor progression in MM cell lines (MMCLs) (Table 2).25 Separately, it was found miRNAs-15a and miRNAs-16 expression levels were often elevated in the PCs from newly diagnosed MM patients in comparison with healthy PCs independent of chromosome 13 deletions by karyotyping.26 The miRNA-17–92 cluster, which targets the apoptosis facilitator Bcl-2, was also reported to confer tumourigenicity in MM.27 MiRNA-29b has been reported to downregulate Mcl-1 and to induce apoptosis of myeloma cells.28 The miRNA-193b-365 cluster is overexpressed in MM and three miRNAs—720, 1308 and 1246—were significantly higher in PCs of myeloma patients than healthy controls.29, 30
A comprehensive miREP study compared PCs from normal donors, those with monoclonal gammopathy of unknown significance (MGUS), MM patients and MMCLs using a custom-made microarray chip with qRT-PCR.31 MGUS patients had a group of upregulated miRNAs with known oncogenic potential including miRNA-21 and the miRNA-106–25 cluster. MiRNA-21 expression is controlled in a STAT-3-interleukin (IL)-6-dependent manner and treatment of IL-6-dependent myeloma cells with IL-6 activated STAT-3, which in turn enhanced miRNA-21 transcription.32 Importantly, ectopic expression of miRNA-21 was sufficient to sustain the growth of IL-6-dependent cells in the absence of exogenous IL-6.
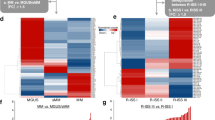
Distinct MM subtypes have been characterized based upon non-overlapping chromosomal translocations detected and correlative gene expression signatures based upon PCs isolated from MM patients.33, 34 Stratification of MM patients by gene expression profiles (GEPs) into subgroups illustrates that MM is not a single disease but represents remarkably distinct forms of the same disease. MM is characterized by very complex cytogenetic aberrations that affect prognosis and affect miRNA expression as well. Not surprisingly, early studies indicate that MM patients can also be stratified based upon distinct miREPs. The upregulation of miRNAs-1 and 133a correlates with MM patients that display the translocation (t)(14;16) to support the premise that deregulated miRNA expression is directly linked to specific chromosomal aberrations.35 miRNAs-221 and 99b were similarly associated with specific chromosomal aberrations.36 Myeloma is characterized by chromosomal abnormalities that vary in prognostic significance. Specific miRNA signatures were associated with the most common translocations seen in MM patients – t(4;14), t(11;14) and del(13q) (Table 337). Distinct miREP patterns were determined from MM patient PCs and compared within the molecular subtypes determined by GEP using highly purified tumor samples from 40 patients.38 The study was designed to correlate the occurrence of specific miRNAs the expression of predicted target mRNAs. Results were then integrated into the myeloma subtypes based upon chromosomal translocations and also correlated to miRNA expression. The miRNA expression patterns that were identified were associated mainly with IgH translocations. Patients that harbored t (4;14) displayed specific overexpression of Let-7e, miRNA-125a-5p and miRNA-99b. However, the occurrence of other chromosomal changes, for example, chromosome 1q gain, chromosome 13q and 17p deletions, and hyperdiploid were weakly correlative with specific miRNA signatures.37, 41
Specific miRNA signatures were also associated with Igh t (4;14) and t (11;14) and Del (13q). Expression levels of miRNAs were distinct between translocation-based subtypes and correctly predicted these abnormalities in >85% of cases. miRNAs associated with different MM isotopes including IgG, IgA light chain MM. miRNA profiles may in fact be a better discriminator of myeloma biology and risk-stratification than GEP or cytogenetic. The studies strongly suggest the potential diagnostic/prognostic utility of miRNA for the identification of patients with MGUS likely to progress to MM.39, 40 Future studies that employ a greater number of patient samples, more advanced miRNA microarray platforms and an integrative analysis based upon computational target prediction and miRNA/mRNA profiling could be useful to correlate miRNA expression with chromosomal translocations specific to MM and to unravel the complex network of functional miRNAs and their mRNA targets.
MiRNA deregulation in MGUS
MGUS is characterized by the restricted clonal expansion of antibody-secreting PCs in BM and progresses to MM at a rate of ∼1% per year.42 GEP has revealed major differences between the PCs isolated from healthy donors and individuals with MGUS but the distinctions between the PCs of MGUS and MM patients were less pronounced.39, 42, 43 miREP studies have also revealed that individual miRNAs were expressed differently in PCs from MGUS compared with normal donors.31 miRNAs-21, 181a/181b and the 106b-25 cluster were upregulated in PCs from both MGUS and MM patients compared with those of healthy donors. In contrast, miRNA-32, 19a/b and the 17–92 cluster were upregulated in MMCLs and PCs from MM patients but not in MGUS or healthy donors. Unsupervised cluster analysis revealed that MM and MGUS samples have miREPs distinct from CD138+ cells from healthy donors.37 A comparison of miRNAs with those aberrantly expressed in other B-cell and T-cell malignancies revealed a surprising (40%) degree of similarity to suggest the existence of a common miRNA signature among these diseases (Table 1). In addition, 39 miRNAs were associated with MGUS, 23 (59%) of which were aberrantly expressed in MM, suggesting common miRNA-dependent events during the progression from MGUS to MM.44
miRNA diagnostics and prognostics in MM
Translational studies suggest that individual tumor types commonly express miRNA signatures that can be useful in categorizing, detecting and predicting disease outcome.45 Transcriptional profiling suggests that miREP may successfully classify tumor types and that the miRNA repertoire is a stable and unique feature that represents distinct tumor types and stages of differentiation.46, 47 Because of their stability in a number of body fluids, for example, plasma, serum, urine and the relative ease for routine quantification, miRNAs are emerging as important diagnostic and prognostic tools. miRNAs hold great promise as readily accessible biomarkers to facilitate early disease detection and to monitor disease progression. miRNAs-21 and 10b have been implicated in the progression of many tumor types, including the lethal, metastatic phase of breast cancer.48, 49 HOX9, c-Myc, Bcl-2 and SHP1/SHP2 represent targets of miRNA-146b, 140, 145, 125a, 151, 223, 155 and Let-7f, and changes in expression of these miRNAs may be involved in myelomagenesis and associated with overall prognosis.50 miRNA-17–92 clusters are other pivotal miRNAs activated by Myc and are highly linked to the progression of MM. miRNA-17–92 cluster expression is not decreased in MM with chromosome 13 deletion and contributes to a poor MM prognosis.27 Gao et al.41 reported that in addition to miRNA-17–92 cluster expression, miRNA-15a and 16–1 are also linked to poor prognosis in MM patients.41 Those with elevated miRNA-17, 20 and 92 levels had shorter progression-free survival than those with reduced miRNA levels. Higher miRNA-20a and 148a levels in myeloma patients are associated with a shorter relapse-free survival, to suggest a possible association between miRNA-20a and poor prognosis.37 Expression of miRNAs-153, 490, 455, 642, 500 and 296 is associated with a better event-free survival, whereas miRNAs-548d, 373, 554 and 888 expression correlates with a poorer outcome.37 Polymorphisms related to miRNAs (miRSNPs) also have a prognostic impact in myeloma, and two miRSNPs—KRT81 and XPO5—are prognostic after melphalan-based autologous stem-cell transplantation.51 The precise role of miRNAs in myeloma needs to be further elucidated, although they are predicted to be involved in PC growth, survival, growth factor response, homing, drug resistance and BM interaction.
Similar to miRNA’s that exist within tumor cells, cell-free, circulating miRNAs have also been detected and proposed to function as 'extracellular communication RNAs' that control proliferation, differentiation and metastasis.30, 52 As miRNAs are freely circulating and highly stable within the bloodstream, there is great interest for their use as biomarkers. miRNAs represent an important new class of biomarkers whose concentration in blood may emulate tissue- and disease-specific expression profiles. Circulating miRNAs escape degradation by residing within microvesicles, exosomes, microparticles and/or apoptotic bodies, although the formation of protein-miRNA complexes has also been proposed.52 In addition, miRNAs have been isolated from circulating tumor cells (CTCs) with tremendous utility as naturally occurring, blood-based biomarkers for non-invasive diagnosis and to monitor therapeutic response. CTCs are detectable within the bloodstream at the earliest stages of MM and can determine the genetic evolution of the tumor during the course of treatment. In MM, malignant PCs have been isolated from BM biopsies and CTCs.53, 54 Although myeloma cells are found throughout BM, these malignant cells are irregularly scattered in patches throughout the skeletal system. Therefore, BM biopsy (repeatedly at the iliac crest) represents an imprecise method to detect and monitor disease status. Technological advances allow for improved cell separation and standardized, reproducible, highly sensitive methods to enumerate CTCs in the peripheral blood. CTCs offer an unprecedented approach to incorporate miRNA studies and unravel the molecular basis of MM, drug resistance and disease progression.55 Although research on CTCs and circulating miRNAs is still in its infancy, robust data will bring rigor to bear in this field and advance the overall miRNA network.
The accurate diagnosis and frequent surveillance of MM can be improved using miRNA obtained from patient BM biopsies, circulating miRNA or miRNA from CTCs. miRNAs offer potential as diagnostics to distinguish the characteristic expression signatures that distinguish MM from normal or MGUS PCs. As each of the MM subtypes also display their own unique miRNA signatures, these predictors may eventually replace the current clinical and biological markers used to diagnose MM. Comprehensive characterization of miRNA signatures will require the inclusion of a maximum number of different miRNAs and will be achieved most readily through high-throughput screening platforms combined with state-of-the art bioinformatics.
miRNA therapeutics for MM
The past decade has witnessed rapid progress in treatment for MM, but the landscape looks even more promising with newer agents that exploit novel targets and previously unaddressed oncogenic pathways. Remarkably, eight regimens have been approved in the past decade—two within the past year—to collectively increase overall survival threefold (with additional gains anticipated).73 Newer agents may be better tolerated, provide more sustained responses and benefit a broader spectrum of patients.57, 58, 59, 60, 61 Although the first human miRNA was discovered in 2000, the miRNA field has rapidly progressed beyond the discovery phase, advanced through translational studies and entered into clinical trial evaluation.57 Certain miRNAs are overexpressed in cancers and demonstrate oncogenic properties (oncomirs), whereas others are lost during tumourigenesis and display tumor-suppressive properties (Figure 1). This provides a two-pronged approach for anti-cancer therapeutic application: to inhibit oncomirs using RNA molecules complementary (antagomirs) to the oncomir or to replace tumor-suppressive miRNAs that are reduced in tumors with the goal of restoring lost function. The artificial introduction of synthetically engineered miRNAs or the inhibition of endogenous miRNAs has provided additional clues to the function of miRNAs in cancer. Expression of exogenous, small RNA molecules, such as anti-oncomirs, has demonstrated a role for many individual miRNAs in cancer.58 Let-7 is the first identified anti-oncomir that functions as a ‘post-transcriptional-gatekeeper’ of specific genes that control cell growth. In lung cancer, certain oncogenes are downregulated by Let-7 to maintain normal cell progression.59 It appears that many miRNAs have a conserved function, and that their deregulation results in functional changes commonly seen in diverse tumor types.
miRNAs that display tumor-suppressive roles to repress the expression of growth-promoting genes are reduced in many tumors.60 Although miRNA antagonists are conceptually similar to other inhibitory strategies, for example, small-molecule enzyme inhibitors, restoring the function of a miRNA by replacement is a less well-characterized approach. MiRNA replacements represent a new research horizon and an innovative technological advancement to investigate the function of an individual miRNA with unmatched specificity and efficacy and to provide uniform potency at low concentrations and superior biological stability. Tumor-suppressor miRNAs that display a loss of function in cancer cells can be replaced by therapies that re-introduce engineered miRNA replacements into diseased tissues to reactivate pathways and drive a therapeutic response. In many cases, reactivation of these miRNA-regulated pathways interferes with oncogenic properties of the cell, blocks uncontrolled proliferation and induces the apoptotic cascade. The rationale for developing miRNA replacements is based upon the premise that aberrantly reduced miRNAs have a key role in the development of human disease, and that correcting these deficiencies by restoring miRNA functions will provide clinical benefit.61, 62 Therapeutically restoring the activity of tumor suppressors for clinical benefit has been investigated previously through gene therapy, but practical, widespread efficacy is lacking. Finally, miRNAs may be beneficial in cancer treatment as has been described for the therapeutic delivery of miRNA mimics in animal models. A recent study reported the antitumour activity of synthetic miRNA-34a as well as anti-proliferative activity, apoptotic effects and modulation of gene expression in clinically relevant xenograft models of myeloma.63
Concluding remarks
Rapidly emerging evidence points to the vast potential of miRNAs as theragnostics to advance the clinical management of MM. A goal is to now identify surrogate miRNAs as reliable diagnostics for early disease detection, as prognostics to detect patients with high-risk disease. The development of more effective diagnostics guided by bioinformatics, genomics, proteomics and pharmacogenomics represents a conceptual departure from traditional practice used in drug discovery. An advantage of miRNA theragnostics is the rational design of agents to identify patient subpopulations most likely to respond to a given therapy. A major challenge now is to decipher the role of individual miRNAs in the development of MM as well as related hematological disorders (Table 1). Importantly, many miRNAs deregulated in MM are similarly deregulated in related hematological disorders and malignancies. Future studies are needed to define the role of the broader family of non-coding (nc) RNAs, for example, long ncRNAs, small nuclear RNAs, in cancer. Clearly, there remains a need for further translational and clinical research to fully integrate the mass of information obtained from diverse technology platforms in meaningful way for physicians, patients and the health-care system. miRNAs represent highly promising, state-of-the-art tools with immediate translational impact and practical application at the point of patient care. MiRNA-based theragnostics will fuse diagnostics with targeted therapy as personalized medicines to improve patient management and outcome.
References
Ozdemir V, Williams-Jones B, Glatt SJ, Tsuang MT, Lohr JB, Reist C . Shifting emphasis from pharmacogenomics to theragnostics. Nat Biotechnol 2006; 24: 942–946.
Pene F, Courtine E, Cariou A, Mira JP . Toward theragnostics. Critical Care Med 2009; 37 (Suppl 1): S50–S58.
Lewis BP, Burge CB, Bartel DP . Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005; 120: 15–20.
Amiel J, de Pontual L, Henrion-Caude A . miRNA, development and disease. Adv Genet 2012; 80: 1–36.
Yu Z, Tozeren A, Pestell RG . Function of miRNAs in tumor cell proliferation, Alahari S (ed.), in MicroRNA in Cancer. Springer: Dodrecht. ISBN:978-94-007-4654-1. 2013, pp 13–27.
Redis RS, Berindan-Neagoe I, Pop VI, Calin GA . Non-coding RNAs as theranostics in human cancers. J Cell Biochem 2012; 113: 1451–1459.
Jansson MD, Lund AH . MicroRNA and cancer. Mol Oncol 2012; 6: 590–610.
Zhang B, Pan X, Cobb GP, Anderson TA . MicroRNAs as oncogenes and tumor suppressors. Dev Biol 2007; 302: 1–12.
He L, Thomson JM, Hemann MT, Hernando-Monge E, Mu D, Goodson S et al. A microRNA polycistron as a potential human oncogene. Nature 2005; 435: 828–833.
Calin GA, Sevignani C, Dumitru CD, Hyslop T, Noc Eh, Croce CM . Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc Natl Acad Sci USA 2004; 101: 2999–3004.
Ventura A, Jacks T . MicroRNAs and cancer: short RNAs go a long way. Cell 2009; 136: 586–591.
Bommert K, Bargou RC, Stuhmer T . Signaling and survival pathways in multiple myeloma. Eur J Cancer 2006; 42: 1574–1580.
Mahindra A, Hideshima T, Anderson KC . Multiple myeloma: biology of the disease. Blood Rev 2010; 24: S5–11.
Kyle RA, Rajkumar SV . Multiple myeloma. Blood 2008; 111: 2962–2972.
San-Miguel JF, Garcia-Sanz R, Gutierrez NC . Prognosis and Staging of Multiple MyelomaWiernik PH, Goldman JM, Dutcher J, Kyle RA, Neoplastic Diseases of the Blood 19 5. Springer: Dodrecht, 2013, pp 615–636.
Barlogie B, Shaughnessy J, Tricot G, Jacobson J, Zangari M, Anaissie E et al. Treatment of multiple myeloma. Blood 2004; 103: 20–32.
Palumbo A . Multiple myeloma. Curr Opin Oncol 2012; 24 (Suppl 2): S1–S2.
Bergsagel PL, Mateos MV, Gutierrez NC, Rajkumar SV, San Miguel JF . Improving overall survival and overcoming adverse prognosis in the treatment of cytogenetically high-risk multiple myeloma. Blood 2013; 121: 884–892.
Calin GA, Croce CM . MicroRNA signatures in human cancers. Nat Rev Cancer 2006; 6: 857–866.
Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Croce CM . Frequent deletions and down-regulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci USA 2002; 99: 15524–15529.
Voorhoeve PM, Agami R . Classifying microRNAs in cancer: the good, the bad and the ugly. Rev Cancer 2007; 1775: 274–282.
Morenos L, Saffery R, Mechinaud F, Ashley D, Elwood N et al. Evaluation of microRNA expression in patient bone marrow aspirate slides. PLoS One 2012; 7: e42951.
Corthals S . MicroRNA signatures characterize multiple myeloma patients. Leukemia 2011; 25: 1784–1789.
Zhou Y, Chen L, Barlogie B, Stephens O, Wu X, Williams DR et al. High risk myeloma is associated with global elevation of miRNAs and overexpression of EIF2C2/AGO2. Proc Natl Acad Sci US 2010; 107: 7904–7909.
Roccaro AM, Sacco A, Thompson B, Leleu X, Anderson KC . MicroRNAs 15a and 16 regulate tumor proliferation in multiple myeloma. Blood 2009; 113: 6669–6680.
Corthals SL, Jongen-Lavrencic M, de Knegt Y, Peeters JK, Beverloo HB, Lokhorst HM et al. Micro-RNA-15a and micro-RNA-16 expression and chromosome 13 deletions in multiple myeloma. Leuk Res 2010; 34: 677–681.
Chen L, Li C, Zhang R, Gao X, Qu X, Zhao M et al. miR-17-92 cluster microRNAs confers tumorigenicity in multiple myeloma. Cancer Lett 2011; 309: 62–70.
Zhang YK, Wang H, Leng Y, Li ZL, Yang YF, Xiao FJ et al. Overexpression of microRNA-29b induces apoptosis of multiple myeloma cells through down regulating Mcl-1. Biochem Biophys Res Commun 2011; 414: 233–239.
Unno K, Zhou Y, Zimmerman T, Platanias LC, Wickrema A . Identification of a novel microRNA cluster miR-193b-365 in multiple myeloma. Leuk Lymphoma 2009; 50: 1865–1871.
Jones CI, Zabolotskaya MV, King AJ, Stewart HJ, Horne GA, Chevassut TJ et al. Identification of circulating microRNAs as diagnostic biomarkers for use in multiple myeloma. Br J Cancer 2012; 107: 1987–1996.
Pichiorri F, De Luca L, Aqeilan RI . MicroRNAs: new players in multiple myeloma. Front Genet 2011; 2: 22.
Loffler D, Brocke-Heidrich K, Pfeifer G, Stocsits C, Hackermuller J, Kretzschmar AK et al. Interleukin-6 dependent survival of multiple myeloma cells involves the Stat3-mediated induction of microRNA-21 through a highly conserved enhancer. Blood 2007; 110: 1330–1333.
Avet-Loiseau H, Attal M, Moreau P, Charbonnel C, Garban F, Hulin C et al. Genetic abnormalities and survival in multiple myeloma: the experience of the Intergroupe Francophone du Myélome. Blood 2007; 109: 3489–3495.
Segges P, Braggio E . Genetic markers used for risk stratification in multiple myeloma. Genet Res Int 2011; 2011: 798089.
Gutiérrez NC, Sarasquete ME, Misiewicz-Krzeminska I, Delgado M, De Las Rivas J, Ticona FV et al. Deregulation of microRNA expression in the different genetic subtypes of multiple myeloma and correlation with gene expression profiling. Leukemia 2010; 24: 629–637.
Huang JJ, Yu J, Li JY, Liu YT, Zhong RQ . Circulating microRNA expression is associated with genetic subtype and survival of multiple myeloma. Med Oncol 2012; 29: 2402–2408.
Chi J, Ballabio E, Chen XH, Kusec R, Taylor S, Hay D et al. MicroRNA expression in multiple myeloma is associated with genetic subtype, isotype and survival. Biol Direct 2011; 6: 23.
Lionetti M, Biasiolo M, Agnelli L, Todoerti K, Mosca L, Fabris S et al. Identification of microRNA expression patterns and definition of a microRNA/mRNA regulatory network in distinct molecular groups of multiple myeloma. Blood 2009; 114: e20–e26.
Davies FE, Dring AM, Li C, Rawstron AC, Shammas MA, O’Connor SM et al. Insights into the multistep transformation of MGUS to myeloma using microarray expression analysis. Blood 2003; 102: 4504–4511.
Calvo KR, Landgren O, Roccaro AM, Ghobrial IM . Role of microRNAs from monoclonal gammopathy of undetermined significance to multiple myeloma. Semin Hematol 2011; 48: 39–45.
Gao X, Zhang R, Qu X, Zhao M, Zhang S, Wu H et al. MiR-15a, miR-16-1 and miR-17-92 cluster expression are linked to poor prognosis in multiple myeloma. Leuk Res 2012; 36: 1505–1509.
Landgren O, Kyle RA, Pfeiffer RM, Katzmann JA, Caporaso NE, Hayes RB et al. Monoclonal gammopathy of undetermined significance (MGUS) consistently precedes multiple myeloma: a prospective study. Blood 2009; 113: 5412–5417.
Zhan F, Hardin J, Kordsmeier B, Bumm K, Zheng M, Tian E et al. Global gene expression profiling of multiple myeloma, monoclonal gammopathy of undetermined significance, and normal bone marrow plasma cells. Blood 2002; 99: 1745–1757.
Anderson KC . Therapeutic advances in relapsed or refractory multiple myeloma. J Natl Compr Canc Netw 2013; 11 (Suppl 5): 676–679.
Tricoli JV, Jacobson JW . MicroRNA: Potential for cancer detection, diagnosis, and prognosis. Canc Res 2007; 67: 4553–4555.
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D et al. MicroRNA expression profiles classify human cancers. Nature 2005; 435: 834–838.
Olson P, Lu J, Zhang H, Shai A, Chun MG, Wang Y et al. MicroRNA dynamics in the stages of tumorigenesis correlate with hallmark capabilities of cancer. Genes Dev 2009; 23: 2152–2165.
Zhu S, Wu H, Wu F, Nie D, Sheng S, Mo YY . MicroRNA-21 targets tumor suppressor genes in invasion and metastasis. Cell Res 2008; 18: 350–359.
Ma L, Teruya-Feldstein J, Weinberg RA . Tumor invasion and metastasis initiated by microRNA-10b in breast cancer. Nature 2007; 449: 682–688.
Adamia S, Avet-Loiseau H, Amin SB, Moreau P, Minvielle SS, Treon S et al. Clinical and biological significance of microRNA profiling in patients with myeloma. J Clin Oncol 2009; 27(suppl 15): (abstract 8539).
de Larrea CF, Navarro A, Tejero R, Tovar N, Díaz T, Cibeira MT et al. Impact of MiRSNPs on survival and progression in patients with multiple myeloma undergoing autologous stem cell transplantation. Clin Cancer Res 2012; 18: 3697–3704.
Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci USA 2008; 105: 10513–10518.
Kiel K, Cremer FW, Rottenburger C, Kallmeyer C, Ehrbrecht E, Atzberger A et al. Analysis of circulating tumor cells in patients with multiple myeloma during the course of high-dose therapy with peripheral blood stem cell transplantation. Bone Marrow Transplant 1999; 23: 1019–1027.
Rigolin GM, Fraulini C, Ciccone M, Mauro E, Bugli EM, De Angeli C et al. Neoplastic circulating endothelial cells in multiple myeloma with 13q14 deletion. Blood 2006; 107: 2531–2535.
Cortez MA, Bueso-Ramos C, Ferdin J, Lopez-Berestein G, Sood AK, Calin GA . MicroRNAs in body fluids—the mix of hormones and biomarkers. Nat Rev Clin Oncol 2011; 8: 467–477.
Zanette DL, Rivadavia F, Molfetta GA, Barbuzano FG, Proto-Sigueira R, Silva-JR WA . miRNA expression profiles in chronic lymphocytic and acute lymphocytic leukemia. Braz J Med Biol Res 2007; 40: 1435–1440.
Yates LA, Norbury CJ, Gilbert RJ . The long and short of microRNA. Cell 2013; 153: 516–519.
Kwak PB, Iwasaki S, Tomari Y . The microRNA pathway and cancer. Cancer Sci 2010; 101: 2309–2315.
Kitade Y, Akao Y . MicroRNAs and their therapeutic potential for human diseases: microRNAs, miR-143 and −145, function as anti-oncomirs and the application of chemically modified miR-143as an anti-cancer drug. J Pharmacol Sci 2010; 114: 276–280.
Broderick JA, Zamore PD . MicroRNA therapeutics. Gene Ther 2011; 18: 1104–1110.
Bader AG, Brown D, Winkler M . The promise of microRNA replacement therapy. Cancer Res 2010; 70: 7027–7030.
Huang X, Schwind S, Yu B, Santhanam R, Wang H, Hoeleebauer P et al. Targeted delivery of microRNA-29b by transferrin-conjugated anionic lipopolyplex nanoparticles: a novel therapeutic strategy in acute myeloid leukemia. Clin Cancer Res 2012; 19: 2355–2367.
Di Martino MT, Leone E, Amodio N, Foresta U, Lionetti M, Pitari MR et al. Synthetic miR-34a mimics as a novel therapeutic agent for multiple myeloma: in Vitro and in vivo evidence. Clin Cancer Res 2012; 18: 6260–6270.
Weng L, Spencer BH, SoohHoo PT, Connors LH, O’Hara CJ, Seldin DC . Dysregulation of miRNAs in AL amyloidosis. Amyloid 2011; 18: 128–135.
Hodge LS, Elsawa SF, Grote DM, Price-Troska TL, Asmann YW, Fonseca R et al. MicroRNA expression in tumor cells from Waldenstrom's macroglobulinemia reflects both their normal and malignant cell counterparts. Blood Cancer J 2011; 1: e24.
Zhao H, Wang D, Du W, Gu D, Yang R . MicroRNA and leukemia: tiny molecule, great function. Crit Rev Oncol Hematol 2010; 74: 149–155.
Calin GA, Liu CG, Sevignani C, Ferracin M, Felli N, Dumitru CD et al. MicroRNA profiling reveals distinct signatures in B cell chronic lymphocytic leukemias. Proc Natl Acad Sci USA 2004; 101: 11755–11760.
Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E et al. Frequent deletions and down-regulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci USA 2002; 99: 15524–15529.
Marcucci G, Radmacher MD, Maharry K, Mrózek K, Ruppert AS, Paschka P et al. MicroRNA expression in cytogenetically normal acute myeloid leukemia. N Engl J Med 2008; 358: 1919–1928.
Takada S, Yamashita Y, Berezikov E, Hatanaka H, Fujiwara S-i, Kurashina K et al. MicroRNA expression profiles of human leukemias. Leukemia 2008; 22: 1274–1278.
Pichiorri F, Suh S-S, Ladetto M, Kuehl M, Palumbo T, Drandi D et al. MicroRNAs regulate critical genes associated with multiple myeloma pathogenesis. Proc Natl Acad Sci USA 2008; 105: 12885–12890.
Korde N, Kristinsson SY, Landgren O . Monoclonal gammopathy of undetermined significance (MGUS) and smoldering multiple myeloma (SMM): novel biological insights and development of early treatment strategies. Blood 2011; 117: 5573–5581.
Richardson PG, Schlossman RL, Alsina M et al. PANORAMA 2: panobinostat in combination with bortezomib and dexamethasone in patients with relapsed and bortezomib-refractory myeloma. Blood 2013; 122 (14): 2331–2337.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Author contributions
All authors have contributed to writing the manuscript. All authors have read and approved the final version of the manuscript.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Ahmad, N., Haider, S., Jagannathan, S. et al. MicroRNA theragnostics for the clinical management of multiple myeloma. Leukemia 28, 732–738 (2014). https://doi.org/10.1038/leu.2013.262
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2013.262
Keywords
This article is cited by
-
Knockdown of lncRNA BDNF-AS inhibited the progression of multiple myeloma by targeting the miR-125a/b-5p-BCL2 axis
Immunity & Ageing (2022)
-
Long non-coding RNA NEAT1 targeting impairs the DNA repair machinery and triggers anti-tumor activity in multiple myeloma
Leukemia (2020)
-
Expanding the repertoire of miRNAs and miRNA-offset RNAs expressed in multiple myeloma by small RNA deep sequencing
Blood Cancer Journal (2019)
-
Dysregulation of EZH2/miR-138 axis contributes to drug resistance in multiple myeloma by downregulating RBPMS
Leukemia (2018)
-
Drugging the lncRNA MALAT1 via LNA gapmeR ASO inhibits gene expression of proteasome subunits and triggers anti-multiple myeloma activity
Leukemia (2018)