Abstract
The poor prognosis of oral cavity squamous cell carcinoma (OCSCC) patients is associated with residual tumor after surgery. Raman spectroscopy has the potential to provide an objective intra-operative evaluation of the surgical margins. Our aim was to understand the discriminatory basis of Raman spectroscopy at a histological level. In total, 127 pseudo-color Raman images were generated from unstained thin tissue sections of 25 samples (11 OCSCC and 14 healthy) of 10 patients. These images were clearly linked to the histopathological evaluation of the same sections after hematoxylin and eosin-staining. In this way, Raman spectra were annotated as OCSCC or as a surrounding healthy tissue structure (i.e., squamous epithelium, connective tissue (CT), adipose tissue, muscle, gland, or nerve). These annotated spectra were used as input for linear discriminant analysis (LDA) models to discriminate between OCSCC spectra and healthy tissue spectra. A database was acquired with 88 spectra of OCSCC and 632 spectra of healthy tissue. The LDA models could distinguish OCSCC spectra from the spectra of adipose tissue, nerve, muscle, gland, CT, and squamous epithelium in 100%, 100%, 97%, 94%, 93%, and 75% of the cases, respectively. More specifically, the structures that were most often confused with OCSCC were dysplastic epithelium, basal layers of epithelium, inflammation- and capillary-rich CT, and connective and glandular tissue close to OCSCC. Our study shows how well Raman spectroscopy enables discrimination between OCSCC and surrounding healthy tissue structures. This knowledge supports the development of robust and reliable classification algorithms for future implementation of Raman spectroscopy in clinical practice.
Similar content being viewed by others
Main
Inadequate surgery (tumor positive (≤1 mm) or close (>1 and ≤5 mm) resection margins) for oral cavity squamous cell carcinomas (OCSCC) varies between 30 and 85%.1 This indicates that currently used visual inspection and tissue palpation by the surgeon are not sufficient to define the borders of the tumor intra-operatively. Frequently, frozen tissue sections are taken during surgery to evaluate whether the margins are tumor free (so-called frozen tissue section procedure). Although state of the art, this procedure has major limitations such as time-consuming whereby only a small percentage (1–5%) of the resection margin can be evaluated.2 The consequences of tumor-positive resection margins are significant since this often leads to revision surgery, the need for adjuvant therapy (post-operative radiotherapy), and higher morbidity and mortality.3, 4
Therefore, several optical diagnostic modalities have entered the field of head and neck oncology in the last decades, aiming to support the surgeon in achieving tumor-free resections. The most studied optical techniques include fluorescence,5, 6 optical coherence tomography,7, 8 elastic light scattering spectroscopy,9 (high-resolution) micro endoscopy10, 11 and Raman spectroscopy.12, 13
Raman spectroscopy is especially suited for intra-operative use since this nondestructive technique does not need any pre-treatment or labeling of the tissue. Previously, several groups investigated the potential application of Raman spectroscopy for tumor detection in the head and neck region,14, 15, 16 including the oral cavity.17, 18, 19, 20, 21, 22, 23, 24 Based on the results of ex vivo and in vivo measurements of oral tissue, it was repeatedly concluded that spectra from normal tissue were recognized by their high lipid content, whereas the spectra from malignant tissue were characterized by their protein features.18, 19, 21, 23 However, the interpretation of the previously published measurements is not straightforward because no exact correlation was made between the histopathological structures in the tissue volume from which the Raman signal was obtained, and the Raman results.
The different layers and structures in healthy oral cavity tissue have their own specific molecular composition and as a consequence their own specific Raman features. OCSCC of the tongue can be surrounded by any of the different healthy tissue structures. In previous research, we developed a method of collecting detailed spectral information of individual histo(patho)logical structures based on in vitro Raman mapping experiments.25
In the current study, we used that method to create a spectral database that comprises OCSCC and healthy tissue structures of the tongue. We investigated how well the tissue spectra enable discrimination between OCSCC and surrounding healthy tissue structures. We discuss how these results would affect development of classification algorithms and future implementation of Raman spectroscopy in clinical practice.
MATERIALS AND METHODS
Sample Handling and Sample Preparation
This study was approved by the Medical Ethics Review Board of the Erasmus MC (number 2011-450). At the Department of Otorhinolaryngology and Head and Neck Surgery of the Erasmus MC cancer institute, University Medical Center Rotterdam, 25 tissue samples were collected from 10 patients who had undergone surgical resection of a primary SCC of the tongue. All patients gave informed consent.
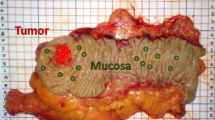
In total, 11 samples contained OCSCC and surrounding healthy tissue structures. These samples were harvested from the surgical resection specimen, more specifically from the center of the macroscopically visible tumor. The 14 normal samples that contained only healthy tissue structures, were harvested from two different locations. Eight samples were harvested from the surgical resection specimen, more specifically from normal-appearing mucosa adjacent to the tumor, i.e., between the tumor border and the resection border (Figure 1). Six samples were collected from the contralateral (not-affected site) edge of the tongue.
All samples were at least 5 × 5 mm in size. Samples from the surgical resection specimen were taken within 60 min after surgical excision. Contralateral samples were taken during surgery. The samples were snap frozen by immersion in isopentane and subsequently in liquid nitrogen, and kept at −80 °C until further use.
For Raman mapping experiments, the frozen tissue samples were mounted on a cryotome stage using CryoCompound (KP-CryoCompound, Klinipath B.V., the Netherlands). Adjacent 6-μm- and 20-μm-thick sections were cut. The 6 μm sections were stained with hematoxylin and eosin (H&E) for histopathological evaluation. The H&E-stained sections were used to select regions for Raman mapping experiments. Corresponding regions were then identified in the unfixed, unstained 20 μm sections that were placed on fused silica windows. These 20 μm sections were allowed to dry at the room temperature and used for the Raman mapping experiments without further treatment. After completing the mapping experiments, the 20 μm sections were also H&E stained.
Raman Spectroscopic Mapping Experiments
Raman spectroscopic mapping experiments were performed using a SpectraCell RA Bacterial Strain Analyzer (RiverD International B.V., Rotterdam, the Netherlands). This instrument was designed as a fully automated inverted confocal Raman microscope for analyzing bacterial samples. After modification of the software, it was used for point-by-point Raman mapping of tissue sections. Each measured point (pixel) represents an individual spectrum.
For each mapping experiment, the selected region was scanned point-by-point in a two-dimensional grid with a step size of 5 μm, and an acquisition time of 1 s per point. About 100 mW of laser light (785 nm) was focused to a spot of 2 μm in diameter, using a custom made infinity corrected objective with an NA of 0.7 and a magnification of 14 (RiverD International B.V.). Raman scattered light was collected in the wavenumber interval from 300 to 2500/cm with a spectral resolution of 4/cm, using a custom made reflection grating with 1650 lines/mm (RiverD International B.V.).
Data Preprocessing
The data preprocessing is described in detail in previous work.25 Briefly, for each individual mapping experiment, the Raman data were processed and analyzed with software developed in-house that operates in a MATLAB environment (MATLAB 7.5.0 (R2007b), MathWorks, MA, USA) and uses the multivariate toolbox PLS-toolbox 7.0.0c (EigenVector Research, WA, USA). After removing cosmic ray events, the raw data were calibrated. Extended multiplicative signal correction combined with spectral interference subtraction26 was used to eliminate all non-informative interfering signals, comprising signal of the fused silica window, CryoCompound, blood, carotenoid (Sigma Aldrich, the Netherlands), hydroxyapatite (Sigma Aldrich), and background fluorescence, as described earlier.25 The contributions of carotenoid and hydroxyapatite represented biological signals that were considered non-informative, since these signals showed a large influence on the scaling procedure, resulting in non-informative differences between similar histopathological structures in different Raman experiments. Finally, the wavenumber range of the spectra that were included in the data-analysis was restricted to the 400–1800/cm.
Data Processing
Processing of the Raman map spectra was done as described in previous work.25 Briefly, principal components analysis (PCA) was used to reduce the data set without significant loss of information.27 All data were mean-centered before PCA. The first 20 PCA scores accounted for >99.5% of the variance present. These PCA scores were grouped using K-means cluster analysis (KCA).28 From the KCA results, a pseudo-color Raman image was generated by assigning a color to each cluster and by plotting the cluster membership of each spectrum as a colored pixel at its measurement position.
Histopathologically Annotated Reference Spectra
After the Raman mapping experiment and H&E staining, a trained pathologist identified the histo(patho)logical structures that were present in the scanned region. For each experiment, the optimal number of K-means clusters was defined, based on the best match between histo(patho)logical structures that were present in the mapped region and the pseudo-color Raman image, as described in detail earlier.25 Figure 2 shows representative examples of H&E-stained sections and their corresponding pseudo-color Raman images (Figure 2). The pathologist was blinded for the corresponding spectra.
H&E stained sections and corresponding pseudo-color Raman images. H&E-stained tissue sections (a, c, e and g) and corresponding pseudo-color images (b, d, f and h). The K-means cluster averages were annotated as one of the following tissue structures: OCSCC (central part, peripheral part, or n.o.s.), squamous epithelium (superficial layers, suprabasal layers, or basal layers), CT (dense and collagen-rich, mixed, or inflammation- and capillary-rich), gland (mucinous or serous), muscle, adipose tissue, or nerve. CT, connective tissue; H&E, hematoxylin and eosin; n.o.s., not otherwise specified; OCSCC, oral cavity squamous cell carcinoma.
These K-means cluster averages were selected on the basis of sufficient signal-to-noise ratio, and on an adequate cluster size to allow histo(patho)logical annotation. The signal was calculated as the peak height at CH2 (1448/cm), and the noise was estimated from the signal part between 1770 and 1800/cm where no signal is present. Clusters with a signal-to-noise ratio <30 were excluded, to ensure that low spectral quality was of no influence to the model, especially for small clusters consisting of few spectra. The cluster size was expressed as percentage (number of representing pixels/total number of pixels in the map). A percentage of ≤5% was used as an exclusion criterion.
The remaining K-means cluster averages were then annotated as one of the following tissue structures: (1) OCSCC, (2) squamous epithelium, (3) connective tissue (CT), (4) gland, (5) muscle, (6) adipose tissue, or (7) nerve, as shown in Figure 2. The K-means cluster averages served as histopathologically annotated reference spectra, and are hereafter called annotated reference spectra.
The annotated reference spectra were further subdivided based on the histopathological heterogeneity of corresponding tissue areas. In this way the OCSCC spectra were subdivided in: (1a) central part, (1b) peripheral part, or (1c) OCSCC n.o.s. (not otherwise specified). Squamous epithelium spectra were subdivided in: (2a) superficial layers, (2b) suprabasal layers, or (2c) basal layers. The CT contains varying amounts of collagen (bundles), blood vessels, and lymphocytes. Accordingly, three subtypes were distinguished: (3a) dense and collagen-rich CT, (3b) mixed CT, and (3c) inflammation- and capillary-rich CT. This second subtype contains loose CT, CT with less collagen, CT with blood vessels, or a mixture of different CT subtypes. Gland spectra were subdivided in: (4a) mucinous and (4a) serous. Figure 3 shows an H&E-stained tissue section which illustrates these subdivisions.
Histopathological structures in H&E-stained tissue section. Micrograph (magnification × 16) of a H&E-stained tissue section of a tumor sample. The squamous epithelium can be divided into three layers; superficial layers (E1), suprabasal layers (E2), and basal layers (E3). The epithelium changes toward the right into dysplastic epithelium (E4). An OCSCC (T1–T3) arises from this dysplasia and can be divided into central part (T1) and peripheral part (T2) or not otherwise specified (n.o.s.) (T3). The underlying lamina propria and submucosa consist of CT that can be divided into dense and collagen-rich CT (CT1), inflammation- and capillary-rich CT (CT2), or mixed CT (CT3). Adipose tissue (A) and muscle tissue (M) are also found in the submucosa. CT, connective tissue; H&E, hematoxylin and eosin; OCSCC, oral cavity squamous cell carcinoma.
A second subdivision of the OCSCC spectra was based on the differentiation grade of the measured tumor: (I) moderately differentiated, (II) moderately to poorly differentiated, or (III) poorly differentiated OCSCC.29 A second subdivision of the squamous epithelium spectra was based on the presence or absence of dysplastic changes; (i) dysplasia, or (ii) no dysplasia.29
Finally, the spectra of the healthy tissue structures ((2) squamous epithelium, (3) CT, (4) gland, (5) muscle, (6) adipose tissue, and (7) nerve) were subdivided depending on their distance to OCSCC. Spectra that were measured in normal (tumor-free) samples were divided in: (A) harvested contralateral (not-affected site of the tongue), (B) harvested adjacent to OCSCC (between tumor border and resection border). Healthy tissue structures that were measured in samples that contained regions of OCSCC were annotated as (C) close to the tumor.
Spectra that could not be histologically annotated were excluded.
Hierarchical Cluster Analysis
The annotated reference spectra of all experiments together were used as input in an unsupervised hierarchical cluster analysis (HCA). The output of HCA is a membership matrix (N × N, where N is the number of spectra) that represents the clustering at each level of agglomeration.28 The HCA results were used to determine the spectral average and s.d. for each histo(patho)logical annotation as described earlier.25 The spectral average represents the overall molecular composition of a histo(patho)logical structure. The s.d. provides insight into the variance in biochemical composition within the histo(patho)logical structure.
Linear Discriminant Analysis
To investigate how well the tissue spectra enabled discrimination between OCSCC and the six surrounding healthy tissue structures, a linear discriminant analysis (LDA) was used.30 Six separate LDA models were made, using the scores of the PCA of the annotated reference spectra as input variables. Each model was tested by leave-one-patient-out (LOPO) analysis, i.e., the models were trained using a subset of nine patients and tested on the remaining one patient. For each model, the optimal number of input parameters was determined by the best result in the LOPO analysis. To prevent overfitting of the LDA model, the amount of input parameters was in all cases smaller than half the number of annotated reference spectra in the smallest group. The classification accuracy (proportion of true results, both true positives and true negatives) served as a measure for discriminative power.
RESULTS
Histopathologically Annotated Database
A total of 127 Raman mapping experiments were performed on 25 samples from 10 patients (11 samples contained OCSCC and surrounding healthy tissue structures, and 14 samples contained only healthy tissue structures). The scanned areas ranged between 405 × 150 μm and 1005 × 470 μm. The step size of 5 μm resulted in 2700 to 24 426 pixels per experiment (mean number of pixels per experiment: 11 538). The optimal number of K-means clusters varied between 4 and 20, resulting in a total of 1114 K-means clusters.
Seven hundred-twenty K-means clusters were annotated based on histo(patho)logical correlation of the scanned tissue regions. In total 88 K-means clusters were annotated as OCSCC, 140 as squamous epithelium, 396 as CT, 41 as muscle, and 33 as adipose tissue. Gland and nerve were both underrepresented in our samples with 17 and 5 annotated reference spectra, respectively. The spectral averages and s.d. of these seven histo(patho)logical tissue structures are shown in Figure 4.
In total, 394 K-means cluster averages were excluded from further analysis. These excluded K-means clusters represented 17% of the total number of pixels of all experiments together. Examples of the excluded K-means spectra are shown in Supplementary Figure S1.
First, 216 K-means clusters did not pass the inclusion criteria. For 62 clusters, the signal-to-noise ratio was <30. The average signal-to-noise ratio of all included spectra was 279 (range: 52–999). The other 154 clusters only represented a few pixels (≤5%) in the scanned regions which made exact histo(patho)logical correlation impossible.
Another 178 K-means clusters were excluded to prevent addition of non-representative contribution to the spectral averages. These include 65 clusters that represented borders between two different tissue structures, and that therefore reveal a composition of the spectral features of both tissue structures. For 83 clusters, no reliable correlation with histopathology was possible, because these clusters consisted of single pixels dispersed throughout several tissue structures in the pseudo-color Raman images. For 20 clusters, contamination or tissue loss during the cutting and H&E staining procedure prevented a reliable correlation. Last, 17 clusters were excluded because they represented structures that were rarely encountered in our samples, such as parakeratotic layers of epithelium and gland’s duct.
Distinction between OCSCC and Its Surrounding Healthy Tissue Structures
To investigate how well OCSCC could be distinguished from the six healthy tissue structures, we made six separate LDA models based on the whole spectrum (i.e., OCSCC vs squamous epithelium; OCSCC vs CT; OCSCC vs gland; OCSCC vs muscle; OCSCC vs adipose tissue, and OCSCC vs nerve). The results of each classification model are given below.
OCSCC vs Squamous Epithelium
Similar to the OCSCC spectrum (pink spectrum Figure 5a), the average spectrum of squamous epithelium (blue spectrum Figure 5a) is dominated by proteins (peaks at 668, 782, 830, 936, 1004, 1156, 1174, 1208, 1302, 1316, 1448, 1514, and 1656/cm).31 The major differences between these spectra are caused by the more intense peaks at 902, 936, and 1656/cm in the squamous epithelium spectrum, and the more intense peaks at 878, 990, 1064, and 1448/cm in the average OCSCC spectrum (enlarged details in Figure 5b). The LDA model classified 39 of the 140 squamous epithelium spectra as OCSCC, and 17 out of 88 OCSCC spectra as squamous epithelium (Figure 5c). This resulted in a classification accuracy of 75%.
OCSCC vs CT
The most discriminative region of the mean CT spectrum (gray spectrum Figure 6a) is attributed to collagen (peaks at 532, 854, 876, 922, 938, 1246, 1266, and 1448/cm).31 The LDA model classified 21 out of 396 CT spectra as OCSCC, and 12 out of 88 OCSCC spectra as CT (Figure 6b). This resulted in a classification accuracy of 93%.
OCSCC vs Gland
Glandular tissue was not often encountered in our samples. Since the mean spectrum was based on spectra of two different types of gland tissue (serous and mucinous) the s.d. was larger than in other tissue types (Figure 4D). Nevertheless the characteristic peaks (440, 718, 874, 1042, 1114, 1156, 1376, and 1518 cm−1) of the mean gland spectrum (teal spectrum Figure 7a) were striking compared with the average OCSCC spectrum (pink spectrum Figure 7a). The LDA model classified 6 out of 17 gland spectra as OCSCC, but none of the 88 OCSCC spectra as gland (Figure 7b), resulting in a classification accuracy of 94%.
OCSCC vs Muscle
The spectral features of OCSCC (pink spectrum Figure 8a) and muscle (red spectrum Figure 8a) are quite similar; however some clear differences are visible. Peaks at 546, 782, 1084, and 1096/cm were present in the average OCSCC spectrum but were absent in the muscle spectrum. The peaks at 936, 1042, and 1396/cm were more intense in the muscle spectrum, compared with the OCSCC spectrum (enlarged details in Figure 8b). The LDA model classified 1 out of 41 muscle spectra as OCSCC, and 3 out of 88 OCSCC spectra as muscle (Figure 8c), resulting in a classification accuracy of 97%.
OCSCC vs Adipose Tissue and OCSCC vs Nerve
The average spectra of nerve (salmon spectrum Figure 9a) and adipose tissue (yellow spectrum Figure 9a) mainly represent lipids (peaks at 962, 1064, 1080, 1302, 1368, 1440, 1558, 1656, and 1744–1748/cm),31 whereas the average spectrum of OCSCC (pink spectrum Figure 9a) is dominated by protein signals (peaks at 668, 782, 830, 936, 1004, 1156, 1174, 1208, 1316, 1448, 1514, and 1656/cm).31 Thanks to these large differences in lipid protein ratios both LDA models showed a classification accuracy of 100% (Figure 9b).
Analysis of Misclassifications by the Use of Subdivisions
Highly accurate discrimination between OCSCC and the majority of healthy tissue structures was possible (CT: 93%, gland: 94%, muscle: 97%, adipose tissue: 100%, and nerve: 100%), with the exception of squamous epithelium (75%). A detailed overview of the misclassifications is given in Table 1.
The spectra of OCSCC, squamous epithelium, CT, and gland, each showed a large variance around their mean (Figure 4A–4D). This reflects the heterogeneity in their histopathological annotations. To understand the cause of the misclassifications, we used further subdivisions of these tissue structures (as described in the materials and methods section).
Misclassifications in Squamous Epithelium
Of the 140 spectra annotated as squamous epithelium, 101 were correctly classified (72%), and 39 were misclassified as OCSCC. The spectra that represented the superficial layers were more often correctly classified as epithelium (84% (37/44)), than the suprabasal layers (78% (31/40)) or basal layers (59% (33/56)), as shown in Supplementary Figure S2.
Spectra that represented squamous epithelium without dysplasia were more often correctly classified as squamous epithelium (78% (87/111)), than the squamous epithelium with dysplasia (48% (14/29)) (Supplementary Figure S2).
The percentage of correct classifications was higher for squamous epithelium in the contralateral normal samples (91% (31/34)), than in the adjacent harvested normal samples (58% (40/69)), or than for squamous epithelium from the samples that contained OCSCC (81% (30/37)) (Table 1). This might be related to previous observation since less dysplasia was seen in the contralateral samples.
Misclassifications in CT
Of the 396 spectra annotated as CT, 375 were correctly classified (95%) and 21 were misclassified as OCSCC. The spectra that represented dense and collagen-rich CT were correctly classified as CT in 100% (62/62), whereas mixed CT was correctly classified in 97% (269/278) and inflammation- and capillary-rich CT in 79% (44/56), as shown in Supplementary Figure S3.
The percentage of correct classifications was lowest for CT in the samples containing OCSCC (92% (144/157)), than for CT measured in normal samples harvested adjacent to OCSCC (between tumor border and resection border) (97% (134/138)) or harvested contralateral (not-affected site of the tongue) (96% (97/101)) (Table 1).
Misclassifications in Gland
Of the 17 spectra annotated as gland, 11 were correctly classified (65%) and 6 were misclassified as OCSCC. The mucinous gland spectra were more often correctly classified as OCSCC (79% (11/14)), than the serous gland spectra (0% (0/3)), as shown in Supplementary Figure S4.
The percentage of correct classifications was higher for gland measured in the contralateral normal samples (100% (5/5)), than measured in adjacent harvested normal samples (86% (6/7)) or than samples containing OCSCC 0% (5/5)) (Table 1).
Misclassifications in OCSCC
OCSCC vs squamous epithelium
Out of 88 spectra annotated as OCSCC, 71 were correctly classified (81%), and 17 were classified as squamous epithelium by the (OCSCC vs squamous epithelium) LDA model.
The spectra that represented the peripheral part were more often correctly classified as OCSCC (91% (21/23)) than the central part (72% (13/18)) or the OCSCC n.o.s. (79% (37/47)), as shown in Supplementary Figure S5A and S5C.
OCSCC vs CT
Out of 88 spectra annotated as OCSCC, 76 were correctly classified (86%) and 12 were classified as CT by the (OCSCC vs CT) LDA model.
The OCSCC spectra that represented the central part were correctly classified in 100% (18/18). More misclassifications occurred in the peripheral part 78% (18/23)), and in OCSCC n.o.s. (85% (40/47)), as shown in Supplementary Figure S5A and S5D.
Differentiation grade
In both distinctions (OCSCC vs squamous epithelium and OCSCC vs CT) the percentage of correct classifications was higher in the poorly differentiated OCSCC (93% (28/30)) and (90% (27/30)), than in the moderately differentiated OCSCC (81% (29/36) and 83% (30/36)) and moderately to poorly differentiated OCSCC (64% (14/22) and 86% (19/22) (Supplementary Figure S5B–5D).
DISCUSSION
Our ultimate goal is to develop an algorithm that helps the surgeon to achieve tumor-free resection margins and as a result improve the outcome of patients with OCSCC. The future implementation of such classification algorithm in clinical practice is dependent on the discriminatory power of Raman spectroscopy. To understand the discriminatory basis of Raman spectroscopy at a histological level, we performed experiments in which spectra and tissue locations (from which the spectra were obtained) were clearly linked. This has enabled us to identify which healthy tissue structures were difficult to distinguish from OCSCC.
OCSCC can be spectrally distinguished from most subepithelial healthy tissue structures with high accuracy (>97% correct discrimination between OCSCC and adipose tissue, nerve, and muscle). Squamous epithelium, CT, and gland were the healthy tissue structures that were most often confused with OCSCC.
Most misclassifications occurred in the distinction between OCSCC and squamous epithelium, especially in the basal layers of epithelium, the dysplastic squamous epithelium, and in the epithelium measured in the samples that contained OCSCC. This is not surprising because OCSCC originates in the epithelial surface (Figure 3). In carcinogenesis, stem cells located in the basal layers of the epithelium acquire genetic alterations, followed by clonal expansion.32 Normal epithelium alters in dysplastic epithelium, which is more often seen adjacent to invasive oral carcinomas33 and is considered to be a pre-malignant disease. As a consequence, the biochemical composition of both tissue structures shows high similarity. Due to clonal expansion of stem cells in the basal layers, the lamina propria is pushed away, and invasion through the basal membrane follows. An OCSCC can develop from well differentiated to moderately differentiated to poorly differentiated OCSCC.34, 35 In each step the similarity with the original squamous epithelium will become smaller. This might explain that moderately differentiated OCSCC is more often correctly classified.
Irrespective of the distance to OCSCC, inflammation- and capillary-rich CT was most often misclassified in the OCSCC vs CT model. This type of CT is more often seen close to tumor (Figure 3). This is caused by two reasons: (1) tumor development is accompanied by an immune response that leads to massive tumor infiltration by inflammatory cells36 and (2) a relatively higher amount of capillaries is present in the CT around the tumor because OCSCC creates new blood supply by stimulating endothelial cell proliferation and formation of new blood vessels.37 This can probably explain why most misclassifications occurred in this type of CT as well as the higher percentage misclassifications in the CT in the samples containing tumor.
Furthermore, the higher percentage of misclassifications in the peripheral part of OCSCC might be explained by the epithelial–mesenchymal transition (EMT). EMT occurs at the periphery of a tumor and is characterized by the loss of epithelial characteristics.38 EMT has a role in invasion and metastasis of cancer cells.39
However, the clinical relevance of these possible misclassifications is questionable. First, the imperfect discrimination between OCSCC and squamous epithelium will not necessarily influence the potential for intra-operative application of Raman spectroscopy. The majority of residual tumor is namely located in the subepithelial, deeper tissue layers. Woolgar et al.40 described that <2% of the resection specimens had solely an involved mucosal margin, whereas 87% of the involved margins was located in the deep soft tissue. In the future it might be an option to develop an algorithm that deals with the real clinical problem by excluding squamous epithelium and discriminating tumor only from the subepithelial healthy structures.
Second, regarding the misclassifications in the CT, it has to be mentioned that in clinical practice a margin of at least 1 cm around the tumor is resected to ensure an adequate resection. A part of inflammation- and capillary-rich CT, as well as the CT close to the tumor will most likely be resected anyway within the 1 cm margin, independent on an intra-operative classification by Raman. However, for algorithm development, this is important to know since over resection (and as a result a less functional organ) by expanding the border beyond the 1 cm margin is also not desirable.
In this study, we determined how well Raman spectroscopy enables discrimination between OCSCC and surrounding healthy tissue structures. This provided knowledge that supports the development of robust and reliable classification algorithms for future implementation of Raman spectroscopy in clinical practice.
References
Smits RWH, Koljenović S, Hardillo J et al. Resection margins in oral cancer surgery: Room for improvement. Head Neck 2015; doi:10.1003/hed.24075.
DiNardo LJ, Lin J, Karageorge LS et al. Accuracy, utility, and cost of frozen section margins in head and neck cancer surgery. Laryngoscope 2000;110:1773–1776.
Priya SR, D'Cruz AK, Pai PS . Cut margins and disease control in oral cancers. J Cancer Res Ther 2012;8:74–79.
Al-Rajhi N, Khafaga Y, El-Husseiny J et al. Early stage carcinoma of oral tongue: prognostic factors for local control and survival. Oral Oncol 2000;36:508–514.
Francisco AL, Correr WR, Pinto CA et al. Analysis of surgical margins in oral cancer using in situ fluorescence spectroscopy. Oral Oncol 2014;50:593–599.
Keereweer S, Hutteman M, Kerrebijn JDF et al. Translational optical imaging in diagnosis and treatment of cancer. Curr Pharm Biotechnol 2012;13:498–503.
Hamdoon Z, Jerjes W, Upile T et al. Optical coherence tomography in the assessment of suspicious oral lesions: an immediate ex vivo study. Photodiagnosis Photodyn Ther 2013;10:17–27.
Wilder-Smith P, Jung WG, Brenner M et al. In vivo optical coherence tomography for the diagnosis of oral malignancy. Lasers Surg Med 2004;35:269–275.
Sharwani A, Jerjes W, Salih V et al. Assessment of oral premalignancy using elastic scattering spectroscopy. Oral Oncol 2006;42:343–349.
Muldoon TJ, Roblyer D, Williams MD et al. Noninvasive imaging of oral neoplasia with a high-resolution fiber-optic microendoscope. Head Neck 2012;34:305–312.
Vila PM, Park CW, Pierce MC et al. Discrimination of benign and neoplastic mucosa with a high-resolution microendoscope (HRME) in head and neck cancer. Ann Surg Oncol 2012;19:3534–3539.
Singh SP, Deshmukh A, Chaturvedi P et al. Raman spectroscopy in head and neck cancers: toward oncological applications. J Cancer Res Ther 2012;8 (Suppl 1):S126–S132.
Srinivasan G. Vibrational Spectroscopic Imaging for Biomedical Applications. Chapter 9: The Current State of Raman Imaging in Clinical Application; 1st (edn). The McGraw-Hill Companies, 2010.
Harris AT, Rennie A, Waqar-Uddin H et al. Raman spectroscopy in head and neck cancer. Head Neck Oncol 2010;2:26.
Hughes OR, Stone N, Kraft M et al. Optical and molecular techniques to identify tumor margins within the larynx. Head Neck 2010;32:1544–1553.
Stone N, Stavroulaki P, Kendall C et al. Raman spectroscopy for early detection of laryngeal malignancy: preliminary results. Laryngoscope 2000;110:1756–1763.
de Veld DCG, Bakker Schut TC, Skurichina M et al. Autofluorescence and Raman microspectroscopy of tissue sections of oral lesions. Lasers Med Sci 2005;19:203–209.
Deshmukh A, Singh SP, Chaturvedi P et al. Raman spectroscopy of normal oral buccal mucosa tissues: study on intact and incised biopsies. J Biomed Opt 2011;16:127004.
Guze K, Pawluk HC, Short M et al. Pilot study: Raman spectroscopy in differentiating premalignant and malignant oral lesions from normal mucosa and benign lesions in humans. Head Neck 2014;37:511–517.
Guze K, Short M, Zeng H et al. Comparison of molecular images as defined by Raman spectra between normal mucosa and squamous cell carcinoma in the oral cavity. J Raman Spectrosc 2011;42:1232–1239.
Malini R, Venkatakrishna K, Kurien J et al. Discrimination of normal, inflammatory, premalignant, and malignant oral tissue: a Raman spectroscopy study. Biopolymers 2006;81:179–193.
Oliveira AP, Bitar RA, Silveira L et al. Near-infrared Raman spectroscopy for oral carcinoma diagnosis. Photomed Laser Surg 2006;24:348–353.
Singh SP, Deshmukh A, Chaturvedi P et al. In vivo Raman spectroscopic identification of premalignant lesions in oral buccal mucosa. J Biomed Opt 2012;17:105002.
Su L, Sun YF, Chen Y et al. Raman spectral properties of squamous cell carcinoma of oral tissues and cells. Laser Phys 2012;22:311–316.
Cals FLJ, Bakker Schut TC, Koljenović S et al. Method development: Raman spectroscopy-based histopathology of oral mucosa. J Raman Spectrosc 2013;44:963–972.
Martens H, Stark E . Extended multiplicative signal correction and spectral interference subtraction: new preprocessing methods for near infrared spectroscopy. J Pharm Biomed Anal 1991;9:625–635.
Jolliffe IT . Principal Component Analysis; 2nd (edn). Springer: New York, NY, USA, 2002, pp 487.
Jain D . Dubes Red Algorithms for Clustering Data. Prentice Hall, 1988, pp 334.
World health Organization Classification of Tumours. In: Barnes LE, Reichart Peds Pathology & Genetics head and Neck Tumours. IARC Press: Lyon, France, 2005, pp 177–180.
Tabachnick Using Multivariate Statistics; 3rd (edn). Harper Colins College Publishers, 1996.
Movasaghi Z, Rehman S, Rehman IU . Raman spectroscopy of biological tissues. Appl Spectrosc Rev 2007;42:493–541.
Braakhuis BJ, Leemans CR, Brakenhoff RH . A genetic progression model of oral cancer: current evidence and clinical implications. J Oral Pathol Med 2004;33:317–322.
Slaughter DP, Southwick HW, Smejkal W . Field cancerization in oral stratified squamous epithelium; clinical implications of multicentric origin. Cancer 1953;6:963–968.
Hanahan D, Weinberg RA . Hallmarks of cancer: the next generation. Cell 2011;144:646–674.
Mills SES . Histology for Pathologists; 3rd (edn). Lippincott Williams & Wilkins, 2004.
Gasparoto TH, de Oliveira CE, de Freitas LT et al. Inflammatory events during murine squamous cell carcinoma development. J Inflamm (Lond) 2012;9:46.
Choi S, Myers JN . Molecular pathogenesis of oral squamous cell carcinoma: implications for therapy. J Dental Res 2008;87:14–32.
Sasahira T, Kirita T, Kuniyasu H . Update of molecular pathobiology in oral cancer: a review. Int J Clin Oncol 2014;19:431–436.
Weber F, Xu YM, Zhang L et al. Microenvironmental genomic alterations and clinicopathological behavior in head and neck squamous cell carcinoma. JAMA 2007;297:187–195.
Woolgar JA, Triantafyllou A . A histopathological appraisal of surgical margins in oral and oropharyngeal cancer resection specimens. Oral Oncol 2005;41:1034–1043.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Laboratory Investigation website
This study shows how well Raman spectroscopy enables discrimination between oral cavity squamous cell carcinoma and surrounding healthy tissue structures. This knowledge supports the development of robust and reliable classification algorithms for future implementation of Raman spectroscopy in clinical practice.
Rights and permissions
About this article
Cite this article
Cals, F., Bakker Schut, T., Hardillo, J. et al. Investigation of the potential of Raman spectroscopy for oral cancer detection in surgical margins. Lab Invest 95, 1186–1196 (2015). https://doi.org/10.1038/labinvest.2015.85
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2015.85
This article is cited by
-
Stimulated Raman histology for histological evaluation of oral squamous cell carcinoma
Clinical Oral Investigations (2023)
-
Raman spectroscopy to discriminate laryngeal squamous cell carcinoma from non-cancerous surrounding tissue
Lasers in Medical Science (2023)
-
Vibrational imaging for label-free cancer diagnosis and classification
La Rivista del Nuovo Cimento (2022)
-
Raman spectroscopy and artificial intelligence to predict the Bayesian probability of breast cancer
Scientific Reports (2021)
-
Risk prediction by Raman spectroscopy for disease-free survival in oral cancers
Lasers in Medical Science (2021)