Abstract
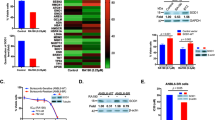
Bortezomib is an effective agent for treating multiple myeloma (MM). To investigate the underlying mechanisms associated with acquired resistance to this agent, we established two bortezomib-resistant MM cell lines, KMS-11/BTZ and OPM-2/BTZ, the 50% inhibitory concentration values of which were respectively 24.7- and 16.6-fold higher than their parental cell lines. No activation of caspase and BH3-only proteins such as Noxa was noted in bortezomib-resistant cells after exposure to the drug. The accumulation of polyubiquitinated proteins was reduced in bortezomib-resistant cells compared with the parental cells, associated with avoidance of catastrophic ER stress as assessed by downregulation of CHOP expression. These resistant MM cells have a unique point mutation, G322A, in the gene encoding the proteasome β5 subunit (PSMB5), likely resulting in conformational changes to the bortezomib-binding pocket of this subunit. KMS-11 parental cells transfected to express mutated PSMB5 also showed reduced bortezomib-induced apoptosis compared with those expressing wild-type PSMB5 or the parental cells. Expression of mutated PSMB5 was associated with the prevention of the accumulation of unfolded proteins. Thus, a fraction of MM cells may acquire bortezomib resistance by suppressing apoptotic signals through the inhibition of unfolded protein accumulation and subsequent excessive ER stress by a mutation of the PSMB5 gene.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Richardson PG, Sonneveld P, Schuster MW, Irwin D, Stadtmauer EA, Facon T et al. Bortezomib or high-dose dexamethasone for relapsed multiple myeloma. N Engl J Med 2005; 352: 2487–2498.
San Miguel JF, Schlag R, Khuageva NK, Dimopoulos MA, Shpilberg O, Kropff M et al. Bortezomib plus melphalan and prednisone for initial treatment of multiple myeloma. N Engl J Med 2008; 359: 906–917.
Kumar S, Rajkumar SV . Many facets of bortezomib resistance/susceptibility. Blood 2008; 112: 2177–2178.
Shah JJ, Orlowski RZ . Proteasome inhibitors in the treatment of multiple myeloma. Leukemia 2009; 23: 1964–1979.
Karin M, Greten FR . NF-kappaB: linking inflammation and immunity to cancer development and progression. Nat Rev Immunol 2005; 5: 749–759.
Fennell DA, Chacko A, Mutti L . BCL-2 family regulation by the 20S proteasome inhibitor bortezomib. Oncogene 2008; 27: 1189–1197.
Perez-Galan P, Roue G, Villamor N, Montserrat E, Campo E, Colomer D . The proteasome inhibitor bortezomib induces apoptosis in mantle-cell lymphoma through generation of ROS and Noxa activation independent of p53 status. Blood 2006; 107: 257–264.
Ri M, Iida S, Ishida T, Ito A, Yano H, Inagaki A et al. Bortezomib-induced apoptosis in mature T-cell lymphoma cells partially depends on upregulation of Noxa and functional repression of Mcl-1. Cancer Sci 2009; 100: 341–348.
Zhu H, Zhang L, Dong F, Guo W, Wu S, Teraishi F et al. Bik/NBK accumulation correlates with apoptosis-induction by bortezomib (PS-341, Velcade) and other proteasome inhibitors. Oncogene 2005; 24: 4993–4999.
Nawrocki ST, Carew JS, Pino MS, Highshaw RA, Dunner Jr K, Huang P et al. Bortezomib sensitizes pancreatic cancer cells to endoplasmic reticulum stress-mediated apoptosis. Cancer Res 2005; 65: 11658–11666.
Obeng EA, Carlson LM, Gutman DM, Harrington Jr WJ, Lee KP, Boise LH . Proteasome inhibitors induce a terminal unfolded protein response in multiple myeloma cells. Blood 2006; 107: 4907–4916.
Adams J . The development of proteasome inhibitors as anticancer drugs. Cancer Cell 2004; 5: 417–421.
Schroder M, Kaufman RJ . The mammalian unfolded protein response. Annu Rev Biochem 2005; 74: 739–789.
Yoshida H . ER stress and diseases. FEBS J 2007; 274: 630–658.
Moenner M, Pluquet O, Bouchecareilh M, Chevet E . Integrated endoplasmic reticulum stress responses in cancer. Cancer Res 2007; 67: 10631–10634.
Lu S, Yang J, Song X, Gong S, Zhou H, Guo L et al. Point mutation of the proteasome beta5 subunit gene is an important mechanism of bortezomib resistance in bortezomib-selected variants of Jurkat T cell lymphoblastic lymphoma/leukemia line. J Pharmacol Exp Ther 2008; 326: 423–431.
Oerlemans R, Franke NE, Assaraf YG, Cloos J, van Zantwijk I, Berkers CR et al. Molecular basis of bortezomib resistance: proteasome subunit beta5 (PSMB5) gene mutation and overexpression of PSMB5 protein. Blood 2008; 112: 2489–2499.
Ruckrich T, Kraus M, Gogel J, Beck A, Ovaa H, Verdoes M et al. Characterization of the ubiquitin-proteasome system in bortezomib-adapted cells. Leukemia 2009; 23: 1098–1105.
Wang Q, Mora-Jensen H, Weniger MA, Perez-Galan P, Wolford C, Hai T et al. ERAD inhibitors integrate ER stress with an epigenetic mechanism to activate BH3-only protein NOXA in cancer cells. Proc Natl Acad Sci USA 2009; 106: 2200–2205.
Hideshima T, Ikeda H, Chauhan D, Okawa Y, Raje N, Podar K et al. Bortezomib induces canonical nuclear factor-kappaB activation in multiple myeloma cells. Blood 2009; 114: 1046–1052.
Groll M, Berkers CR, Ploegh HL, Ovaa H . Crystal structure of the boronic acid-based proteasome inhibitor bortezomib in complex with the yeast 20S proteasome. Structure 2006; 14: 451–456.
Acknowledgements
We thank Ms Chiori Fukuyama for her skillful technical assistance. This work was supported by Grants-in-Aid for Scientific Research on Priority Areas (No. 17016065 & 16062101 for RU) from the Ministry of Education, Culture, Science, Sports and Technology, Japan; and Grants-in-Aid for Cancer Research from the Ministry of Health, Labor and Welfare, Japan (No. 17S-1, 17-16 anf 21-8-5 for SI). This research was also funded in part by Kyowa Hakko Kirin Co., Ltd, Tokyo, Japan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
TN, HM and YS are employees of Kyowa Hakko Kirin Co., Ltd., Japan. SI received research funding from Kyowa Hakko Kirin. SI declares honoraria from Janssen Pharmaceutical K.K., Dainippon Sumitomo Pharmaceutical Co., Ltd, Chugai Pharmaceutical Co., Ltd and Novartis Pharma K.K.
Additional information
Supplementary Information accompanies the paper on the Leukemia website
Rights and permissions
About this article
Cite this article
Ri, M., Iida, S., Nakashima, T. et al. Bortezomib-resistant myeloma cell lines: a role for mutated PSMB5 in preventing the accumulation of unfolded proteins and fatal ER stress. Leukemia 24, 1506–1512 (2010). https://doi.org/10.1038/leu.2010.137
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2010.137
Keywords
This article is cited by
-
Acute suppression of translation by hyperthermia enhances anti-myeloma activity of carfilzomib
International Journal of Hematology (2024)
-
Negligible role of TRAIL death receptors in cell death upon endoplasmic reticulum stress in B-cell malignancies
Oncogenesis (2023)
-
A clinically relevant pulse treatment generates a bortezomib-resistant myeloma cell line that lacks proteasome mutations and is sensitive to Bcl-2 inhibitor venetoclax
Scientific Reports (2022)
-
The combination of the tubulin binding small molecule PTC596 and proteasome inhibitors suppresses the growth of myeloma cells
Scientific Reports (2021)
-
LncRNA H19 overexpression induces bortezomib resistance in multiple myeloma by targeting MCL-1 via miR-29b-3p
Cell Death & Disease (2019)