Abstract
Variable degrees of molecular degradation occur in human surgical specimens before clinical examination and severely affect analytical results. We therefore initiated an investigation to identify protein markers for tissue degradation assessment. We exposed 4 cell lines and 64 surgical/autopsy specimens to defined periods of time at room temperature before procurement (experimental cold ischemic time (CIT)-dependent tissue degradation model). Using two-dimensional fluorescence difference gel electrophoresis in conjunction with mass spectrometry, we performed comparative proteomic analyses on cells at different CIT exposures and identified proteins with CIT-dependent changes. The results were validated by testing clinical specimens with western blot analysis. We identified 26 proteins that underwent dynamic changes (characterized by continuous quantitative changes, isoelectric changes, and/or proteolytic cleavages) in our degradation model. These changes are strongly associated with the length of CIT. We demonstrate these proteins to represent universal tissue degradation indicators (TDIs) in clinical specimens. We also devised and implemented a unique degradation measure by calculating the quantitative ratio between TDIs’ intact forms and their respective degradation-modified products. For the first time, we have identified protein TDIs for quantitative measurement of specimen degradation. Implementing these indicators may yield a potentially transformative platform dedicated to quality control in clinical specimen analyses.
Similar content being viewed by others
Main
Mammalian tissues are maintained in a homeostatic environment by the circulation of blood. When circulation ends—as by excision of tissue from a patient—homeostasis is disrupted and molecular degradation ensues. Lack of oxygen supply causes respiratory distress and electrolyte imbalance, followed by disintegration of protein, RNA, DNA, and other biomolecules. Recent technological advances have elevated surgical cancer specimens to crucially important resources, to obtain individual bioinformation from patients, which is decisive for personalized treatment.1, 2, 3 Variable degrees of degradation, however, occur in specimens before the degradation process can be effectively halted (usually by fixation or freezing techniques). Moreover, a large number of known and unknown pre-analytical variables (PAVs) determine the rapidity of the degradation process. Therefore, the exact state of molecular degradation of individual human biospecimens is unknown.4, 5, 6
The degradation stage of mammalian tissues after extended time of exposure to PAVs has been a central topic of forensic research.7, 8, 9, 10 Forensic research developed various methodologies to assess decay stage by measurement of chemicals, fatty acid, and PCR length.9, 11, 12 These efforts, however, predominantly investigated degradation periods in excess of several days and have minor applicability to surgical specimens. Only more recently, evidence accumulated that biochemical degradation of tissues can be observed within hours or even minutes after tissue excision.13, 14, 15, 16 Although recent work has focused on developing protocols/guidelines for better tissue manipulation and developing anti-degradation reagents such as RNA stabilizers and protease inhibitor cocktails,3, 17, 18, 19, 20 no degradation measurement tool exists currently.
Detrimental effects on structural and biomolecular quality of human biospecimens have been widely acknowledged,19, 21, 22, 23, 24, 25, 26 and analysis of human biospecimens may therefore be affected and impact patient care. Because of lack of standards, the degree of tissue degradation of a specimen cannot be measured, and no comparison can be performed between specimens. Consequently, biodata obtained from specimens of unknown degradation stage may be biased or simply inaccurate. With this study, we attempted to develop an objective standard for tissue degradation measurement. To accomplish this, we pursued the hypothesis that degradation-sensitive markers can be identified via comparative proteomic analysis using experimental degradation models. We attempted to identify degradation-sensitive proteins that may serve as tissue degradation indicators (TDIs) and allow for molecular integrity assessment of clinical tissue specimens before or simultaneously with diagnostic testing.
MATERIALS AND METHODS
Cell Lines
Four human cell lines, 293T/17, MDA-MB-231 (HTB-26), Jurkat (TIB-152), and LNCap (CRL-1740), were acquired from American Type Culture Collection (ATCC, Manassas, VA, USA). Cell lines were cultured according to ATCC’s recommendations. Cell lines were grown until ∼80% confluent and trypsinized. Trypsinized cells were washed three times in PBS (pH 7.4). Equal amounts of cells were assigned to tissue degradation model.
Tissue Materials
Tissues specimens were collected by IRB-approved Yale Pathology Tissue Services (Yale Human Investigation Committee approved protocol 0304025173, expiration date: 4 November 2013). Homogeneous tumors/tissues were collected from excess surgical pathology material including five intracranial meningiomas, one intracranial glioma, five uterine leiomyomas, two non-tumor kidney tissues, and two non-tumor liver tissues. Multiple organs (cerebrum, cerebellum, lung, liver, kidney, heart, spleen, small intestines, large intestines, prostate, uterus, and skeletal muscle) were collected at autopsy from two patients with 13 and 15 h post mortem interval (PMI), and cerebral middle frontal gyrus was collected from 25 autopsy cases with PMIs between 7 and 38 h. Multiple organs were collected from mice with C57BL/6J genetic background. Organ and tissue diagnoses were verified by histological examination.
Experimental Cold Ischemic Time-Dependent Tissue Degradation Model
Collected cells were divided into aliquots; obtained fresh tissues were divided into fractions of similar size. One aliquot/fraction of each subject was immediately snap-frozen, stored in liquid nitrogen, and assigned as cold ischemic time (CIT) time 0. The remaining aliquots/fractions were exposed to 23 °C in a Petri dish to mimic clinical CIT and sequentially snap-frozen at assigned CIT intervals. For cells used for proteomic analysis, the assigned CITs were 0, 0.5, 4, 8, 24, and 48 h; for tissues used for validation studies, the assigned CITs varied (0∼120 h) depending on how many fractions an individual specimen was allowed to be divided into. For validation studies on formalin-fixed paraffin-embedded (FFPE) specimens, tissue fractions were fixed in neutral balanced 4% paraformaldehyde for 12 h immediately after completion of the CIT course, followed by standard paraffin embedding.
Cell Lysis and Protein Preparation
For proteomic analysis, frozen cells were lysed in lysis buffer (7 M urea, 2 M thiourea, 4% CHAPS, 30 mM Tris, 5 mM magnesium acetate, pH 8.5). The lysates were incubated on ice for 0.5 h with occasional vortexing. For western blot analysis, cells/tissues were homogenized with a THQ-handheld homogenizer (Omni International, GA, USA) in RIPA cell lysis buffer with protease inhibitor (Cocktail; Pierce, IL, USA). For proteomic analysis, proteins were purified with 2D Clean-up Kit and quantified with 2D Quant Kit (GE Healthcare, NJ, USA); for other usages, protein concentrations were directly measured with Bradford approach using Protein Assay (Bio-Rad, CA, USA).
Proteomic Comparison between Cells of Different CITs
Two-dimensional fluorescence difference gel electrophoresis (2D DIGE) was used for proteomic profiling following GE Healthcare’s instruction. In brief, 50 μg protein from each experimental group were pre-labeled with both Cy3 and Cy5 fluorescent dyes (two repeats for each experiment) using Minimal CyDyes Kit (GE Healthcare). In addition, an internal standard was prepared by mixing 25 μg of each sample and pre-labeled with Cy2 dye. A schematic gel design is shown in Table 1. Protein (50 μg) of two different groups, together with 50 μg internal standard, were separated on one large format IPG gel stripe (24 cm, pH 3–10) using Ettan IPGphor3 platform (GE Healthcare) for isoelectric focusing. The 2D separation was performed on 12.5% SDS-PAGE gels using Ettan DALTtwelve System (GE Healthcare). Protein spots were visualized with Typhoon 9410 Imager (GE Healthcare). The spectrometric density of spots was analyzed with DeCyder software (GE Healthcare) using automated BVA mode, which assigns a quantitative (numeric) value for each protein spot after normalization with its counterpart in the universal control and creates a statistical value using reads from repeats with built-in ANOVA tools. Protein spots, which satisfied the following conditions, were selected for sequencing: (1) the quantity was continuously increasing or decreasing throughout all observed CIT time points;(2) the quantity was more than 1.5-fold different from the reads of the same spot at neighboring time points with statistical differences (P<0.05); (3) the quantity was in the highest 20% range (likely ‘housekeeping protein’); and (4) the tendency of quantitative change was consistent in all four cell lines. One large protein spot, which had constant quantity across the observation period, was selected for sequencing to serve as a loading control for later validation studies.
Mass Spectrometric Protein Sequencing
The protein candidates were picked up from gels with the robotic Ettan Spot Picker (GE Healthcare) and digested with trypsin using automated Ettan TA Digester (GE Healthcare), followed by matrix-assisted laser desorption/ionization time-of-flight/time-of-flight mass spectrometry (MALDI-Tof/Tof MS) analysis. The instrumentation used was the Applied Biosystems (CA, USA) Model 4800 MALDI-Tof/Tof MS, following manufacturer’s protocol. Reflectron analysis summed 1250 laser shots to generate the peptide fingerprint map, and masses were chosen by the Applied Biosystems 4000 Series Explorer software (version 3.0) for MS/MS acquisition. In parallel, another set of gel pieces were trypsin-digested and loaded for liquid chromatography-MS/MS (LC-MS/MS) analysis; LC was performed on an Agilent 1100 Nano-flow system following standard protocol. The LC was interfaced to a dual-pressure linear ion-trap MS (LTQ Velos, Thermo Fisher, MA, USA) via nano-electrospray ionization. An electrospray voltage of 1.8 kV was applied to a pre-column tee. The MS was programmed to acquire tandem mass spectra from the top 15 ions in the full scan from 400–1400 m/z. MS RAW data files were converted to MGF format using msconvert and searched using X!Hunter against the latest library available on the GPM at the time. Proteins were required to have two or more unique peptides with E-value scores of 0.01 or less. Protein clusters were generated using the Search Tool for the Retrieval of Interacting Genes (STRING) database resource via its web portal (http://string-Db.org). The interactions included direct (physical) and indirect (functional) associations, and they were derived from genomic context, high-throughput experiments, co-expression, and previous knowledge (PubMed).
Validation of Proteomic Findings using Antibody-Based Immunoassays
Duplicated MS protein sequencing generated two independent data sets. Proteins with consensus IDs from both approaches were selected for antibody-based validation studies using one-dimensional (1D) and 2D western blotting following standard procedures. Briefly, ReadyStrip IPG Strip (pH 4–7), mini-Protean-TGX gel (12%), and 0.2 μm nitrocellulose membrane sandwich were obtained from Bio-Rad and used for protein separation and solid-phase transfer. The following antibodies were obtained from Abcam (MA, USA): mouse monoclonal anti-AHNAK (1:500), anti-alpha fodrin (D8B7, 1:1000), anti-actin (1:400), anti-desmoplakin I+II (1:100), rabbit monoclonal anti-alpha fodrin (EPR3017, 1:2000), anti-EEF2–C-terminal (1:10 000), rabbit polyclonal anti-EEF2 (phosphor T56) (1:300), and anti-vimentin (1:1000). Rabbit monoclonal anti-p53 (1:1000), rabbit polyclonal anti-cleaved alpha fodrin (Asp1185; 1:1000), anti-NPM (1:1000), and anti-UBE1a (1:1000) were obtained from Cell Signaling Technology (MA, USA). Mouse monoclonal anti-PSMC3 (1:500), anti-PEX19 (1:1000), rabbit polyclonal anti-STRAP (AB1, 1:2000), anti-EIF3I (1:500), anti-SFRS1, antiDDAH1 (1:1000), anti-HSPB1 (AB-82; 1:1000), anti-UBA1 (N-terminal; 1:1000), antiDCTN2 (1:1000), and goat polyclonal anti-PPP2CA/PPP2CB (1:2500) were obtained from Sigma-Aldrich (MO, USA). Goat polyclonal anti-tropomyosin (E-16, 1:200), anti-hnRNPC1/C2 (N-16, 1:200), anti-tropomyosin (E-16, 1:200), and anti-NACA (N-14, 1:200) were obtained from Santa Cruz Biotechnology (CA, USA). Mouse monoclonal anti-B23 (1:1000) was obtained from Millipore, and mouse monoclonal anti-EGFR (Ab-10, 1:250) was obtained from Thermo Scientific (MA, USA).
Analysis of Western Blotting Data
Independent of MS findings, western blot analysis on tissue specimens created new data sets. To correlate antibody-revealed quantitative change of candidate TDIs and CIT exposure, we generated degradation reference curves for individual candidates. Protein quantity was measured by using the densitometry data of the signals on western blot films via histogram analysis. Signals from candidates’ TDIs or from a degradation-unaffected protein were used as internal controls. The quantitative ratio between a TDI’s signal, and the signal from its internal control and its corresponding CIT length were used to generate antibody-based tissue degradation reference curve. Prism’s (GraphPad Software, Inc.) built-in statistical functions were used for construction of trend lines and calculation of statistical parameters.
RESULTS
Tissue Degradation Frequently Occurs Before Surgical Specimens Reach Appropriate Storage Condition
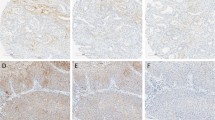
Variable degrees of degradation occur in human specimens during the interval from surgical resection to chemical fixation or snap-freezing. Here we provide additional exemplary evidence that tissue degradation affects the diagnostic features in specimens used for this study. By histology, significant morphological changes occur during tissue degradation, including altered intensity of nuclear staining and loss of cell border definition (Figure 1a). To observe the impact of tissue degradation on commonly used diagnostic markers, we investigated alteration of EGFR and p53 quantity during specimen degradation. EGFR was detected at CIT time point 0, but absent at any consecutive time points in a meningioma specimen (Figure 1b). Continuous decrease of p53 was also observed in cells that underwent the CIT degradation model throughout the observation time period (0 to 120 h; Figure 1c).
Impact of tissue degradation on diagnostic features of surgical specimens (numeric values indicate cold ischemic time (CIT) hours). One intracranial meningioma specimen with homogeneous histology was divided into four equal-sized fractions. Tissue fractions were randomly assigned to 0-, 4-, 24-, and 36-h CIT exposure, followed by OCT embedding to form a frozen tissue micro-array (TMA). (a) H&E staining, × 20. Decrease of histological quality at time points 0 h (left upper), 4 h (right upper), 24 h (left lower), and 36 h (right lower), including reduction of nuclear staining and loss of distinct cytoplasmic borders. A low power image of the respective tumor sample is demonstrated in insets showing the entire TMA section with respective tumor sample in green frame. (b) Tissues collected from individual pieces of the TMA were subjected to western blotting analysis using anti-EGFR antibody. A 170-kDa EGFR expression was detected at time point 0, but undetectable at later time points; lower panel: beta-actin for loading control. (c) 293T cell line was used to observe p53 changes during tissue degradation. Western blot analysis exhibits significant p53 degradation during CIT exposure; lower panel: beta-actin for loading control.
Cell Degradation Levels Quantitatively Correlate with the Degradation Status of a Subset of Proteins
We assigned four cell lines to a CIT-dependent tissue degradation experimental model and utilized a 2D DIGE-based proteomic approach to determine protein changes between cells at different CIT exposure (Figure 2a). For each cell line, six levels of CIT exposure were conducted and evaluated on twelve 2D gel images (experiments were performed in duplicate). In average, 2431 of 2798 protein spots (86.9%) showed no significant (<1.5-fold) quantitative changes during 48 h of CIT exposure. Fulfilling our criteria of candidate selection, a total of 50 protein spots had a continuous loss or increase during CIT in all four cell lines and were selected for MS sequencing (Figures 2b–e).
Two-dimensional fluorescence difference gel electrophoresis (2D DIGE) proteomic profiling and comparison. (a) One of the twelve 2D DIGE images obtained from Typhoon scanner using 293T cells (cold ischemic time (CIT)=0 h). Red-framed region of all 12 experimental groups is demonstrated in high resolution with DeCyder software (b and c). (b) After image alignment, the same protein spot is pink-circled in all 12 gels (gridded windows). (c) A neighboring protein spot is highlighted in 12 gels. (d) Illustration of gel layout in b and c. Quantitative decrease of protein spot in b and quantitative accumulation of protein spot in c are observed during CIT exposure. (e) A reciprocal change between protein spot in b (highlighted peak) and protein spot in c (right most peak) is illustrated in a three-dimensional format.
Identification of TDIs
Candidate protein spots from 2D gel analysis were subjected to both MALDI- and LC-based peptide sequencing. Mass spectrometric findings from both platforms are presented in Table 2. Twenty-six proteins of interest were successfully sequenced by both platforms. Of these proteins, 19 had shown continuous quantity loss during cell degradation, whereas 7 had shown continuous quantity accumulation; the additional prominent spot that had remained quantitatively stable at all time points was identified as beta-actin (no. 20, ACTB). In six instances, two or more separate gel spots were identified as parts of the same protein (no. 1 TDI: AHNAK, nucleoprotein; no. 2 TDI: SPTAN1, alpha-II spectrin; no. 3 TDI: EEF2, eukaryotic translation elongation factor 2, no. 8 TDI: EIF3I, eukaryotic translation initiation factor 3; no. 17 TDI: TPM3, tropomyosin 3; and no. 19 TDI: HSPB1, heat shock 27 kDa protein 1), and different spots of the same protein sometimes revealed reciprocal intensity changes with increasing CIT. Reciprocal intensity changes were observed in protein spots of either different isoelectric point (eg, no. 3 EEF2) or different molecular size (eg, no. 2 alpha-II spectrin). Computerized STRING analysis revealed the protein cohort of TDI candidates to mainly function as protein binding elements and/or structural constituents of the cytoskeleton (statistical significance P<0.01).
Validation of TDIs in Tissue Specimens
Western blot analysis was selected for TDI validation using 64 clinical human specimens and 12 mouse specimens. As degradation-induced protein changes included intact/breakdown conversions (isoelectric changes or molecular weight changes) in some cases, we applied both 1D and 2D western blot analysis to monitor these potential alterations.
The 2D western blot analysis successfully validated proteomic findings on isoelectric point (PI)-based TDI degradation for 11 proteins, EEF2 (no. 3), EIF3I (no. 8), PPP2C1 (no. 9), NPM1P21 (no. 13), HNRNPC (no. 14), SFRS1 (no. 15), DDAH1 (no. 16), TPM3 (no. 17), NACA (no. 18), PSMC3 (no. 24), and DCTN2 (no. 27); Table 2). Figure 3 shows successful validation of degradation-induced PI changes of EEF2; 2D western blotting exhibits a dynamic conversion from EEF2’s original form to a degraded form beginning at 0.5 h of CIT (Figure 3a). The quantitative ratio between the two spots strongly correlates with respective CIT impact (Figure 3c); when the same protein samples are evaluated by 1D western blot analysis alone, the CIT impact on EEF2 cannot be detected (Figures 3b and d). The protein PPP2C1 (no. 9 TDI) also exhibits a dynamic conversion between its two forms with different PIs (indicated with blue and red arrows in Figure 3e). The quantitative ratio between these forms strongly correlates with CIT impact with a R2-value of 0.98 for the logarithmical curve (Figure 3f).
Two-dimensional (2D) western blotting demonstrates proteins’ isoelectric point (PI) changes during tissue degradation. (a) Two EEF2 (eukaryotic translation elongation factor 2) isoforms show reciprocal changes during tissue degradation (cold ischemic time (CIT) hours are labeled on the left). Quantitative ratio between the left spot (left arrow) and the right spot (right arrow) is associated with respective CIT impact, as indicated by the logarithmical trend line in c. In contrast, b and d, using identical cell lysate under identical experimental condition, one-dimensional western blot is not able to distinguish EEF2 changes during tissue degradation. (e and f) A reciprocal change of the two isoforms (spot A and B, indicated with left and right arrow, respectively) of protein PPP2C1 is observed during tissue degradation. The logarithmical trend line indicates the association between the quantitative ratio of PPP2C1 isoforms and the CIT impact.
Quantitative changes of TDIs can be validated using 1D western blotting. The 1D western blotting results can be summarized as follows: (1) candidate TDIs were represented by correct molecular size and showed expected decrease of expression intensity with increased CIT exposure. These proteins included GSN (no. 4), VIM (no. 5), SET (no. 10), PEX (no. 11), TPM1 (no. 12), HSPB1 (no. 19), and B23 (no. 13, Figure 4a); (2) some antibodies could not validate candidate TDIs at proper molecular size and possible explanations include lack of specificity of applied antibodies or presence of TDI breakdown products. These proteins include AHNAK (no. 1, Figure 4b), NUP37 (no. 7), ATXN10 (no. 22), TUBA1B (no. 23), PDIA6 (no. 25), and PSMC4 no. 26); (3) candidate TDIs with quantitatively observed continuous increase during CIT exposure (Figure 4c; (4) beta-actin (no. 20) was successfully validated by 1D western blot analysis as an unaffected protein during CIT exposure (Figure 4d); (5) one protein (no. 2, alpha-II spectrin) revealed consistent quantitative decline of its native (intact) form and consistent quantitative increase of its breakdown product.
One-dimensional western blotting demonstrates protein quantity change during degradation (numeric values indicate cold ischemic time (CIT) hours). (a) Quantity of B23 protein decreases during CIT exposure in three meningioma specimens (tumor 1: 1 and 2 lane, tumor 2: 3 and 4 lanes, and tumor 3: 5 and 6 lanes). (b) Commercially available anti-AHNAK antibody cannot yield a discrete immune-signal of this super-sized protein (630 kDa) in a meningioma specimen, but reacts with protein products of smaller molecular size (likely AHNAK breakdown products). A relatively intact pattern of AHNAK can still be observed in early CIT exposure, although this intact pattern fades in later stages of tissue degradation. (c) Western blot analysis demonstrates an accumulation of ubiquitin-activating enzyme E1 (UBA1) at its proper 110-kDa size during 293T cell degradation. (d) Beta-actin is strongly and continuously expressed at all time points observed.
The Quantitative Ratio of Intact Alpha-II Spectrin and its Breakdown Products: a Better Tool for Degradation Assessment
By 1D western blot analysis, alpha-II spectrin exhibits a continuous and dynamic conversion between its intact form and its breakdown form during tissue degradation. Several commercially available spectrin-antibodies, as well as antibodies generated in our lab (data not shown), were used to simultaneously examine the quantity of the intact spectrin and its breakdown products (Figure 5). Intact spectrin and its breakdown product revealed continuous reciprocal change during CIT exposure in human surgical specimens (Figures 6a–c) and in mouse tissues of varied types (Figures 6d–g). An exponential correlation (y=aebx, a=2.897±0.07435 (0.9834∼3.158), b=−0.05076±0.002458 (−0.3005∼−0.01646)) was detected between spectrin breakdown and CIT stage in specimens that have undergone CIT degradation experiments (Figure 6h).
Dynamic conversion between spectrin’s intact and breakdown forms is confirmed in multiple tissues of varied origins via western analysis (cold ischemic time (CIT) hours are labeled on top of each lane) (a) in a non-tumorous human kidney specimen; (b) in three human meningioma specimens (tumor 1: 1 and 2 lane, tumor 2: 3 and 4 lanes, and tumor 3: 5 and 6 lanes); (c) in a human uterine leiomyoma specimen; (d–g) mouse uterus, intestine, lung, and bladder tissues, respectively; (h) an exponential trend line indicates the overall association between the spectrin intact/breakdown ratio and the CIT impact in all tissues observed.
Exploration of Spectrin-Based Degradation Assessment with Autopsy and FFPE Tissues
Compared with surgical specimens, autopsy specimens degrade in a different environment. A PMI elapses before tissue samples can be chemically fixed or frozen. Encouraged by the strong correlation between spectrin breakdown and CIT in surgical specimens, we tested for correlation between spectrin breakdown and PMI. We detected an association between spectrin breakdown and respective PMI of specimens using autopsy brain tissues (Figures 7a and b). These data highlight the potential of TDI application to autopsy tissue quality assessment and to studies in forensic medicine.
Exploration of using spectrin intact/breakdown ratio as a tissue degradation indicator (TDI) for autopsy tissues and formalin-fixed paraffin-embedded (FFPE) tissues. (a) Western blot. Hours of post mortem intervals (PMIs) are labeled on top of each lane. (b) The association between spectrin breakdown ratio and PMI is indicated by an exponential trend line. (c) Tissue fractions of a uterine leiomyoma specimen were exposed to assigned cold ischemic times (CITs) followed by 12-h formalin fixation and routine paraffin embedding. Western analysis shows that spectrin is difficult to be detected as clear immuno signals in FFPE tissues, compared with the frozen tissue at lane one (c, upper panel); low molecular weight beta-actin can be detected using the same blot (c, lower panel). However, using the vague immune signals at expected spectrin sizes (arrowheads indicate the intact spectrins and arrows indicate spectrin breakdowns), we can still determine the correlation between spectrin breakdown ratio and the CIT impact in these FFPE tissues, as illustrated with exponential trend line (d).
In analogy to frozen specimens, varied degrees of degradation may have occurred in FFPE tissues before formalin fixation. Pre-fixation degradation in FFPE specimens may affect biochemical integrity of specimens. To reveal pre-fixation degradation, we assigned fresh surgical tissues into experimental degradation models and conducted formalin fixation and paraffinization. In general, the efficiency of protein extraction from FFPE tissue was low, and the protein quality was poor because of formalin fixation and paraffinization. Higher molecular weight bands, such as spectrin, were displayed as vague bands (Figures 7c and d), whereas lower molecular weight proteins, such as beta-actin, remained better preserved and could still be revealed as discrete bands of acceptable quality. Although the signal of spectrin breakdown is not optimal, we were able to extract quantitative information about spectrin breakdown by quantifying the total signals at expected molecular sizes from western blot and determined the association between spectrin breakdown and CIT impact in FFPE tissues.
DISCUSSION
In agreement with many previous studies, we have demonstrated tissue degradation to occur in surgical pathology specimens immediately after tissue resection. Tissue degradation may alter the results of morphological evaluation and biomolecule examination and misguide patient care in severe instances.19, 21, 22, 23, 24, 25, 26 Therefore, degradation assessment on clinical specimens is critical to maximize confidence in clinical data interpretation.
To identify candidate proteins for degradation assessment, we conducted gel-based proteomic analysis on cultured cells exposed to different CITs. Our approach allowed for identification of approximately 3000 protein spots only, and most proteins of the cellular proteome remained undetected. However, this approach allowed for reliable identification of the most highly expressed proteins that are particularly attractive candidates for universal degradation indicators across many tissue types. Furthermore, it allowed for simultaneous observation of proteins and their breakdown products. To overcome heterogeneity issues of tissue specimens, we chose to use cultured cells of varied types for initial TDI discovery, followed by validation of candidate TDIs in human and mouse tissues.
After selection and excision of candidate protein spots from 2D gels, consensus IDs were identified for 26 by both LC/MS and MALDI/TOF (Table 2). Most of these proteins are highly expressed and known to participate in degradation-related cellular events, such as membrane repair (no. 1 AHNAK),27, 28 membrane/skeleton stabilization (no. 2 SPTAN1, no. 4 GSN, no. 5 VIM, no. 12 TPM1, no. 17 TPM3, no. 20 ACTB, no. 23 TUBA1B, no. 27 DCTN2),29, 30, 31, 32, 33, 34, 35 RNA/protein synthesis (no. 3 EEF2, no. 8 EIF3I, no. 13 B23, no. 14 NHRNPC, no. 15 SFRS1, no. 18 NACA, no. 25 PDIA6),36, 37, 38, 39, 40, 41, 42, 43, 44 cell death (no. 6 STRAP, no. 9 PPP2CA, no. 10 SET),45, 46, 47 chromosome segregation (no. 7 NUP37),48 free radical detoxification (no. 11 PEX19),49 and cell protection (no. 16 DDAH1, no. 19 HSPB1).50, 51 In addition, identified were key elements in the ubiquitin–proteasome pathway, including ubiquitin-like modifier activating enzyme 1 (no. 21 UBA1), which catalyzes the first step in ubiquitin conjugation to mark cellular proteins for degradation, and two ATPases (no. 24 PSMC3, no. 26 PSMC4) on the 26S proteasome, which cleaves peptides in an ATP/ubiquitin-dependent process.52, 53, 54
Proteomic findings in cells were validated by western analysis of an independent cohort of tissue specimens (Figures 3 and 4). These experiments confirmed the degradation-associated change of TDI candidates, which is characterized by quantitative loss, accumulation of degradation products, and/or altered isoelectric points. Of particular interest, we were able to utilize the quantitative ratio between intact TDIs and their respective breakdown form(s) for degradation assessment. We found establishment of intensity ratios between intact protein and breakdown forms(s) to be a particularly precise and sensitive way for degradation measurement. Further exploration of ratio alterations during CIT may be attractive as measurements can be obtained without reference to any other proteins.
Among the identified TDIs, alpha-II spectrin exhibited particularly stable and reproducible kinetics of intact-breakdown conversion within 48 h of tissue degradation, which can be considered as the most representative and relevant time frame for surgical and autopsy tissues (Figures 5 and 6). Analysis of 64 tissue specimens revealed a strong association between spectrin degradation and the impact of CIT that ranged from 0.81 to 0.96 (R2-value). Alpha-II spectrin is a 285 kDa scaffolding protein abundant in most cells. It forms the spectrin heterodimer with any of the five β-spectrins to carry out a wide variety of cellular functions, such as formation of plasma membrane, maintenance of cell shape,55, 56 and participation in signal transduction and molecular trafficking.57, 58 Calpains and caspases mediate alpha-II spectrin proteolysis under a number of pathological conditions in cells, including necrosis, apoptosis, traumatic damage, and cytotoxicity.59, 60, 61, 62, 63, 64 The initial calpain- and caspase-mediated cleavages yield four breakdown products at about 145–150 kDa.61, 65, 66 By applying a calpain inhibitor to our degradation model, we have successfully blocked degradation-induced spectrin breakdown, and by applying caspase cleavage-specific antibody (Asp1185) in western blotting, we did not detect the degradation-induced spectrin breakdown products (data not shown). These data suggest that the degradation-induced spectrin breakdown is calpain-mediated.
To further explore the usage of TDIs in tissue quality assessment, we examined the correlation between spectrin cleavage ratio and the degradation stage of autopsy tissues in reference to clinically documented PMIs (Figures 7a and b). In spite of numerous unknown contributing variables, such as circumstances of death, ambient conditions, as well as the accuracy of PMI determination, we were able to detect a correlation between spectrin cleavage and PMI.
Another attractive and important application of TDIs may be the evaluation of FFPE tissues. We examined degradation-induced spectrin cleavage in FFPE specimens of different pre-fixation CIT exposures (Figures 7c and d). In western blot analysis, although we detected accumulation of immunoreactive signal for spectrin at predicted sizes, we failed to detect clear bands due to formalin fixation. Nevertheless, comparative analysis of the intensity of intact spectrin with the intensity of its breakdown product demonstrated a correlation between spectrin degradation and length of CIT exposed in FFPE specimens. However, gel-based approaches may need to be complemented by other immunoassay strategies, such as ELISA and immunohistochemical staining, which do not require accurate protein separation.
Our identification of universal TDIs may yield a transformative platform dedicated to specimen quality control. This effort addresses critical, yet unmet needs for developing a universal standard for specimen degradation measurement. With this standard, specimen users will be able to define/scale the stage of decay in any specimens. An obvious and direct advantage of our findings is more informed comparability of data obtained from human biospecimens. In particular, if degradation-sensitive molecules are investigated, specimens with equal degradation stage should be considered.
Furthermore, TDI-based specimen assessment may also allow for extrapolation of the original integrity status (at specimen resection) of biomolecules of interest: individual specimen users may measure the quantity of their molecules of interest at multiple time points in multiple specimens of a certain tissue type; they may then correlate the quantity of the molecules with the respective degradation stages (scaled by our degradation indicators) and thereby conclude the degradation speed of the molecules of interest. For improved experimental analysis of new specimens, the users will be able to retrospectively recover the original quantity of the molecules of interest by adding/subtracting the degradation impact to the observed measurement via computing the observed quantity of the molecules, the degradation speed, and the corresponding degradation stage.
References
Mann GB, Fahey VD, Feleppa F et al. Reliance on hormone receptor assays of surgical specimens may compromise outcome in patients with breast cancer. J Clin Oncol 2005;23:5148–5154.
Chang JW, Chou CL, Huang SF et al. Erlotinib response of EGFR-mutant gefitinib-resistant non-small-cell lung cancer. Lung Cancer 2007;58:414–417.
Loi S, Symmans WF, Bartlett JM et al. Proposals for uniform collection of biospecimens from neoadjuvant breast cancer clinical trials: timing and specimen types. Lancet Oncol 2012;12:1162–1168.
Anagnostou VK, Welsh AW, Giltnane JM et al. Analytic variability in immunohistochemistry biomarker studies. Cancer Epidemiol Biomarkers Prev 2010;19:982–991.
Bai Y, Tolles J, Cheng H et al. Quantitative assessment shows loss of antigenic epitopes as a function of pre-analytic variables. Lab Invest 2011;91:1253–1261.
Tolles J, Bai Y, Baquero M et al. Optimal tumor sampling for immunostaining of biomarkers in breast carcinoma. Breast Cancer Res 2011;13:R51.
Zhou C, Byard RW . Factors and processes causing accelerated decomposition in human cadavers—An overview. J Forensic Leg Med 2011;18:6–9.
Mann RW, Bass WM, Meadows L . Time since death and decomposition of the human body: variables and observations in case and experimental field studies. J Forensic Sci 1990;35:103–111.
Vass AA, Barshick SA, Sega G et al. Decomposition chemistry of human remains: a new methodology for determining the postmortem interval. J Forensic Sci 2002;47:542–553.
Lee Goff M . Early post-mortem changes and stages of decomposition in exposed cadavers. Exp Appl Acarol 2009;49:21–36.
Szathmary SC, von Tamaska L, Steigel A . [Postmortem decomposition of neutral lipids. Use of modern methods of analysis (HPLC, capillary GC, GC–MS and NMR) in adipocere formation]. Z Rechtsmed 1985;94:273–287.
Moreno LI, Mills D, Fetscher J et al. The application of amplicon length heterogeneity PCR (LH-PCR) for monitoring the dynamics of soil microbial communities associated with cadaver decomposition. J Microbiol Methods 2011;84:388–393.
Srinivasan M, Sedmak D, Jewell S . Effect of fixatives and tissue processing on the content and integrity of nucleic acids. Am J Pathol 2002;161:1961–1971.
Espina V, Edmiston KH, Heiby M et al. A portrait of tissue phosphoprotein stability in the clinical tissue procurement process. Mol Cell Proteomics 2008;7:1998–2018.
Jung K, Lein M, Brux B et al. Different stability of free and complexed prostate-specific antigen in serum in relation to specimen handling and storage conditions. Clin Chem Lab Med 2000;38:1271–1275.
Tammen H . Specimen collection and handling: standardization of blood sample collection. Methods Mol Biol 2008;428:35–42.
Wolff AC, Hammond ME, Schwartz JN et al. American Society of Clinical Oncology/College of American Pathologists guideline recommendations for human epidermal growth factor receptor 2 testing in breast cancer. J Clin Oncol 2007;25:118–145.
Chin JL, Srigley J, Mayhew LA et al. Guideline for optimization of surgical and pathological quality performance for radical prostatectomy in prostate cancer management: evidentiary base. Can Urol Assoc J 2012;4:13–25.
Hammond ME, Hayes DF, Wolff AC . Clinical Notice for American Society of Clinical Oncology-College of American Pathologists guideline recommendations on ER/PgR and HER2 testing in breast cancer. J Clin Oncol 2012;29:e458.
Park NJ, Yu T, Nabili V et al. RNAprotect saliva: an optimal room- temperature stabilization reagent for the salivary transcriptome. Clin Chem 2006;52:2303–2304.
Pinhel IF, Macneill FA, Hills MJ et al. Extreme loss of immunoreactive p-Akt and p-Erk1/2 during routine fixation of primary breast cancer. Breast Cancer Res 2010;12:R76.
De Cecco L, Musella V, Veneroni S et al. Impact of biospecimens handling on biomarker research in breast cancer. BMC Cancer 2009;9:409.
Cappuzzo F, Magrini E, Ceresoli GL et al. Akt phosphorylation and gefitinib efficacy in patients with advanced non-small-cell lung cancer. J Natl Cancer Inst 2004;96:1133–1141.
Sauter G, Lee J, Bartlett JM et al. Guidelines for human epidermal growth factor receptor 2 testing: biologic and methodologic considerations. J Clin Oncol 2009;27:1323–1333.
Bartlett JM, Starczynski J, Atkey N et al. HER2 testing in the UK: recommendations for breast and gastric in-situ hybridisation methods. J Clin Pathol 2012;64:649–653.
Moatamed NA, Nanjangud G, Pucci R et al. Effect of ischemic time, fixation time, and fixative type on HER2/neu immunohistochemical and fluorescence in situ hybridization results in breast cancer. Am J Clin Pathol 2012;136:754–761.
Huang Y, Laval SH, van Remoortere A et al. AHNAK, a novel component of the dysferlin protein complex, redistributes to the cytoplasm with dysferlin during skeletal muscle regeneration. FASEB J 2007;21:732–742.
Lee IH, Lim HJ, Yoon S et al. Ahnak protein activates protein kinase C (PKC) through dissociation of the PKC-protein phosphatase 2A complex. J Biol Chem 2008;283:6312–6320.
De Matteis MA, Morrow JS . Spectrin tethers and mesh in the biosynthetic pathway. J Cell Sci 2000;113 (Pt 13):2331–2343.
Sun HQ, Yamamoto M, Mejillano M et al. Gelsolin, a multifunctional actin regulatory protein. J Biol Chem 1999;274:33179–33182.
Nekrasova OE, Mendez MG, Chernoivanenko IS et al. Vimentin intermediate filaments modulate the motility of mitochondria. Mol Biol Cell 2011;22:2282–2289.
Gourlay CW, Carpp LN, Timpson P et al. A role for the actin cytoskeleton in cell death and aging in yeast. J Cell Biol 2004;164:803–809.
Lawlor MW, Dechene ET, Roumm E et al. Mutations of tropomyosin 3 (TPM3) are common and associated with type 1 myofiber hypotrophy in congenital fiber type disproportion. Hum Mutat 2010;31:176–183.
Procaccio V, Salazar G, Ono S et al. A mutation of beta-actin that alters depolymerization dynamics is associated with autosomal dominant developmental malformations, deafness, and dystonia. Am J Hum Genet 2006;78:947–960.
Dutcher SK . The tubulin fraternity: alpha to eta. Curr Opin Cell Biol 2001;13:49–54.
Kaul G, Pattan G, Rafeequi T . Eukaryotic elongation factor-2 (eEF2): its regulation and peptide chain elongation. Cell Biochem Funct 2011;29:227–234.
Srivastava S, Verschoor A, Frank J . Eukaryotic initiation factor 3 does not prevent association through physical blockage of the ribosomal subunit-subunit interface. J Mol Biol 1992;226:301–304.
Okuda M . The role of nucleophosmin in centrosome duplication. Oncogene 2002;21:6170–6174.
Choi YD, Grabowski PJ, Sharp PA et al. Heterogeneous nuclear ribonucleoproteins: role in RNA splicing. Science 1986;231:1534–1539.
Gorlach M, Wittekind M, Beckman RA et al. Interaction of the RNA-binding domain of the hnRNP C proteins with RNA. EMBO J 1992;11:3289–3295.
Lin S, Fu XD . SR proteins and related factors in alternative splicing. Adv Exp Med Biol 2007;623:107–122.
Spreter T, Pech M, Beatrix B . The crystal structure of archaeal nascent polypeptide-associated complex (NAC) reveals a unique fold and the presence of a ubiquitin-associated domain. J Biol Chem 2005;280:15849–15854.
Kaiser BK, Yim D, Chow IT et al. Disulphide-isomerase-enabled shedding of tumour-associated NKG2D ligands. Nature 2007;447:482–486.
Echeverri CJ, Paschal BM, Vaughan KT et al. Molecular characterization of the 50-kD subunit of dynactin reveals function for the complex in chromosome alignment and spindle organization during mitosis. J Cell Biol 1996;132:617–633.
Poloz YO, O'Day DH . Determining time of death: temperature-dependent postmortem changes in calcineurin A, MARCKS, CaMKII, and protein phosphatase 2A in mouse. Int J Legal Med 2009;123:305–314.
Fan Z, Beresford PJ, Zhang D et al. HMG2 interacts with the nucleosome assembly protein SET and is a target of the cytotoxic T-lymphocyte protease granzyme A. Mol Cell Biol 2002;22:2810–2820.
Fan Z, Beresford PJ, Oh DY et al. Tumor suppressor NM23-H1 is a granzyme A-activated DNase during CTL-mediated apoptosis, and the nucleosome assembly protein SET is its inhibitor. Cell 2003;112:659–672.
Loiodice I, Alves A, Rabut G et al. The entire Nup107-160 complex, including three new members, is targeted as one entity to kinetochores in mitosis. Mol Biol Cell 2004;15:3333–3344.
Sacksteder KA, Jones JM, South ST et al. PEX19 binds multiple peroxisomal membrane proteins, is predominantly cytoplasmic, and is required for peroxisome membrane synthesis. J Cell Biol 2000;148:931–944.
Pope AJ, Druhan L, Guzman JE et al. Role of DDAH-1 in lipid peroxidation product-mediated inhibition of endothelial NO generation. Am J Physiol Cell Physiol 2007;293:C1679–C1686.
Bryantsev AL, Kurchashova SY, Golyshev SA et al. Regulation of stress-induced intracellular sorting and chaperone function of Hsp27 (HspB1) in mammalian cells. Biochem J 2007;407:407–417.
Ramser J, Ahearn ME, Lenski C et al. Rare missense and synonymous variants in UBE1 are associated with X-linked infantile spinal muscular atrophy. Am J Hum Genet 2008;82:188–193.
Ciechanover A . The ubiquitin-proteasome pathway: on protein death and cell life. EMBO J 1998;17:7151–7160.
Rivkin E, Kierszenbaum AL, Gil M et al. Rnf19a, a ubiquitin protein ligase, and Psmc3, a component of the 26S proteasome, tether to the acrosome membranes and the head-tail coupling apparatus during rat spermatid development. Dev Dyn 2009;238:1851–1861.
Bennett V, Baines AJ . Spectrin and ankyrin-based pathways: metazoan inventions for integrating cells into tissues. Physiol Rev 2001;81:1353–1392.
Kizhatil K, Yoon W, Mohler PJ et al. Ankyrin-G and beta2-spectrin collaborate in biogenesis of lateral membrane of human bronchial epithelial cells. J Biol Chem 2007;282:2029–2037.
Muresan V, Stankewich MC, Steffen W et al. Dynactin-dependent, dynein-driven vesicle transport in the absence of membrane proteins: a role for spectrin and acidic phospholipids. Mol Cell 2001;7:173–183.
Nicolas G, Fournier CM, Galand C et al. Tyrosine phosphorylation regulates alpha II spectrin cleavage by calpain. Mol Cell Biol 2002;22:3527–3536.
Cardali S, Maugeri R . Detection of alphaII-spectrin and breakdown products in humans after severe traumatic brain injury. J Neurosurg Sci 2006;50:25–31.
Knoblach SM, Alroy DA, Nikolaeva M et al. Caspase inhibitor z-DEVD-fmk attenuates calpain and necrotic cell death in vitro and after traumatic brain injury. J Cereb Blood Flow Metab 2004;24:1119–1132.
Zhang Z, Larner SF, Liu MC et al. Multiple alphaII-spectrin breakdown products distinguish calpain and caspase dominated necrotic and apoptotic cell death pathways. Apoptosis 2009;14:1289–1298.
Dutta S, Chiu YC, Probert AW et al. Selective release of calpain produced alphalI-spectrin (alpha-fodrin) breakdown products by acute neuronal cell death. Biol Chem 2002;383:785–791.
Newcomb JK, Pike BR, Zhao X et al. Concurrent assessment of calpain and caspase-3 activity by means of western blots of protease-specific spectrin breakdown products. Methods Mol Biol 2000;144:219–223.
Glantz SB, Cianci CD, Iyer R et al. Sequential degradation of alphaII and betaII spectrin by calpain in glutamate or maitotoxin-stimulated cells. Biochemistry 2007;46:502–513.
Wang KK, Posmantur R, Nath R et al. Simultaneous degradation of alphaII- and betaII-spectrin by caspase 3 (CPP32) in apoptotic cells. J Biol Chem 1998;273:22490–22497.
Nakajima E, Hammond KB, Rosales JL et al. Calpain, not caspase, is the causative protease for hypoxic damage in cultured monkey retinal cells. Invest Ophthalmol Vis Sci 2011;52:7059–7067.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Cell lines and human tissue specimens were exposed to an experimental cold ischemic time-dependent tissue degradation model. Using two-dimensional fluorescence gel electrophoresis in conjunction with mass spectrometry, tissue degradation indicators were identified for quantitative measurement of specimen degradation. These findings may allow for development of novel tests for quality control of human biospecimens.
Rights and permissions
About this article
Cite this article
Li, J., Kil, C., Considine, K. et al. Intrinsic indicators for specimen degradation. Lab Invest 93, 242–253 (2013). https://doi.org/10.1038/labinvest.2012.164
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2012.164