Abstract
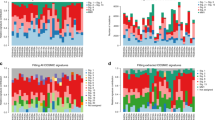
Bone disease in myeloma occurs as a result of complex interactions between myeloma cells and the bone marrow microenvironment. A custom-built DNA single nucleotide polymorphism (SNP) chip containing 3404 SNPs was used to test genomic DNA from myeloma patients classified by the extent of bone disease. Correlations identified with a Total Therapy 2 (TT2) (Arkansas) data set were validated with Eastern Cooperative Oncology Group (ECOG) and Southwest Oncology Group (SWOG) data sets. Univariate correlates with bone disease included: EPHX1, IGF1R, IL-4 and Gsk3β. SNP signatures were linked to the number of bone lesions, log2 DKK-1 myeloma cell expression levels and patient survival. Using stepwise multivariate regression analysis, the following SNPs: EPHX1 (P=0.0026); log2 DKK-1 expression (P=0.0046); serum lactic dehydrogenase (LDH) (P=0.0074); Gsk3β (P=0.02) and TNFSF8 (P=0.04) were linked to bone disease. This assessment of genetic polymorphisms identifies SNPs with both potential biological relevance and utility in prognostic models of myeloma bone disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hideshima T, Mitsiades C, Tonon G, Richardson PG, Anderson KC . Understanding multiple myeloma pathogenesis in the bone marrow to identify new therapeutic targets. Nat Rev Cancer 2007; 7: 585–598.
Giuliani N, Rizzoli V, Roodman GD . Multiple myeloma bone disease: pathophysiology of osteoblast inhibition. Blood 2006; 108: 3992–3996.
Durie BGM, Salmon SE, Mundy GR . Relation of osteoclast activating factor production to extent of bone disease in multiple myeloma. Br J Hematol 1981; 47: 21–30.
Harada S, Rodan G . Control of osteoblast function and regulation of bone mass. Nature 2003; 423: 349–355.
Westendorf JJ, Kahler RA, Schroeder TM . Wnt signaling in osteoblasts and bone diseases. Gene 2004; 341: 19–39.
Tian E, Zhan F, Walker R, Rasmussen E, Ma Y, Barlogie B et al. The role of the Wnt-signaling antagonist DKK1 in the development of osteolytic lesions in multiple myeloma. N Engl J Med 2003; 349: 2483–2494.
Yaccoby S, Ling W, Zhan F, Walker R, Barlogie B, Shaughnessy Jr JD . Antibody-based inhibition of DKK1 suppresses tumor-induced bone resorption and multiple myeloma growth in vivo. Blood 2007; 109: 2106–2111.
Colla S, Zhan F, Xiong W, Wu X, Xu H, Stephens O et al. The oxidative stress response regulates DKK1 expression through the JNK signaling cascade in multiple myeloma plasma cells. Blood 2007; 109: 4470–4477.
Qian J, Xie J, Hong S, Yang J, Zhang L, Han X et al. Dickkopf-1 (DKK-1) is a widely expressed and potent tumor-associated antigen in multiple myeloma. Blood 2007; 110: 1587–1594.
Choi SJ, Oba Y, Gazitt Y, Alsina M, Cruz J, Anderson J et al. Antisense inhibition of macrophage inflammatory protein 1-alpha blocks bone destruction in a model of myeloma bone disease. J Clin Invest 2001; 108: 1833–1841.
Lentzsch S, Gries M, Janz M, Bargou R, Dorken B, Mapara MY . Macrophage inflammatory protein 1-alpha (MIP-1 alpha) triggers migration and signaling cascades mediating survival and proliferation in multiple myeloma (MM) cells. Blood 2003; 101: 3568–3573.
Vallet S, Raje N, Ishitsuka K, Hideshima T, Podar K, Chhetri S et al. MLN3897, a novel CCR1 inhibitor, impairs osteoclastogenesis and inhibits the interaction of multiple myeloma cells and osteoclasts. Blood 2007; 110: 3744–3752.
Zhan F, Huang Y, Colla S, Stewart JP, Hanamura I, Gupta S et al. The molecular classification of multiple myeloma. Blood 2006; 108: 2020–2028.
Walker R, Barlogie B, Haessler J, Tricot G, Anaissie E, Shaughnessy Jr JD et al. Magnetic resonance imaging in multiple myeloma: diagnostic and clinical implications. J Clin Oncol 2007; 25: 1121–1128.
Johnson DC, Corthals S, Ramos C, Hoering A, Cocks K, Dickens NJ et al. Genetic associations with thalidomide mediated venous thrombotic events in myeloma identified using targeted genotyping. Blood 2008; 112: 4924–4934.
Van Ness B, Ramos C, Haznadar M, Hoering A, Haessler J, Crowley J et al. Genomic variation in myeloma: design, content, and initial application of the bank on a cure SNP panel to analysis of survival. BMC Med 2008; 6: 26 [pages not specified].
Terry MT, Elizabeth J, Atkinson R . An introduction to recursive partitioning using the rpart routines. 1997.Technical report 61, Mayo Clinic. 2Available at http://mayoresearch.mayo.edu/mayo/research/biostat/techreports.cfm##R package available at http://cran.r-project.org/src/contrib/Descriptions/rpart.html.
Agresti A . An introduction to categorical data analysis. Wiley: NJ, USA, 1996.
Breiman L . Random forests. Machine Learning 2001; 45: 5–32.
Efron B, Tibshirani R . An introduction to the Bootstrap. Chapman & Hall/CRC: FL, USA, 1994.
Kaplan EL, Meier P . Nonparametric estimation from incomplete observations. J Am Stat Assoc 1958; 53: 457–481.
Hirschhorn JN, Lohmueller K, Byrne E, Hurschhorn K . A comprehensive review of genetic association studies. Genet Med 2002; 4: 45–61.
Moghaddam MF, Grant DF, Cheek JM, Green JF, Williamson KC, Hammock BD . Bioactivation of leukotoxins to their toxic diols by epoxide hydrolase. Nature Med 1997; 3: 562–566.
Burchiel SW, Thompson TA, Lauer FT, Oprea TI . Activation of dioxin response element (DRE)-associated genes by benzo (A) pyrene3,6-quinone and benzo (A) pyrene1,6-quinone in MCF-10A human mammary epithelial cells. Toxicol Appl Pharmacol 2007; 221: 203–214.
Shi C-S, Huang N-N, Harrison K, Han S-B, Kehrl JH . The mitogen-activated protein kinase kinase kinase kinase GCKR positively regulates canonical and noncanonical Wnt signaling in B lymphocytes. Mol Cell Biol 2006; 26: 6511–6521.
Robinson JA, Chatterjee-Kishore M, Yaworsky PJ, Cullen DM, ZhaoW LC et al. Wnt/β signaling is a normal physiological response to mechanical loading in bone. J Biol Chem 2006; 281: 31720–31728.
Shi C-S, Tuscano JM, Witte ON, Kehrl JH . GCKR links the Bcr-Abl oncogene and Ras to the stress-activated protein kinase pathway. Blood 1999; 93: 1338–1345.
Knobloch J, Reimann K, Klotz L-O, Ruther U . Thalidomide resistance is based upon the capacity of the glutathione-dependent antioxidant defense. Mol Pharmaceutics 2008; 5: 1138–1144.
Edwards CM, Edwards JR, Lwin ST, Mundy GR . Target Wnt signaling in myeloma in vivo; differential effects on tumor burden and myeloma bone disease. Blood 2008; 111: 2833–2842.
Caspi M, Zilberberg A, Eldar-Finkelman H, Rosin-Arbesfeld R . Nuclear GSK-3β inhibits the canonical Wnt signalling pathway in a β-catenin phosphorylation-independent manner. Oncogene 2008; 27: 3546–3555.
Staal FJT, Luis TC, Tiemessen MM . WNT signalling in the immune system: WNT is spreading its wings. Immunology 2008; 8: 581–593.
Qiang Y-W, Chen Y, Stephens O, Brown N, Chen B, Epstein J et al. Myeloma-derived Dixkkopf-1 disrupts Wnt-regulated osteoprotegerin and RANKL production by osteoblasts: a potential mechanism underlying osteolytic bone lesions in multiple myeloma. Blood 2008; 112: 196–207.
Qiang Y-W, Shaughnessy JD, Yaccoby S . Wnt3a signaling within bone inhibits multiple myeloma bone disease and tumor growth. Blood 2008; 112: 374–382.
Edward CM . Wnt signaling: bone's defense against myeloma. Blood 2008; 112: 216–218.
Ferlin M, Noraz N, Hertogh C, Brochier J, Taylor N, Klein B . Insulin-like growth factor induces the survival and proliferation of myeloma cells through an interleukin-6-independent transduction pathway. Br J Hematology 2000; 111: 626–634.
Ge N-L, Rudikoff S . Insulin-like growth factor 1 is a dual effector of multiple myeloma cell growth. Blood 2000; 96: 2856–2861.
Mitsiades CS, Mitsiades N, Poulaki V, Schlossman R, Akivama M, Chauhan D et al. Activation of NF-kB and upregulation of intracellular anti-apoptotic proteins via the IGF-1/Akt signaling in human multiple myeloma calls: therapeutic implications. Oncogene 2002; 21: 5673–5683.
Podar K, Tai Y-T, Cole CE, Hideshima T, Sattler M, Hamblin A et al. Essential role of caveolae in interleukin-6 and insulin-like growth factor I-triggered Akt-1-mediated survival of multiple myeloma cells. J Biol Chem 2003; 278: 5794–5801.
DiGirolamo DJ, Mukherjee A, Fulzele K, Gan Y, Cao X, Frank SJ et al. Mode of growth hormone action in osteoblasts. J Biol Chem 2007; 282: 31666–31674.
Brechter AB . Kinins-important regulators in inflammation induced bone resorption. Umea Univ Odontological Dissertation. Department of Oral Cell Biology, Umea University: Umea, Sweden, 2006 (ISBN 91-7264-195-9).
Fujisawa T, Ikegami H, Kawaguchi Y, Ogihata T . Meta-analysis of the association of Trp Arg polymorphism of β3-adrenergic receptor gene with body mass index. J Clin Endocrinol Metab 1998; 83: 2441–2444.
Wang CY, Nguyen ND, Morrison NA, Eisman JA, Center JR, Nguyen TV . β3-adrenergic receptor gene, body mass index, bone mineral density and fracture risk in elderly men and women: the dubbo osteoporosis epidemiology study (DOES). BMC Med Genet 2006; 7: 57.
Riancho JA, Zarrabeitia MT, Olmos JM, Amado JA, Gonzalez MJ . Effects of interleukin-4 on human osteoblast-like cells. Bone Miner 1993; 21: 53–61.
Durie BGM, Urnovitz HB, Murphy WH . RT-PCR amplicons in the plasma of multiple myeloma patients, clinical relevance and molecular pathology. Acta Oncologica 2000; 39: 789–796.
Hassett C, Robinson KB, Beck NB, Omiecinski CJ . The human microsomal expoxide hydrolase gene (EPHX1): complete nucleotide sequence and structural characterization. Genomics 1994; 23: 433–442.
Fretland AJ, Omiecinski CJ . Epoxide hydrolases: biochemistry and molecular biology. Chem Biol Interact 2000; 129: 41–59.
Omiecinski CJ, Hassett C, Hosagrahara V . Epoxide hydrolase- polymorphism and role in toxicology. Toxicol Lett 2000; 112–113: 365–370.
Berndt SI, Johnson D, Crowley J, Durie BGM, Hoover R, Katz M et al. Large scale evaluation of genetic variation and the risk of multiple myeloma. Blood 2008; 112, Abstract # 1679.
Acknowledgements
This investigation was supported in part by an unrestricted grant from the International Myeloma Foundation (Bank on a Cure project), as well as by the following PHS Cooperative Agreement grant numbers awarded by the National Cancer Institute, DHHS: CA32102 and CA38926 (SWOG); and CA21115 (ECOG); plus CA 97513 (JDS and BB).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Durie, B., Van Ness, B., Ramos, C. et al. Genetic polymorphisms of EPHX1, Gsk3β, TNFSF8 and myeloma cell DKK-1 expression linked to bone disease in myeloma. Leukemia 23, 1913–1919 (2009). https://doi.org/10.1038/leu.2009.129
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2009.129
Keywords
This article is cited by
-
Pathogenesis of bone disease in multiple myeloma: from bench to bedside
Blood Cancer Journal (2018)