Abstract
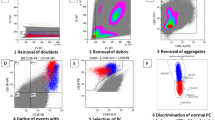
Detection of minimal residual disease (MRD) in chronic lymphocytic leukaemia (CLL) is becoming increasingly important as treatments improve. An internationally harmonised four-colour (CLR) flow cytometry MRD assay is widely used but has limitations. The aim of this study was to improve MRD analysis by identifying situations where a less time-consuming CD19/CD5/κ/λ analysis would be sufficient for detecting residual CLL, and develop a six-CLR antibody panel that is more efficient for cases requiring full MRD analysis. In 784 samples from CLL patients after treatment, it was possible to determine CD19/CD5/κ/λ thresholds that identified cases with detectable MRD with 100% positive predictive value (PPV). However, CD19/CD5/κ/λ analysis was unsuitable for predicting iwCLL/NCI response status or identifying cases with no detectable MRD. For the latter cases requiring a full MRD assessment, a six-CLR assay was designed comprising CD19/CD5/CD20 with (1) CD3/CD38/CD79b and (2) CD81/CD22/CD43. There was good correlation between four-CLR and six-CLR panels in dilution studies and clinical samples, with 100% concordance for detection of residual disease at the 0.01% (10−4) level (n=59) and good linearity even at the 0.001–0.01% (10−5–10−4) level. A six-CLR panel therefore provides equivalent results to the four-CLR panel but it requires fewer reagents, fewer cells and a much simpler analysis approach.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Gribben JG, O’Brien S . Update on therapy of chronic lymphocytic leukemia. J Clin Oncol 2011; 29: 544–550.
Abrisqueta P, Pereira A, Rozman C, Aymerich M, Giné E, Moreno C et al. Improving survival in patients with chronic lymphocytic leukemia (1980–2008): the Hospital Clinic of Barcelona experience. Blood 2009; 114: 2044–2050.
Moreton P, Kennedy B, Lucas G, Leach M, Rassam SMB, Haynes A et al. Eradication of minimal residual disease in B-cell chronic lymphocytic leukemia after alemtuzumab therapy is associated with prolonged survival. J Clin Oncol 2005; 23: 2971–2979.
Hallek M, Fischer K, Fingerle-Rowson G, Fink AM, Busch R, Mayer J et al. Addition of rituximab to fludarabine and cyclophosphamide in patients with chronic lymphocytic leukaemia: a randomised, open-label, phase 3 trial. Lancet 2010; 376: 1164–1174.
Böttcher S, Ritgen M, Fischer K, Stilgenbauer S, Busch RM, Fingerle-Rowson G et al. Minimal residual disease quantification is an independent predictor of progression-free and overall survival in chronic lymphocytic leukemia: a multivariate analysis from the randomized GCLLSG CLL8 trial. J Clin Oncol 2012; doi:10.1200/JCO.2011.36.9348.
Böttcher S, Ritgen M, Pott C, Brüggemann M, Raff T, Stilgenbauer S et al. Comparative analysis of minimal residual disease detection using four-color flow cytometry, consensus IgH-PCR, and quantitative IgH PCR in CLL after allogeneic and autologous stem cell transplantation. Leukemia 2004; 18: 1637–1645.
Moreno C, Villamor N, Colomer D, Esteve J, Giné E, Muntañola A et al. Clinical significance of minimal residual disease, as assessed by different techniques, after stem cell transplantation for chronic lymphocytic leukemia. Blood 2006; 107: 4563–4569.
Rawstron AC, Villamor N, Ritgen M, Böttcher S, Ghia P, Zehnder JL et al. International standardized approach for flow cytometric residual disease monitoring in chronic lymphocytic leukaemia. Leukemia 2007; 21: 956–964.
Lenormand B, Bizet M, Fruchart C, Tilly H, Daliphard S, Thouret F et al. Residual disease in B-cell chronic lymphocytic leukemia patients and prognostic value. Leukemia 1994; 8: 1019–1026.
Vuillier F, Claisse JF, Vandenvelde C, Travade P, Magnac C, Chevret S et al. Evaluation of residual disease in B-cell chronic lymphocytic leukemia patients in clinical and bone-marrow remission using CD5-CD19 markers and PCR study of gene rearrangements. Leuk Lymphoma 1992; 7: 195–204.
Robertson LE, Huh YO, Butler JJ, Pugh WC, Hirsch-Ginsberg C, Stass S et al. Response assessment in chronic lymphocytic leukemia after fludarabine plus prednisone: clinical, pathologic, immunophenotypic, and molecular analysis. Blood 1992; 80: 29–36.
Tam CS, O’Brien S, Wierda W, Kantarjian H, Wen S, Do K-A et al. Long-term results of the fludarabine, cyclophosphamide, and rituximab regimen as initial therapy of chronic lymphocytic leukemia. Blood 2008; 112: 975–980.
Hillmen P, Skotnicki AB, Robak T, Jaksic B, Dmoszynska A, Wu J et al. Alemtuzumab compared with chlorambucil as first-line therapy for chronic lymphocytic leukemia. J Clin Oncol 2007; 25: 5616–5623.
Hallek M, Cheson BD, Catovsky D, Caligaris-Cappio F, Dighiero G, Döhner H et al. Guidelines for the diagnosis and treatment of chronic lymphocytic leukemia: a report from the International Workshop on Chronic Lymphocytic Leukemia updating the National Cancer Institute-Working Group 1996 guidelines. Blood 2008; 111: 5446–5456.
Catovsky D, Richards S, Matutes E, Oscier D, Dyer MJS, Bezares RF et al. Assessment of fludarabine plus cyclophosphamide for patients with chronic lymphocytic leukaemia (the LRF CLL4 Trial): a randomised controlled trial. Lancet 2007; 370: 230–239.
Kennedy B, Rawstron A, Carter C, Ryan M, Speed K, Lucas G et al. Campath-1H and fludarabine in combination are highly active in refractory chronic lymphocytic leukemia. Blood 2002; 99: 2245–2247.
Böttcher S, Stilgenbauer S, Busch R, Brüggemann M, Raff T, Pott C et al. Standardized MRD flow and ASO IGH RQ-PCR for MRD quantification in CLL patients after rituximab-containing immunochemotherapy: a comparative analysis. Leukemia 2009; 23: 2007–2017.
Bosch F, Abrisqueta P, Villamor N, Terol MJ, González-Barca E, Ferra C et al. Rituximab, fludarabine, cyclophosphamide, and mitoxantrone: a new, highly active chemoimmunotherapy regimen for chronic lymphocytic leukemia. J Clin Oncol 2009; 27: 4578–4584.
Bosch F, Ferrer A, Villamor N, González M, Briones J, González-Barca E et al. Fludarabine, cyclophosphamide, and mitoxantrone as initial therapy of chronic lymphocytic leukemia: high response rate and disease eradication. Clin Cancer Res 2008; 14: 155–161.
Marti GE, Rawstron AC, Ghia P, Hillmen P, Houlston RS, Kay N et al. Diagnostic criteria for monoclonal B-cell lymphocytosis. Br J Haematol 2005; 130: 325–332.
Böttcher S, Ritgen M, Buske S, Gesk S, Klapper W, Hoster E et al. Minimal residual disease detection in mantle cell lymphoma: methods and significance of four-color flow cytometry compared to consensus IGH-polymerase chain reaction at initial staging and for follow-up examinations. Haematologica 2008; 93: 551–559.
Perez-Andres M, Paiva B, Nieto WG, Caraux A, Schmitz A, Almeida J et al. Human peripheral blood B-cell compartments: a crossroad in B-cell traffic. Cytometry B Clin Cytom 2010; 78 (Suppl 1): S47–S60.
Bomberger C, Singh-Jairam M, Rodey G, Guerriero A, Yeager AM, Fleming WH et al. Lymphoid reconstitution after autologous PBSC transplantation with FACS-sorted CD34+ hematopoietic progenitors. Blood 1998; 91: 2588–2600.
Rawstron AC, Kennedy B, Evans PA, Davies FE, Richards SJ, Haynes AP et al. Quantitation of minimal disease levels in chronic lymphocytic leukemia using a sensitive flow cytometric assay improves the prediction of outcome and can be used to optimize therapy. Blood 2001; 98: 29–35.
Le Roy C, Varin-Blank N, Ajchenbaum-Cymbalista F, Letestu R . Flow cytometry APC-tandem dyes are degraded through a cell-dependent mechanism. Cytometry A 2009; 75: 882–890.
Hillmen P, Cohen DR, Cocks K, Pettitt A, Sayala HA, Rawstron AC et al. A randomized phase II trial of fludarabine, cyclophosphamide and mitoxantrone (FCM) with or without rituximab in previously treated chronic lymphocytic leukaemia. Br J Haematol 2011; 152: 570–578.
Logan AC, Gao H, Wang C, Sahaf B, Jones CD, Marshall EL et al. High-throughput VDJ sequencing for quantification of minimal residual disease in chronic lymphocytic leukemia and immune reconstitution assessment. Proc Natl Acad Sci USA 2011; 108: 21194–21199.
Pedreira CE, Costa ES, Almeida J, Fernandez C, Quijano S, Flores J et al. A probabilistic approach for the evaluation of minimal residual disease by multiparameter flow cytometry in leukemic B-cell chronic lymphoproliferative disorders. Cytometry A 2008; 73A: 1141–1150.
Acknowledgements
We are grateful to BD Biosciences, particularly Frans Nauwelaers and Lucia Testolin, for provision of antibodies used in this study. PG supported by Cariplo Foundation (Milan, Italy), Program Molecular Clinical Oncology-5 per mille number 9965 and Investigator Grant, Associazione Italiana per la Ricerca sul Cancro (AIRC Milano, Italy), Progetti di Ricerca di Interesse Nazionale (PRIN - Ministry of education, University and Research - MIUR, Rome, Italy) and Ricerca Finalizzata 2010 (Ministry of Health, Rome, Italy); AR supported by LLR (www.leukaemialymphomaresearch.org.uk). We are grateful to Linda Falck, Elke Harbst, Jamileh Hanani, Lada Henseleit, Heike Hinrichs and Abdelmalek Dahmani for excellent technical support.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
The antibodies used for MRD analysis in this study were provided by BD Biosciences. AR has received royalties from BD for an unrelated product (IntraSure). Other companies were contacted at the time of publication of the international standardised approach8 but did not express an interest in providing antibodies for development of MRD detection. BD Biosciences had no scientific input into the project and there were no significant conflicts of interest that could be perceived to bias the work. The remaining authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Rawstron, A., Böttcher, S., Letestu, R. et al. Improving efficiency and sensitivity: European Research Initiative in CLL (ERIC) update on the international harmonised approach for flow cytometric residual disease monitoring in CLL. Leukemia 27, 142–149 (2013). https://doi.org/10.1038/leu.2012.216
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2012.216
Keywords
This article is cited by
-
Measurable Residual Disease in Chronic Lymphocytic Leukemia: Current Understanding and Evolving Role in Clinical Practice
Current Treatment Options in Oncology (2023)
-
Bcl2 inhibitor venetoclax +/− Anti-CD20: what do deep remissions mean?
memo - Magazine of European Medical Oncology (2022)
-
Identification of CD105 (endoglin) as novel risk marker in CLL
Annals of Hematology (2022)
-
Measurable residual disease in chronic lymphocytic leukemia: expert review and consensus recommendations
Leukemia (2021)
-
Prognostic value of high-sensitivity measurable residual disease assessment after front-line chemoimmunotherapy in chronic lymphocytic leukemia
Leukemia (2021)