Abstract
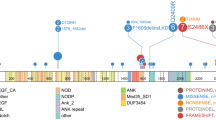
Acute lymphoblastic leukemia (ALL), the most common malignant disorder in childhood, is typically associated with numerical chromosomal aberrations, fusion genes or small focal deletions, thought to represent important pathogenetic events in the development of the leukemia. Mutations, such as single nucleotide changes, have also been reported in childhood ALL, but these have only been studied by sequencing a small number of candidate genes. Herein, we report the first unbiased sequencing of the whole exome of two cases of pediatric ALL carrying the ETV6/RUNX1 (TEL/AML1) fusion gene (the most common genetic subtype) and corresponding normal samples. A total of 14 somatic mutations were identified, including four and seven protein-altering nucleotide substitutions in each ALL. Twelve mutations (86%) occurred in genes previously described to be mutated in other types of cancer, but none was found to be recurrent in an extended series of 29 ETV6/RUNX1-positive ALLs. The number of single nucleotide mutations was similar to the number of copy number alterations as detected by single nucleotide polymorphism arrays. Although the true pathogenetic significance of the mutations must await future functional evaluations, this study provides a first estimate of the mutational burden at the genetic level of t(12;21)-positive childhood ALL.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Hanahan D, Weinberg RA . The hallmarks of cancer. Cell 2000; 100: 57–70.
Forbes SA, Bhamra G, Bamford S, Dawson E, Kok C, Clements J et al. The catalogue of somatic mutations in cancer (COSMIC). Curr Protoc Hum Genet 2008; 57: 10.11.1–10.11.26.
Mitelman F, Johansson B, Mertens F . Mitelman Database of Chromosome Aberrations in Cancer 2010. http://cgap.nci.nih.gov/Chromosomes/Mitelman.
Ley TJ, Mardis ER, Ding L, Fulton B, McLellan MD, Chen K et al. DNA sequencing of a cytogenetically normal acute myeloid leukemia genome. Nature 2008; 456: 66–72.
Mardis ER, Ding L, Dooling DJ, Larson DE, McLellan MD, Chen K et al. Recurring mutations found by sequencing an acute myeloid leukemia genome. N Engl J Med 2009; 361: 1058–1066.
Shah SP, Morin RD, Khattra J, Prentice L, Pugh T, Burleigh A et al. Mutational evolution in a lobular breast tumour profiled at single nucleotide resolution. Nature 2009; 461: 809–813.
Pleasance ED, Stephens PJ, O'Meara S, McBride DJ, Meynert A, Jones D et al. A small-cell lung cancer genome with complex signatures of tobacco exposure. Nature 2010; 463: 184–190.
Pleasance ED, Cheetham RK, Stephens PJ, McBride DJ, Humphray SJ, Greenman CD et al. A comprehensive catalogue of somatic mutations from a human cancer genome. Nature 2010; 463: 191–196.
Ng SB, Buckingham KJ, Lee C, Bigham AW, Tabor HK, Dent KM et al. Exome sequencing identifies the cause of a mendelian disorder. Nat Genet 2010; 42: 30–35.
Choi M, Scholl UI, Ji W, Liu T, Tikhonova IR, Zumbo P et al. Genetic diagnosis by whole exome capture and massively parallel DNA sequencing. Proc Natl Acad Sci USA 2009; 106: 19096–19101.
Gilissen C, Arts HH, Hoischen A, Spruijt L, Mans DA, Arts P et al. Exome sequencing identifies WDR35 variants involved in sensenbrenner syndrome. Am J Hum Genet 2010; 87: 418–423.
Greif PA, Yaghmaie M, Konstandin NP, Ksienzyk B, Alimoghaddam K, Ghavamzadeh A et al. Somatic mutations in acute promyelocytic leukemia (APL) identified by exome sequencing. Leukemia 2011; 25: 1519–1522.
Li H, Durbin R . Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009; 25: 1754–1760.
Li H, Ruan J, Durbin R . Mapping short DNA sequencing reads and calling variants using mapping quality scores. Genome Res 2008; 18: 1851–1858.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009; 25: 2078–2079.
Crowe M . SeqDoC: rapid SNP and mutation detection by direct comparison of DNA sequence chromatograms. BMC Bioinformatics 2005; 6: 133.
Hedges DJ, Hedges D, Burges D, Powell E, Almonte C, Huang J et al. Exome sequencing of a multigenerational human pedigree. PLoS ONE 2009; 4: e8232.
Sherry ST, Ward MH, Kholodov M, Baker J, Phan L, Smigielski EM et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res 2001; 29: 308–311.
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D et al. Circos: an information aesthetic for comparative genomics. Genome Res 2009; 19: 1639–1645.
Lilljebjörn H, Soneson C, Andersson A, Heldrup J, Behrendtz M, Kawamata N et al. The correlation pattern of acquired copy number changes in 164 ETV6/RUNX1-positive childhood acute lymphoblastic leukemias. Hum Mol Genet 2010; 19: 3150–3158.
Stratton MR, Campbell PJ, Futreal PA . The cancer genome. Nature 2009; 458: 719–724.
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P et al. A method and server for predicting damaging missense mutations. Nat Methods 2010; 7: 248–249.
Ng PC, Henikoff S . SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res 2003; 31: 3812–3814.
Stirewalt DL, Radich JP . The role of FLT3 in haematopoietic malignancies. Nat Rev Cancer 2003; 3: 650–665.
Ouhtit A, Gaur RL, Abd Elmageed ZY, Fernando A, Thouta R, Trappey AK et al. Towards understanding the mode of action of the multifaceted cell adhesion receptor CD146. Biochim Biophys Acta 2009; 1795: 130–136.
Ma C, Quesnelle KM, Sparano A, Rao S, Park MS, Cohen MA et al. Characterization CSMD1 in a large set of primary lung, head and neck, breast and skin cancer tissues. Cancer Biol Ther 2009; 8: 907–916.
Cano CE, Gommeaux J, Pietri S, Culcasi M, Garcia S, Seux M et al. Tumor protein 53–induced nuclear protein 1 is a major mediator of p53 antioxidant function. Cancer Res 2009; 69: 219–226.
Acknowledgements
This work was supported by grants from the Swedish Cancer Society, Swedish Childhood Cancer Foundation, Swedish Research Council (personal project grant to TF; Hemato-Linne and BioCARE strategic research program grants), Inga-Britt and Arne Lundberg Foundation, Gunnar Nilsson Cancer Foundation, and the Medical Faculty of Lund University.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Leukemia website
Rights and permissions
About this article
Cite this article
Lilljebjörn, H., Rissler, M., Lassen, C. et al. Whole-exome sequencing of pediatric acute lymphoblastic leukemia. Leukemia 26, 1602–1607 (2012). https://doi.org/10.1038/leu.2011.333
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2011.333
Keywords
This article is cited by
-
The evolution of relapse of adult T cell acute lymphoblastic leukemia
Genome Biology (2020)
-
ETV6/RUNX1-positive childhood acute lymphoblastic leukemia in China: excellent prognosis with improved BFM protocol
Italian Journal of Pediatrics (2018)
-
Cytogenetic and Molecular Predictors of Outcome in Acute Lymphocytic Leukemia: Recent Developments
Current Hematologic Malignancy Reports (2012)