Abstract
Ultraviolet radiation (UVR) mutagenesis causes nearly all cutaneous melanomas, however, since UVR signatures are largely absent in acral melanoma, as well as melanoma in sun-protected sites, the cause of these melanomas is unknown. Whole-genome sequencing data generated as part of the Australian Melanoma Genome Project was supplemented with a detailed histopathological assessment with the melanomas then classified as UVR or non-UVR related, based on their mutation signatures. The clinicopathological characteristics of melanomas with mutation signatures for their subtype were compared. Three (of 35=8.6%) acral melanomas, all clinically and pathologically verified as arising from acral or subungual locations, had predominant UVR mutation burden, whereas four (of 140=2.9%) cutaneous melanomas showed predominant non-UVR mutations. Among the acral melanomas, the few that were UVR dominant occurred in younger patients, had a higher mutation load and a proportion of mutation burden due to UVR, which was similar to that in melanomas from intermittently UVR-exposed skin. Acral melanomas with a UVR signature occurred most frequently in subungual sites and included tumors harboring BRAF or NF1 mutations. Cutaneous melanomas dominated by non-UVR signatures had lower mutation burdens counts and their primary tumors were thicker and had more mitoses than in other cutaneous melanomas. No histopathological features predicted UVR dominance in acral melanomas or non-UVR dominance in cutaneous melanomas. Our finding of acral/subungual melanomas with predominant UVR mutagenesis suggests that the nail plate and acral skin do not provide complete protection from UVR. Our data also confirm that cutaneous melanomas not caused by UVR are infrequent. Identifying where mutation burden is discordant with primary tumor anatomical site is likely to be clinically significant when determining treatment options for metastatic acral and cutaneous melanoma patients.
Similar content being viewed by others
Main
Malignant tumors arise as a result of the accumulation of multiple genetic abnormalities and melanoma has the highest mutation load of any malignancy.1 It has been established that melanomas arising in different anatomical locations with varying degrees of exposure to ultraviolet radiation (UVR) are caused by different genetic alterations.2
For many decades it has been recognized that cutaneous melanoma incidence is associated with fair skin3 and UVR exposure,4, 5 however, this relationship is not simple.6 Furthermore, melanomas arising in areas of intermittent UVR exposure, such as the trunk, back and abdomen, and those arising in sites of chronic UVR exposure, such as head and neck, have different associations with risk factors, histological appearances, and mutation profiles.7, 8 Melanomas from intermittently UVR-exposed sites tend to occur in younger patients, more commonly arise in pre-existing benign nevi, more often have superficial spreading histology and more often BRAF mutated, compared with melanomas arising in chronically exposed skin.7 In contrast, the latter tend to occur in older people, are more commonly of lentigo maligna histological subtype and are less likely to carry BRAF mutation.2
Cutaneous melanomas are defined as those arising non-glabrous skin, thereby excluding the plantar or palmar and the nail apparatus. They are predominantly caused by exposure to UVR, which acts as a mutagen by forming covalent links between two adjacent pyrimidines.9 The subsequent conformation change in the DNA is detected and normally repaired by DNA damage response mechanisms. However, inefficiencies in DNA damage repair can lead to mutations, most frequently C>T or CC>TT transitions, which are the dominant point mutations of UVR.1, 10, 11
The incidence of acral melanoma, defined as those arising on glabrous skin of the palms and soles and in the nail apparatus, is similar across different populations.12 Furthermore, acral melanomas are thought to be protected from UVR by a thick strata corneum13 or nail plate.14 Therefore, it has been hypothesized that acral and sun-shielded mucosal melanomas have different genetic causal pathways unrelated to UVR exposure. This is typically reflected in low mutation burden, which is offset by numerous focused gene amplifications and deletions or structural abnormalities.2, 13, 15 They also have characteristic histological features, including a predominantly lentiginous growth pattern.8, 16
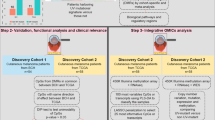
We recently performed the first high coverage whole-genome sequencing study of a large cohort of cutaneous, acral, and mucosal melanomas, which enable a high-resolution assessment of mutation signatures in melanomas arising in these different anatomical sites (currently submitted for review). Here we focus comprehensive clinical, pathological, and molecular analysis on the features of melanomas with mutation signatures that are discordant with their anatomical site, namely cutaneous melanomas dominated by non-UVR mutation signatures and acral melanomas dominated by UVR mutation signatures. Since a tumor’s molecular profile, rather than the primary tumor anatomical site, will determine the most appropriate treatment options for individual melanoma patients, these features are likely to be clinically significant.
MATERIALS AND METHODS
Patients and Specimens
The study was undertaken with Human Ethics Review Committee approval and patient informed consent. One hundred and eighty-three melanomas were whole-genome sequenced and data processed as part of the Australian Melanoma Genome Project17 (currently submitted for review). These cases included melanomas arising from cutaneous, acral, and mucosal primary anatomical sites and also melanomas with unknown primary sites. Fresh surgical specimens were macro-dissected and tumor tissues were procured (with as little contaminating normal tissue as possible) and snap frozen in liquid nitrogen within 1 h of surgery. All samples were pathologically assessed before inclusion into the study, with samples requiring greater than 80% tumor content and less than 30% necrosis to be included.17 All samples were independently reviewed by a pathologist to confirm the presence of melanoma and fulfillment of the above criteria. Samples requiring tumor enrichment underwent macrodissection or frozen tissue coring (Cryoxtract, Woburn, MA, USA) using a marked H&E slide as a reference.
The histopathology of all mucosal and acral samples was reviewed by RVR and RAS to confirm the diagnosis and anatomical site of the primary tumor. Acral melanomas were classified as occurring within acral skin of the palm of the hand, sole of the foot, and in subungual locations under nail beds. Clinical site, lack of hair follicles, and thickened stratum corneum/nail apparatus was confirmed for all acral cases with reference to clinical photos and notes whenever possible. Primary mucosal melanomas were defined as occurring in the mucosal membranes lining oral, respiratory, gastrointestinal, and urogenital tracts. The H&E slides of the primary melanomas were reviewed for all mucosal and acral tumors and their anatomical location confirmed histologically.
Patient demographics, primary tumor site, primary tumor characteristics (Breslow thickness, Clark level, lymphatic invasion, vascular invasion, neurotropism, mitotic rate, regression status, satellite status, T stage, sentinel node status, ulceration, and time to first recurrence) and follow-up data were retrieved from the MIA Research Database. In patients with multiple primary melanomas, the culprit melanoma was assigned using a previously described algorithm.18
Whole-Genome Sequencing
Whole-genome sequencing (WGS) was performed as previously described.17 All the WGS data mentioned in this manuscript has been made publically available on the ICGC Data Portal.
Mutation Signatures
Mutation signatures were identified from WGS data as described.1 Tumors were classified as ‘UVR’-dominant when UVR signature accounted for more than 60% of the total mutation burden. Tumors were defined as ‘non-UVR’ dominant when the mutation burden did not fulfill the above definition.
Histopathological Features
For all cases, corresponding formalin-fixed, paraffin-embedded (FFPE) tumor specimens of the primary tumor were sought from the laboratories where the sequenced tissue was sourced and when identified, H&E-stained sections were obtained and reviewed (n=74) and a semi-quantative analysis was performed on a range of histopathological features as follows:
Tumor-Infiltrating Lymphocyte (TIL) Grade
TIL-grade was assessed by a seven-tier semi-quantitative grading scheme, which was been adapted by us from a previous published study19 for pathological evaluation of cases in The Cancer Genome Atlas Melanoma Project (TCGA).20 The scheme is based on a sum of the assessment of TIL density from 0 to 3 (negative, mild, moderate, or marked) and TIL distribution from 0 to 3 (negative, focal, multifocal, or diffuse) across the entire extent of the invasive tumor (no TILs to marked-diffuse TILs, 0–6, respectively).
Pigmentation
Pigmentation was assessed by a seven-tier semi-quantitative grading scheme based on the sum of an assessment of pigment distribution from 0 to 3 (0, 0–25, 25–50, or >50% of cells pigmented) and average pigment density assessed from 0 to 3 (absent, mild, moderate, or severe) across the extent of the invasive tumor (no pigmentation to diffuse severely pigmented, 0–6, respectively), as previously described.21 Melanophages were not considered in interpretation.
Regression
Regression was assessed as to the percentage of tumor affected from 0 to 3 (0, 0–25, 25–50, or >50%).
Size and Shape of Melanoma Cells
The size and shape of the melanoma cells were assessed utilizing a × 20 objective and based on a semi-quantitative grading scheme adapted from a previously published study.21, 22 The size of the lesional cells was compared with that of a lymphocyte and graded from 0 to 2 (<2 ×, 2 × , and >2 × the diameter of a lymphocyte, respectively). The shape of the cells was determined by the length of the long and short axis of the cells and graded from 0 to 2 (epithelioid with long and short axis approximately equal, mixed epithelioid and spindled, spindled with the long axis at least double the length of the short axis).
Tumor Border
The tumor border was assessed at × 20 power as to the predominant pattern at the advancing edge of the melanoma (pushing, infiltrative).
Solar Elastosis
The severity of solar elastosis was assessed in a semi-quantitative grading system as judged at a × 20 objective from 0 to 3 (absent, mild, moderate, or severe/homogeneous).
Upward Scatter
The amount of upward scatter of melanocytes within the epidermis was assessed in a semi-quantitative grading system based on a grading system previously reported,22 which in short assess the amount of epidermal melanocytes located within the basal layer of the epidermis and graded 0 to 3 (100, 75–100, 50–75, or <50%)
Lateral Circumscription
The amount of lateral circumscription within the epidermal component was assessed in a semi-quantitative grading system previously reported,22 which in short assess the abruptness of the transition from the intraepidermal component of the lesion to adjacent uninvolved epidermis. This is graded from 0 to 2 (areas of apparently uninvolved tumor interspersed with normal, gradual decrease in melanocytes with a transition that is difficult to pin point, abrupt transition from involved epidermis to normal appearing skin over a distance of 1 or 2 rete ridges or 0.1 mm).
Statistical Methods
Descriptive statistics were provided for UVR/non-UVR signature and acral/cutaneous melanomas. Continuous variables were summarized by their mean (s.d.), median, first and third quartiles. For categorical variables frequency and percentage are reported. No inference was performed given the limited sample size, however, difference in means and their 95% confidence interval are reported to assess the magnitude of the difference between the continuous variables
RESULTS
Proportion with UVR Dominance
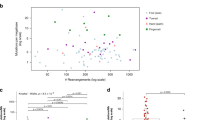
Of the 183 melanomas, 35 were from acral sites including 10 from subungual locations, eight were mucosal melanomas, and the remaining 140 were from other cutaneous, non-acral skin, or had arisen from an unknown primary site. Melanomas from unknown primary sites were arbitrarily considered to be cutaneous melanomas as they have been shown to have similar mutation profiles to cutaneous melanoma in prior studies.23, 24 Cases were then classified as UVR or non-UVR based on the proportion of UVR signatures in the overall mutation burden, as defined in ‘Methods’ section. As expected, no mucosal melanomas were found to be UVR dominant. Seven tumors had mutation burdens that were discordant with their origin: three of the 35 (9%) acral melanomas (Figure 1) were found to be UVR dominant while four of the 140 (3%) (Figure 2) cutaneous melanomas lacked UVR signatures (Table 1 and Figure 3). All non-acral melanomas arising from the hands and feet had dominant UVR signatures (Table 4).
Cohort of 175 melanomas (140 cutaneous, 35 acral) illustrating total mutations of each melanoma against proportion of UVR signature in total mutation burden. (a) Depicts the total number of mutations (SNVs/indels) per sample, patients are colored based on the melanoma subtype/mutation signature. (b) Depicts the relationship between number of structural variants and the proportion of UVR-related mutations.
UVR Dominant Acral Melanoma
The three acral melanomas dominated by UVR signatures are described in Table 2. Two had arisen as primaries located on a subungual location involving the thumb, while the third occurred on the sole of the foot. All were males, aged 58–64 years (median 61 years) at the time of diagnosis of the primary. At the time of diagnosis, metastatic disease was present within regional lymph nodes in all cases with no distant metastatic disease, ie, American Joint Committee on Cancer (AJCC) 7th edition stage IIIA–IIIC.25 All cases were of acral lentiginous melanoma subtype with Breslow thickness 1.3–4.0mm (median 3.2), tumor mitotic rate 2–3 mm/2 (median 3) and two of the cases were ulcerated.
The 32 acral melanomas lacking UVR signatures are described in Table 3. In brief, 21 of the cases were female and 11 male, aged 34–90 years (median 72 years) at the time of primary diagnosis. Breslow thickness ranged from 0.8 to 14.5 mm (median 4.3 mm), tumor mitotic rate 0 to 15 mm/2 (median 4.0 mm/2), 14 (44%) cases were ulcerated, and 13 (41%) of cases were AJCC stage either III or IV at time of diagnosis.
Even though no P-value was calculated, due to the small sample size, the 95% confidence intervals (CI) have shown non-UVR acral melanomas are more likely in older patients and have more mitoses than acral melanoma with a UVR signature (Table 3).
Cutaneous Melanoma Cases without Dominant UVR Signatures
The four cases of cutaneous melanoma without UVR signatures, two male and two females aged 49–68 years (median 60 years), are described in Table 2. In two cases, the primary melanoma was located on the thorax, one was on the abdomen, and one was on the lower ankle/lateral aspect of the foot. At the time of diagnosis, three patients had metastatic disease within regional lymph nodes with no distant metastatic disease, AJCC 7th edition stage IIIB–IIIC25 while the other case was stage IB at the time of diagnosis. Breslow thickness ranged from 0.5 to 20 mm (median 8.65 mm), tumor mitotic rate 5 to 30 mm/2 (median 10.5 mm/2) with two cases ulcerated.
The 136 cutaneous melanomas with a UVR signature are described in Table 3. In brief, 46 of the cases were female and 89 male aged 17–92 years (median 60 years). The Breslow thicknesses ranged from 0.3 to 20 mm (median 2.7 mm), tumor mitotic rate ranged from 0 to 40 mm/2 (median 6.0 mm/2), 42 (31%) cases were ulcerated, and 41 (30%) of cases were AJCC stage III or IV at time of diagnosis.
Although the Breslow thickness and tumor mitotic rate were higher in the non-UVR group than the UVR group there was no significant difference as the 95% CI includes zero (Table 3).
Histopathological Features
H&E-stained sections of the primary tumors were available for review in three (100%) and 23 (72%) of the acral melanomas with a UVR signature and non-UVR signature, respectively, and the findings are outlined in Table 4. Forty-four (32%) and four (100%) of the cutaneous melanomas with a UVR signature and non-UVR signature, respectively, had H&E-stained sections available for review and the findings are outlined in Table 4.
All cutaneous melanomas with a non-UVR signature and all non-cutaneous melanoma with a UVR signature, where the sections of the primary were available, were reviewed to confirm anatomical location of the primary tumor. For example, in case 5 the primary site was described as the lateral ankle, however, acral skin was noted at one end of the histological specimen but not overlying the primary tumor. Clinical photos confirmed this to be correctly classified as a cutaneous melanoma, which occurred in close proximity to the transition to acral skin (Figure 2).
No histopathological feature predicted a discordant classification by UVR status. Of note, there was no solar elastosis present in the three UVR-dominant acral melanomas or the four cutaneous melanomas without UVR dominance.
Tumor Mutation Burden and UVR Dominance
The total burden of point mutations, comprising of single nucleotide variations (SNV) and indels, detected in the three cases of acral melanoma with a UVR signature and the four cases of cutaneous melanoma with a non-UVR signature are detailed in Table 4 and compared with those of the comparison groups of acral and cutaneous melanomas. Figure 3 relates total count of SNV and structural variants (SV) to the relative burden of UVR-related point mutation signatures.
For three acral melanomas with dominant UVR signatures, the total mutation burden per case was 7175 to 45 751 (median 26 271), including 4405 to 36 139 (median 16 717) C>T transitions, 41 to 310 (median 244) CC>TT transitions, and 71 to 234 (median 118) SV. One case was found to harbor a BRAF mutation, one a NF1 mutation, and the third had no mutation detected in BRAF, RAS, or NF1. The proportion of the mutation burden due to UVR was 72.5 to 76.9%. In contrast, the acral melanomas without a dominant UVR signature had a total mutation burden from 1610 to 19 128 per case (median 5779), including 202 to 8416 (median 1709) C>T transitions, 1 to 431 (median 4) CC>TT transitions, 41 to 1132 (median 335) SV; the proportion of the mutation burden due to UVR signatures was much lower in these cases, ranging from 0 to 41.5% (median 4.2%).
For the four cutaneous melanomas with no dominant UVR signature, the total mutation burden per case was 2122 to 11 583, including 138 to 3469 (median 1870) C>T transitions, 2 to 203 (median 6.5) CC>TT transitions, and 52 to 1123 (median 79) SV. Two of the four cases harbored a BRAF mutation while the other two cases harbored no mutation in BRAF, RAS, or NF1. The proportion of the mutation burden due to UVR was 0 to 26.3%. In contrast, the cutaneous melanomas with dominant UVR signatures had a total mutation burden from 13 111 to 775 848 (median 92 605), including 12 258 to 645 517 (median 76 817) C>T transitions, 124 to 6189 (median 932) CC>TT transitions, and 3 to 657 (median 63.5) SV; the proportion of the mutation burden due to UVR signature was much higher in these cases ranging from 70.3 to 100% (median 98.6%).
DISCUSSION
Melanoma has the highest rate of somatic mutation of any malignancy.1 Exome and whole-genome sequencing studies have confirmed that this mutation burden is dominated (median 82.5% in one study) by cytosine to thymidine transition (C>T),10, 11, 26, 27 which results from UVR exposure. In contrast, UVR signatures account for a much lower proportion of the mutation burden in acral melanomas; 25–49% in one study. The total number of mutations correlates with UVR exposure, ranging from greater than 100 000 mutations for those in chronic sun-exposed locations to 30 000 mutations in skin of intermittently UVR-exposed locations to <1000 mutations for those in acral/mucosal sites. Conversely, acral and mucosal melanomas have been shown to harbor more chromosomal aberrations and other structural variations2 than cutaneous melanomas.
Utilizing data generated in the first high coverage whole-genome sequencing study of a large cohort of melanomas (183 cases, currently under review), we have analyzed the characteristics of acral and cutaneous melanomas with mutation burdens that are discordant with these patterns: three acral melanomas with a dominant UVR signature and four cutaneous melanomas without a dominant UVR signature.
Acral and Subungual Melanomas with a Dominant UVR Signature
The acral melanomas with a dominant UVR signature had a high number of mutations when compared with the acral melanomas without a dominant UVR signature. In previous exome and whole-genome sequencing studies, mutations per Mb in acral melanomas have been reported to range from 1.02 to 3.68 (5 cases),15 1.95, 28 and 2.79 to 14 (two cases).27 In contrast, the mean mutation rate of The Cancer Genome Atlas Network study,20 which consisted entirely of cutaneous melanomas, was 16.8 mutations/Mb. Two of the cases we identified in the current study, cases 2 and 3, displayed a mutation burden of 8.76 and 15.25 mutation/Mb, respectively. Although mutation rates are not generally comparable from one study to the next due to different methodologies, it would appear that these cases have a mutation burden more in keeping with cutaneous melanomas. The only other reported similar rate of mutation burden in an acral melanoma was that in case ME032.27
There have been recent conflicting reports on UVR signatures in sequenced acral melanomas. Some studies report no dominant UVR signature within acral melanomas10, 15, 26 while other studies report a dominant UVR signature in cases of acral melanoma: one case report of whole-genome sequencing;28 two of six cases of melanoma cell lines examined;29 and one of two cases of whole-genome sequencing.27 In our view, these findings probably relate to the case selection; some cases designated as arising from acral sites may have involved the non-acral surfaces of the hands or feet. However, in one study,10 the authors postulated that this discrepancy may be due to a low threshold used for C>T transitions used to constitute a UVR signature and the percent of total mutations at a dipyrimidine and the percent of C>T transitions that were at a dipyrimidine was not reported. In the three cases, we describe the proportion of UVR signature constituting the total mutation burden ranged from 72.5 to 76.9%. These levels, while at the lower end of the range for UVR signature (Figure 3), appear sufficient to provide confidence that they are correctly identified as primarily UVR driven. Further evidence of this is the elevated total mutation load within these cases and the generally lower level of structural variations when compared with acral melanomas with a non-UVR signature.
It has been shown that the mean age of diagnosis for acral melanoma is higher than that of cutaneous melanoma.30 In our series of acral cases with a dominant UVR signature, we report the age of the patient at the time of diagnosis of the primary to be lower than those without a dominant UVR signature. This finding provides further evidence that these acral melanomas possibly behave more like cutaneous melanomas than acral melanomas due to the different mutation processes that are driving them.
Interestingly, two of the three cases of acral melanoma with a dominant UVR signature occurred in subungual locations on the thumb and these two cases had the more elevated mutation burden. Although the eight other subungual melanomas, which had no UVR signature, had a mutation burden more in keeping with an acral melanoma (Table 4). Examining of human cadaveric nails has previously showed that all of UVB and the vast majority of UVA light is blocked by the nail plate.14 The fact that two of the cases we reported in the thumb and a previous case report in the toe28 of subungual melanoma with a dominant UVR signature appears to refute these findings. It would therefore appear that in some patients sufficient UVR can penetrate the nail plate to cause mutation effects in the melanocytes in the underlying subungual tissue. The other acral melanoma with a dominant UVR signature was noted to arise on the sole of the foot. This would also seem to argue against the traditional theory that the thickened stratum corneum on the soles of the feet and the palms of the hand, in addition to the fact that in the majority of the population they represent anatomical locations rarely exposed to UVR, provides a protective environment from UVR.13 However, it should be noted that in some people through various occupational or recreational activities, such as excessive sun bathing, these areas could be exposed to a significant amount of UVR. In case 1, which involved the sole of the foot, we could not confirm that the patient had been involved in any such activities.
This is the first study, as far as we are aware, to compare histopathological features of acral and cutaneous melanomas with and without a dominant UVR signature. Within all the parameters examined, there were no particular histopathological features which appeared predictive of an acral melanoma with a dominant UVR signature compared with a non-UVR dominant signature. Of note, no histopathological evidence of solar damage was noted in the three acral melanomas with a UVR signature.
In summary, the intermediate total mutation count, lower number of structural variations, and lack of solar damage histologically indicate that these acral melanomas with a dominant UVR signature have a similar pathogenesis, and therefore possibly have similar clinical behavior, to melanomas arising in intermittently exposed skin, such as the torso or abdomen. This novel hypothesis appears to broaden our understanding of the heterogeneous nature of mutagenesis in acral melanoma.
Cutaneous Melanomas Lacking a UVR Signature
Of the cutaneous melanomas with a non-UVR dominant signature, the mutation load, C>T, and CC>TT transitions are lower than those with a UVR dominant signature and in the range of acral melanomas with a non-UVR dominant signature (Figure 3 and Table 3). Interestingly, the number of structural variations in case 4, 5, and 7 are low when compared with acral melanomas with a non-UVR dominant signature with case 6 harboring an exceedingly large burden of structural variations. These findings lead us to believe that there is considerable heterogeneity in melanomas with a non-UVR dominant signature with multiple, at this stage unknown, causative factors.
Case 5 was anatomically located in the ankle and clinically not an acral melanoma. Histopathology revealed that the tumor was centered on non-acral skin, which at the inferior aspect of lesion merged with acral skin. The remainder of these cases were from sites which would generally only receive intermittent UVR exposure (torso and abdomen).
Interestingly, in cutaneous melanomas with a non-UVR dominant signature, the tumor mitotic rate and Breslow thickness appeared higher than in the comparison group of cutaneous melanomas with a dominant UVR signature, although this difference was found not to be significant. These two criteria along with tumor ulceration are known to represent the most important prognostic factors for primary melanomas.25 Results from the TCGA melanoma project,20 which looked only at cutaneous melanoma, were also reviewed for any differences in high-risk features between UVR and non-UVR dominant melanomas (Supplementary Figure 1). These findings support our results with a greater Breslow thickness (P=0.0223) noted in the non-UVR group. Also, although no statistical significance was found, the non-UVR group were more frequently ulcerated (P=0.0691) and had a greater mitotic rate than the UVR group. Previously, cutaneous melanomas have been found to be thinner than acral melanomas and also to have a better prognosis.30 Extrapolating these findings, although the numbers we have reviewed are small and no statistical significance was found between the two groups, it could indicate that cutaneous melanomas with a non-UVR dominant signature are more similar in pathogenesis and possible clinical behavior to acral melanoma than cutaneous melanoma with a UVR signature.
Also of interest was the finding that, two of these four cutaneous melanoma cases (case 4 and 5) harbored a BRAF V600E mutation and these same two cases had a larger proportion of UVR signature compared with the two BRAF wild-type cases. As BRAF V600E mutations occur early in mutagenesis and are associated with exposure to intermittent UVR exposure rather than chronic UVR exposure,2, 7, 31 it is possible that early low-dose intermittent UVR exposure is sufficient to enable BRAF mutation after which other, at this point unknown, mutation drivers become the predominant mutagenic process in this subset of melanomas.
Within the histopathological parameters examined, there was no particular feature found that was predictive of cutaneous melanoma with a non-UVR dominant signature compared with a dominant UVR signature.
Following the first high coverage whole-genome sequencing study of a large cohort of melanomas, we identified interesting cases which appeared to have contradictory UVR signatures based on the anatomical site of the primary tumor. These findings seem to resolve an area of uncertainty in the literature that a proportion of acral melanomas, particularly from subungual locations, are driven by UVR mutagenic process. We hypothesize that, based on mutation load, number of structural variations, age of onset and at times BRAF mutation status, these types of acral melanoma might more appropriately be grouped and viewed with melanomas arising from sites of intermittent UVR exposure. Conversely, some melanomas from cutaneous sites are not caused by UVR-associated mutation processes and could be more similar to acral melanomas in terms of mutagenesis, epidemiological features, and clinical behavior. Interestingly, in comparison with acral melanoma with a non-UVR dominant signature, three of the four cases of non-UVR cutaneous melanoma had relatively low structural variations. This highlights the genetic heterogeneity in non-UVR signature melanoma. Further research to investigate the spectrum of potential causative mechanisms in these melanomas will be of great interest.
In addition, we are the first study, as far as we are aware, to perform in-depth histopathological correlation of these interesting cases with seeming mismatch of mutation signatures. Taken together, these findings add to an expanding knowledge of the heterogeneous nature of melanoma. In particular, the novel findings we have presented could lead to further investigation into the poorly understood non-UVR signature mutagenic processes in melanoma, which could help guide preventative measures, ensure appropriate management, and guide research into future treatment options for melanoma.
References
Alexandrov LB, Nik-Zainal S, Wedge DC et al. Signatures of mutational processes in human cancer. Nature 2013;500:415–421.
Curtin JA, Fridlyand J, Kageshita T et al. Distinct sets of genetic alterations in melanoma. N Engl J Med 2005;353:2135–2147.
McGovern V . Melanoblastoma. Med J Aust 1952;1:139–142.
Lancaster HO . Some geographical aspects of the mortality from melanoma in Europeans. Med J Aust 1956;43:1082–1087.
Bulliard JL, Cox B, Elwood JM . Latitude gradients in melanoma incidence and mortality in the non-Maori population of New Zealand. Cancer Causes Control 1994;5:234–240.
Whiteman DC, Pavan WJ, Bastian BC . The melanomas: a synthesis of epidemiological, clinical, histopathological, genetic, and biological aspects, supporting distinct subtypes, causal pathways, and cells of origin. Pigment Cell Melanoma Res 2011;24:879–897.
Bauer J, Buttner P, Murali R et al. BRAF mutations in cutaneous melanoma are independently associated with age, anatomic site of the primary tumor, and the degree of solar elastosis at the primary tumor site. Pigment Cell Melanoma Res 2011;24:345–351.
Broekaert SM, Roy R, Okamoto I et al. Genetic and morphologic features for melanoma classification. Pigment Cell Melanoma Res 2010;23:763–770.
Daya-Grosjean L, Sarasin A . The role of UV induced lesions in skin carcinogenesis: an overview of oncogene and tumor suppressor gene modifications in xeroderma pigmentosum skin tumors. Mutat Res 2005;571:43–56.
Krauthammer M, Kong Y, Ha BH et al. Exome sequencing identifies recurrent somatic RAC1 mutations in melanoma. Nat Genet 2012;44:1006–1014.
Pleasance ED, Cheetham RK, Stephens PJ et al. A comprehensive catalogue of somatic mutations from a human cancer genome. Nature 2010;463:191–196.
Bastian BC . The molecular pathology of melanoma: an integrated taxonomy of melanocytic neoplasia. Annu Rev Pathol 2014;9:239–271.
Bastian BC, Kashani-Sabet M, Hamm H et al. Gene amplifications characterize acral melanoma and permit the detection of occult tumor cells in the surrounding skin. Cancer Res 2000;60:1968–1973.
Stern DK, Creasey AA, Quijije J et al. UV-A and UV-B penetration of normal human cadaveric fingernail plate. Arch Dermatol 2011;147:439–441.
Furney SJ, Turajlic S, Stamp G et al. The mutational burden of acral melanoma revealed by whole-genome sequencing and comparative analysis. Pigment Cell Melanoma Res 2014;27:835–838.
Kuchelmeister C, Schaumburg-Lever G, Garbe C . Acral cutaneous melanoma in caucasians: clinical features, histopathology and prognosis in 112 patients. Br J Dermatol 2000;143:275–280.
Wilmott JS, Field MA, Johansson PA et al. Tumour procurement, DNA extraction, coverage analysis and optimisation of mutation-detection algorithms for human melanoma genomes. Pathology 2015;47:683–693.
Murali R, Brown PT, Kefford RF et al. Number of primary melanomas is an independent predictor of survival in patients with metastatic melanoma. Cancer 2012;118:4519–4529.
Azimi F, Scolyer RA, Rumcheva P et al. Tumor-infiltrating lymphocyte grade is an independent predictor of sentinel lymph node status and survival in patients with cutaneous melanoma. J Clin Oncol 2012;30:2678–2683.
Cancer Genome Atlas Network. Genomic classification of cutaneous melanoma. Cell 2015;161:1681–1696.
Long GV, Wilmott JS, Haydu LE et al. Effects of BRAF inhibitors on human melanoma tissue before treatment, early during treatment, and on progression. Pigment Cell Melanoma Res 2013;26:499–508.
Viros A, Fridlyand J, Bauer J et al. Improving melanoma classification by integrating genetic and morphologic features. PLoS Med 2008;5:e120.
Dutton-Regester K, Kakavand H, Aoude LG et al. Melanomas of unknown primary have a mutation profile consistent with cutaneous sun-exposed melanoma. Pigment Cell Melanoma Res 2013;26:852–860.
Egberts F, Bergner I, Kruger S et al. Metastatic melanoma of unknown primary resembles the genotype of cutaneous melanomas. Ann Oncol 2014;25:246–250.
Balch CM, Gershenwald JE, Soong SJ et al. Final version of 2009 AJCC melanoma staging and classification. J Clin Oncol 2009;27:6199–6206.
Hodis E, Watson IR, Kryukov GV et al. A landscape of driver mutations in melanoma. Cell 2012;150:251–263.
Berger MF, Hodis E, Heffernan TP et al. Melanoma genome sequencing reveals frequent PREX2 mutations. Nature 2012;485:502–506.
Turajlic S, Furney SJ, Lambros MB et al. Whole genome sequencing of matched primary and metastatic acral melanomas. Genome Res 2012;22:196–207.
Furney SJ, Turajlic S, Fenwick K et al. Genomic characterisation of acral melanoma cell lines. Pigment Cell Melanoma Res 2012;25:488–492.
Bradford PT, Goldstein AM, McMaster ML et al. Acral lentiginous melanoma: incidence and survival patterns in the United States, 1986–2005. Arch Dermatol 2009;145:427–434.
Maldonado JL, Fridlyand J, Patel H et al. Determinants of BRAF mutations in primary melanomas. J Natl Cancer Inst 2003;95:1878–1890.
Acknowledgements
This work was supported by Melanoma Institute Australia (MIA), Bioplatforms Australia, New South Wales Ministry of Health, Cancer Council New South Wales, Program Grants of the National Health and Medical Research Council of Australia (NHMRC) and Cancer Institute New South Wales, and the Australian Cancer Research Foundation. RVR is supported by Cameron Fellowship through Melanoma Institute Australia. NKH, NW, RAS, and JSW are supported by NHMRC Fellowships. Sample collection for the Australian Melanoma Genome Project was supported by MIA, the Victorian Cancer Agency, the Victorian Cancer Biobank, Victorian State Government Operational Infrastructure Support Program, the Melanoma Research Alliance, and the Melbourne Melanoma Project. Samples were contributed to these banks, thanks to the efforts of patients, clinicians, and other staff at many health services in Australia. The authors thank Doug Stetner for computing assistance and gratefully acknowledge the support of colleagues at MIA, Royal Prince Alfred Hospital, NSW Health Pathology, The Westmead Institute for Medical Research, Peter MacCallum Cancer Centre, and Olivia Newton-John Cancer Research Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Laboratory Investigation website
Whole genome sequencing data generated as part of the Australian Melanoma Genome Project identified 8.6% of acral melanomas with a predominant UVR mutation burden and 2.9% of cutaneous melanomas with a predominant non-UVR mutation burden. This study performs in depth clinicopathological correlation of these interesting cases with contradictory UVR signatures based on the anatomical site of the primary tumor.
Supplementary information
Rights and permissions
About this article
Cite this article
Rawson, R., Johansson, P., Hayward, N. et al. Unexpected UVR and non-UVR mutation burden in some acral and cutaneous melanomas. Lab Invest 97, 130–145 (2017). https://doi.org/10.1038/labinvest.2016.143
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2016.143
This article is cited by
-
Arsenic is a potent co-mutagen of ultraviolet light
Communications Biology (2023)
-
Clinical features, molecular pathology, and immune microenvironmental characteristics of acral melanoma
Journal of Translational Medicine (2022)
-
Whole-genome sequencing of acral melanoma reveals genomic complexity and diversity
Nature Communications (2020)
-
Patterns of genomic evolution in advanced melanoma
Nature Communications (2018)