Abstract
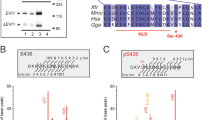
The ecotropic viral integration site-1 (EVI-1) is a nuclear transcription factor and has an essential function in the proliferation/maintenance of haematopoietic stem cells. Aberrant expression of EVI-1 has been frequently found in myeloid leukaemia as well as in several solid tumours, and is associated with a poor patient survival. It was recently shown that EVI-1 associates with two different histone methyltransferases (HMTs), SUV39H1 and G9a. However, the functional roles of these HMTs in EVI-1-mediated leukemogenesis remain unclear. In this study, we showed that EVI-1 physically interacts with SUV39H1 and G9a, but not with Set9. Immunofluorescence analysis revealed that EVI-1 colocalizes with these HMTs in nuclei. We also found that the catalytically inactive form of SUV39H1 abrogates the transcriptional repression mediated by EVI-1, suggesting that SUV39H1 is actively involved in EVI-1-mediated transcriptional repression. Furthermore, RNAi-based knockdown of SUV39H1 or G9a in Evi-1-expressing progenitors significantly reduced their colony-forming activity. In contrast, knockdown of these HMTs did not impair bone marrow immortalization by E2A/HLF. These results indicate that EVI-1 forms higher-order complexes with HMTs, and this association has a role in the transcription repression and bone marrow immortalization. Targeting these HMTs may be of therapeutic benefit in the treatment for EVI-1-related haematological malignancies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Morishita K, Parker DS, Mucenski ML, Jenkins NA, Copeland NG, Ihle JN . Retroviral activation of a novel gene encoding a zinc finger protein in IL-3-dependent myeloid leukemia cell lines. Cell 1988; 54: 831–840.
Mucenski ML, Taylor BA, Ihle JN, Hartley JW, Morse III HC, Jenkins NA et al. Identification of a common ecotropic viral integration site, Evi-1, in the DNA of AKXD murine myeloid tumors. Mol Cell Biol 1988; 8: 301–308.
Valk PJ, Verhaak RG, Beijen MA, Erpelinck CA, Barjesteh van Waalwijk van Doorn-Khosrovani S, Boer JM et al. Prognostically useful gene-expression profiles in acute myeloid leukemia. N Engl J Med 2004; 350: 1617–1628.
Lugthart S, van Drunen E, van Norden Y, van Hoven A, Erpelinck CA, Valk PJ et al. High EVI1 levels predict adverse outcome in acute myeloid leukemia: prevalence of EVI1 overexpression and chromosome 3q26 abnormalities underestimated. Blood 2008; 111: 4329–4337.
Kurokawa M, Mitani K, Irie K, Matsuyama T, Takahashi T, Chiba S et al. The oncoprotein Evi-1 represses TGF-beta signalling by inhibiting Smad3. Nature 1998; 394: 92–96.
Kurokawa M, Mitani K, Yamagata T, Takahashi T, Izutsu K, Ogawa S et al. The evi-1 oncoprotein inhibits c-Jun N-terminal kinase and prevents stress-induced cell death. EMBO J 2000; 19: 2958–2968.
Tanaka T, Nishida J, Mitani K, Ogawa S, Yazaki Y, Hirai H . Evi-1 raises AP-1 activity and stimulates c-fos promoter transactivation with dependence on the second zinc finger domain. J Biol Chem 1994; 269: 24020–24026.
Buonamici S, Li D, Chi Y, Zhao R, Wang X, Brace L et al. EVI1 induces myelodysplastic syndrome in mice. J Clin Invest 2004; 114: 713–719.
Jin G, Yamazaki Y, Takuwa M, Takahara T, Kaneko K, Kuwata T et al. Trib1 and Evi1 cooperate with Hoxa and Meis1 in myeloid leukemogenesis. Blood 2007; 109: 3998–4005.
Watanabe-Okochi N, Kitaura J, Ono R, Harada H, Harada Y, Komeno Y et al. AML1 mutations induced MDS and MDS/AML in a mouse BMT model. Blood 2008; 111: 4297–4308.
Yuasa H, Oike Y, Iwama A, Nishikata I, Sugiyama D, Perkins A et al. Oncogenic transcription factor Evi1 regulates hematopoietic stem cell proliferation through GATA-2 expression. EMBO J 2005; 24: 1976–1987.
Goyama S, Yamamoto G, Shimabe M, Sato T, Ichikawa M, Ogawa S et al. Evi-1 is a critical regulator for hematopoietic stem cells and transformed leukemic cells. Cell Stem Cell 2008; 3: 207–220.
Fears S, Mathieu C, Zeleznik-Le N, Huang S, Rowley JD, Nucifora G . Intergenic splicing of MDS1 and EVI1 occurs in normal tissues as well as in myeloid leukemia and produces a new member of the PR domain family. Proc Natl Acad Sci USA 1996; 93: 1642–1647.
Nitta E, Izutsu K, Yamaguchi Y, Imai Y, Ogawa S, Chiba S et al. Oligomerization of Evi-1 regulated by the PR domain contributes to recruitment of corepressor CtBP. Oncogene 2005; 24: 6165–6173.
Sood R, Talwar-Trikha A, Chakrabarti SR, Nucifora G . MDS1/EVI1 enhances TGF-beta1 signaling and strengthens its growth-inhibitory effect but the leukemia-associated fusion protein AML1/MDS1/EVI1, product of the t(3;21), abrogates growth-inhibition in response to TGF-beta1. Leukemia 1999; 13: 348–357.
Izutsu K, Kurokawa M, Imai Y, Maki K, Mitani K, Hirai H . The corepressor CtBP interacts with Evi-1 to repress transforming growth factor beta signaling. Blood 2001; 97: 2815–2822.
Palmer S, Brouillet JP, Kilbey A, Fulton R, Walker M, Crossley M et al. Evi-1 transforming and repressor activities are mediated by CtBP co-repressor proteins. J Biol Chem 2001; 276: 25834–25840.
Vinatzer U, Taplick J, Seiser C, Fonatsch C, Wieser R . The leukaemia-associated transcription factors EVI-1 and MDS1/EVI1 repress transcription and interact with histone deacetylase. Br J Haematol 2001; 114: 566–573.
Rea S, Eisenhaber F, O'Carroll D, Strahl BD, Sun ZW, Schmid M et al. Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature 2000; 406: 593–599.
Tachibana M, Sugimoto K, Fukushima T, Shinkai Y . Set domain-containing protein, G9a, is a novel lysine-preferring mammalian histone methyltransferase with hyperactivity and specific selectivity to lysines 9 and 27 of histone H3. J Biol Chem 2001; 276: 25309–25317.
Cattaneo F, Nucifora G . EVI1 recruits the histone methyltransferase SUV39H1 for transcription repression. J Cell Biochem 2008; 105: 344–352.
Spensberger D, Delwel R . A novel interaction between the proto-oncogene Evi1 and histone methyltransferases, SUV39H1 and G9a. FEBS Lett 2008; 582: 2761–2767.
Kitamura T, Koshino Y, Shibata F, Oki T, Nakajima H, Nosaka T et al. Retrovirus-mediated gene transfer and expression cloning: powerful tools in functional genomics. Exp Hematol 2003; 31: 1007–1014.
Takeshita M, Ichikawa M, Nitta E, Goyama S, Asai T, Ogawa S et al. AML1-Evi-1 specifically transforms hematopoietic stem cells through fusion of the entire Evi-1 sequence to AML1. Leukemia 2008; 22: 1241–1249.
Goyama S, Kurokawa M . Pathogenetic significance of ecotropic viral integration site-1 in hematological malignancies. Cancer Sci 2009; 100: 990–995.
Sato T, Goyama S, Nitta E, Takeshita M, Yoshimi M, Nakagawa M et al. Evi-1 promotes para-aortic splanchnopleural hematopoiesis through up-regulation of GATA-2 and repression of TGF-b signaling. Cancer Sci 2008; 99: 1407–1413.
Chakraborty S, Senyuk V, Sitailo S, Chi Y, Nucifora G . Interaction of EVI1 with cAMP-responsive element-binding protein-binding protein (CBP) and p300/CBP-associated factor (P/CAF) results in reversible acetylation of EVI1 and in co-localization in nuclear speckles. J Biol Chem 2001; 276: 44936–44943.
Peters AH, Kubicek S, Mechtler K, O'Sullivan RJ, Derijck AA, Perez-Burgos L et al. Partitioning and plasticity of repressive histone methylation states in mammalian chromatin. Mol Cell 2003; 12: 1577–1589.
Laricchia-Robbio L, Nucifora G . Significant increase of self-renewal in hematopoietic cells after forced expression of EVI1. Blood Cells Mol Dis 2008; 40: 141–147.
Smith KS, Rhee JW, Cleary ML . Transformation of bone marrow B-cell progenitors by E2a-Hlf requires coexpression of Bcl-2. Mol Cell Biol 2002; 22: 7678–7687.
Gyory I, Wu J, Fejer G, Seto E, Wright KL . PRDI-BF1 recruits the histone H3 methyltransferase G9a in transcriptional silencing. Nat Immunol 2004; 5: 299–308.
Greiner D, Bonaldi T, Eskeland R, Roemer E, Imhof A . Identification of a specific inhibitor of the histone methyltransferase SU(VAR)3-9. Nat Chem Biol 2005; 1: 143–145.
Kubicek S, O'Sullivan RJ, August EM, Hickey ER, Zhang Q, Teodoro ML et al. Reversal of H3K9me2 by a small-molecule inhibitor for the G9a histone methyltransferase. Mol Cell 2007; 25: 473–481.
Acknowledgements
We thank T Kitamura for Plat-E packaging cells, H Nakauchi and M Onodera for pGCDNsam-eGFP retroviral vector, T Inaba, M Tachibana, Y Shinkai, K Wright and D Reinberg for cDNAs (see Materials and methods), Y Shimamura for expert technical assistance, and KYOWA KIRIN for cytokines. This work was supported in part by a Grant-in-Aid for Scientific Research from the Japan Society for the Promotion of Science, and by Health and Labour Sciences Research grants from the Ministry of Health, Labour and Welfare.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Leukemia website (http://www.nature.com/leu)
Rights and permissions
About this article
Cite this article
Goyama, S., Nitta, E., Yoshino, T. et al. EVI-1 interacts with histone methyltransferases SUV39H1 and G9a for transcriptional repression and bone marrow immortalization. Leukemia 24, 81–88 (2010). https://doi.org/10.1038/leu.2009.202
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2009.202
Keywords
This article is cited by
-
SUV39H1 regulates the progression of MLL-AF9-induced acute myeloid leukemia
Oncogene (2020)
-
Aging increases vulnerability to stress-induced depression via upregulation of NADPH oxidase in mice
Communications Biology (2020)
-
Quinazolines as inhibitors of chromatin-associated proteins in histones
Medicinal Chemistry Research (2019)
-
The SUV39H1 inhibitor chaetocin induces differentiation and shows synergistic cytotoxicity with other epigenetic drugs in acute myeloid leukemia cells
Blood Cancer Journal (2015)
-
Evi1 defines leukemia-initiating capacity and tyrosine kinase inhibitor resistance in chronic myeloid leukemia
Oncogene (2014)