Abstract
Tumor initiation (TI) in xenotransplantation models of head and neck squamous cell carcinoma (HNSCC) is an inefficient process. Poor TI could be due to (1) posttransplant cell loss, (2) a rare sub-population of cancer stem cells or (3) a requirement for specific cellular interactions, which rely on cell number. By tracking GFP-expressing HNSCC cells, we conclude that the posttransplant loss of cancer cells is minimal in the xenotransplant model. Furthermore, an examination of putative cancer stem cell markers (such as CD133, CD44, SP and label retention) in HNSCC cell lines revealed no correlation between marker expression and tumorigenicity. In addition, single-cell clones randomly isolated from HNSCC cell lines and then transplanted into mice were all capable of initiating tumors with efficiencies varying almost 34-fold. As the observed variation in the clones was both more and less tumorigenic than the parental cells, a combination of two clones, at suboptimal cell numbers for TI, was implanted into mice and was found to modulate the tumor-initiating activity, thus indicating that TI is dependent on a ‘critical’ number of cells and, for the first time, that interactions between clonal variants within tumors can modulate the overall tumor-initiating activity. Put in context with previous literature on tumorigenic activity, we believe that interactions between clonal variants within a tumor as well as (1) stromal interactions, (2) angiogenic activity, (3) immunocompetence and (4) cancer stem cells may all contribute to tumorigenic potential and the propensity for tumor growth and recurrence.
Similar content being viewed by others
Main
Squamous cell carcinoma of the head and neck (HNSCC) is the sixth most common cancer globally and is associated with a mortality rate of 40–50% in the developed world.1 Surgical and radiation therapies are effective in removing localized disease, and the addition of chemotherapy helps to control disseminated disease and prevent relapse.2 However, despite evidence of complete clinical remission in many patients, tumor recurrence still occurs in approximately 1–3% of patients per year and disease recurrence and subsequent spread to local and distant sites is a frequent and fatal complication of HNSCC.2 To improve cure rates in HNSCC, it is important to develop therapies that can control tumor recurrence and metastasis.
An obligate step in tumor recurrence is the ability of residual cancer cells to re-establish a tumor at the primary or secondary site. Thus, to understand tumor recurrence, we need to understand the biological constraints involved in initiating tumors. The major biological constraint on tumor initiation (TI) is the inherent tumorigenic potential of tumor cells combined with microenvironmental constraints imposed on the tumor. The inherent ability of cells to initiate a tumor can be explained by two models, namely the clonal evolution model (stochastic model)3 and the cancer stem cell model.4 The clonal evolution model states that cancer cells are genetically unstable, leading to heterogeneity within tumors with respect to their genomes and their tumorigenic potential.3, 5 In addition, selective pressure placed on the tumor by the tumor–host environment can enrich cells with enhanced tumorigenic potential.3, 5 In contrast, the cancer stem cell model states that the ability to self-renew and to initiate tumors is the sole domain of a sub-population of tumor-initiating cells (referred to as cancer stem cells). The ability of these models to explain TI is the subject of considerable debate at present.6, 7, 8, 9, 10, 11, 12, 13 Identifying which of these models describe TI in HNSCC will lead to a better understanding of tumor recurrence and perhaps better targeted therapies.
In addition to transforming lesions acquired by tumor cells, another determinant of tumorigenic potential is the microenvironment. Microenvironmental constraints on tumor progression include stromal interactions,14, 15, 16 immunoediting17, 18, 19 and angiogenesis.15 The significant contribution by the immune system in regulating TI is clearly demonstrated in immunosuppressed patients (eg, transplant recipients) who are at a high risk of developing SCCs.20 Studies in immunocompetent patients/mice indicate that immune-directed tumor cell killing involves both the innate and the acquired immune system.17, 18 Interactions between cancer cells and their surrounding stroma have also been demonstrated to support and facilitate the progression of tumors.14, 15, 16 For example, it has been reported that progression from ductal carcinoma in situ to invasive carcinoma is enhanced by stromal elements, such as myofibroblasts.21 In addition, tumors with high growth rates have been shown to enhance the recruitment of mesenchymal stem cells into the tumor stroma22 and to stimulate neoangiogenesis.15, 23 Moreover, tumor growth/progression can be stimulated by macrophages.23 In contrast, there is also conflicting evidence that suggests that carcinoma-associated fibroblasts do not promote tumor progression.24 Taken together, the weight of evidence suggests that TI is a complex process that results from the combination of inherent cancer cell properties and microenvironmental interactions. Given the heterogeneity between tissue types (eg, mesenchyme vs bone marrow vs epithelia), it is likely that the relative contribution of cancer cells and the microenvironment will be tumor type specific, ie, the constraints on leukemic development and progression are likely to differ from those of the oral mucosae.
In this study, we examine the biological basis for TI of HNSCC cells in a murine xenotransplantation model. On the basis of stem cell marker analyses and clonal cell analyses, we conclude that the initiation of HNSCC in a xenotransplant model is most consistent with a stochastic model of TI. We examined clonal variants derived from HNSCC cell lines and found evidence that all variants retained an ability to initiate tumors. Significantly, we found that clonal variants differed in their transcript profiles, their TI activity and their histomorphology in vivo. We also found that the clonal variants localized as foci within the xenotransplants and this was confirmed in human tumors. In addition, we showed that an admixture of the clonal variants could significantly modulate TI activity. Thus we report, for the first time, that clonal variants within HNSCCs can modulate TI activity.
MATERIALS AND METHODS
Cell Culture, TI and Tumor Collection
All HNSCC cell lines were obtained from the ATCC and cultured as per ATCC recommendations (VA, USA). For TI studies, single-cell suspensions were prepared from subconfluent or confluent cultures and injected (subcutaneously, s.c.) into the neck scruff or along the lateral aspect of the back of NOD/SCID mice (ARC, Murdoch, Western Australia). Tumor size was monitored once a palpable mass was detected. Once tumors reached 0.8 cm3, they were collected and either stored in PBS for viable cell isolation or in 4% formaldehyde (Australian Biostain, VIC, Australia) for 24 h and then in 70% ethanol until prepared for sectioning and histopathological analyses.
Side Population Analysis
Side population analysis was estimated as described previously.25 The only alteration to this procedure was a preincubation of 30 min, with or without 100 μM Verapamil (Sigma-Aldrich, NSW, Australia), followed by incubation with 15 μM Hoechst 33342 for 60 min. The SP was analyzed on a Cytomation Moflo (Beckmann Coulter, CA, USA) equipped with a UV laser and high-speed four-way sorter and summit v.4.3 software (Dako, Denmark).
Label Retaining Cell Analysis
Single-cell suspensions were incubated with 1.5 μM CFDA (final concentration; Molecular Probes, OR, USA) in 1 ml PBS for 20 min at 37°C, and then 5 × the volume of media was added and cells were washed repeatedly in PBS. Cells were plated and kept away from any light source until they were trypsinized and analyzed on a FACSCanto 8 days later (BD Bioscience Pharmingen, FL1 channel).
CD44 and CD133 Expression Analysis
Expression levels for CD44-PE/APC (BD Bioscience, Australia) and CD133-APC (Miltenyi Biotec, NSW, Australia) were estimated by FACS as described previously described26 using FACSCanto. In both instances, an isotype control was included as a negative control.
Infection of Cell Lines with Lentivirus pLL 3.7 GFP Vector
A Detroit 562 cell line stably expressing GFP, was a generous gift from the McMillan Laboratory (Diamantina Institute, UQ, QLD, Australia) and was generated using a lentivirus containing the pLL 3.7 GFP vector (Addgene plasmid 11795) as described previously.27
In Vivo Tumor Imaging
Mice were anesthetized in the imaging chamber of a Kodak FX imaging system (Kodak, NSW, Australia) using isoflurane gas, an X-ray image was taken and GFP fluorescence was estimated using an excitation wavelength of 465 nm and an emission wavelength of 535 nm.
Single Clone Isolation and TI Assays
Single-cell suspensions were prepared as described previously,28 and then serially diluted in PBS. Cells were plated at a concentration of 1.5 cells per well in a 48-well plate (Corning Life Sciences, MA, USA). Cells were monitored and all wells containing more than a single colony were excluded. Cells were expanded then injected into mice within 2–5 passages of initial isolation from the cell line.
Microarray Analysis and Conformation of Gene Expression
RNA was collected from cells cultured in vitro and then purified using the RNeasy mini kit (Qiagen, MD, USA) as per the manufacturer's instruction. The quality and quantity of RNA was assessed using the Agilent 2100 BioAnalyser (Agilent, CA, USA). Only RNA with a reading of ≥7.5 was used. Biotinylated cRNA probes were then generated using an Illumina TotalPrep RNA amplification kit (Applied Biosystems, Scoresby, VIC, Australia). Probes were then hybridized to Illumina Human HT-12 BeadChip whole-genome arrays, as per the manufacturer's instructions. Data were then analyzed using R and Bioconductor (Applied Biosystems)29 and genes upregulated or downregulated threefold, or more, with a B value of ≥3 (ie, exceeding the 95% confidence interval) compared with the parental cell line were considered to be differentially expressed.
Cell and Tumor Immunohistochemistry
Cells were cultured on chamber slides (Lab-tek, Thermo Fisher, NY, USA) as described previously,28 then fixed in 4% formaldehyde for 10 min and stored in 70% ethanol. For chamber slide staining, cells were permeablized in PBS/0.2% Triton X-100 (Sigma-Aldrich) for 5 min and then washed twice in PBS before incubation with antibody. Tumor samples were paraffin embedded and 5 μm sections incubated in a heating oven (56°C) for 1 h before rehydration. Sections were rehydrated through an alcohol series before antigen retrieval (10 mM sodium citrate buffer in a microwave for 5 min on high). Sections were then allowed to rest at room temperature for 20 min, before immunohistochemistry. For immunohistochemistry, cells/tumor sections were incubated overnight with CEACAM6 (Biogenex, Australia) or MMP9 (R&D Systems, MN, USA) antibody (15 μg/ml) at 4°C. After this, sections were washed thoroughly in PBS and then incubated with secondary and tertiary antibodies (LSAB2 System HRP, Dako) as per the manufacturer's instructions. Visualization was performed after incubation with Cardassian DAB Chromagen (Applied Medical, QLD, Australia) as per the manufacturer's recommendations. All images were taken on a Zeiss axiovert 135 microscope with a × 20 (EC Plan-NEOFLUAR) or × 40 (Zeiss, LD achroplan) objective lens (AxioVision Release 4.7.2, Carl Zeiss Pty Ltd, Australia).
RESULTS
We examined the biological constraints on HNSCC recurrence using a xenotransplant model of HNSCC. Immunocompetence is an established determinant of SCC development and progression,19, 20 and thus to exclude this variable, we examined TI in the context of an immunocompromised mouse (NOD/SCID). TI of HNSCC cell lines injected s.c. in NOD/SCID mice required 104 to 3 × 105 cells to initiate tumors (Table 1). To exclude the possibility that inefficient TI was simply due to cell loss after implantation, we injected GFP-expressing Detroit 562 cells s.c. into NOD/SCID mice and measured fluorescence intensity over time until tumors reached 0.8 cm3 (Figure 1). The intensity and location of the fluorescence indicated that ∼60% of GFP positivity remained by day 2 after which GFP positivity increased with tumor growth (Figure 1a and b). These data indicate that the majority of cells injected survive (Figure 1a and b). This survival rate in vivo is similar to the ability of these cells to establish colonies in vitro (53% for Detroit 562 cells), suggesting that the loss of cells upon plating or injection may simply reflect the loss of viability after trypsinization. To determine whether humanizing the tumor microenvironment would enhance tumor-initiating activity, GFP-positive Detroit 562 cells were coinjected with human dermal fibroblasts or human epidermal keratinocytes (Figure 1c). These experiments indicated that neither tumor onset nor tumor growth was significantly altered by the presence of normal human stromal cells or keratinocytes (Figure 1c). These data indicate that the inefficient initiation of Detroit 562 cells in NOD/SCID mice cannot be attributed to posttransplant loss of cells or the lack of signals provided by human fibroblasts or keratinocytes.
Determining whether cell loss or coinjection with fibroblasts or keratinocytes alters tumor-initiating activity and tumor cell viability. (a) Detroit 562 cells stably expressing GFP were implanted subcutaneously in NOD/SCID mice and fluorescence was monitored over time (days). The site of injection is indicated with the arrow. Spurious fluorescence outside the injection site is due to autofluorescence of any remaining fur in the surrounding area. (b) Fluorescent intensity was measured in mice after injection of 107 GFP-expressing Detroit 562 cells. Initial fluorescence was arbitrarily set at 100%. Data are presented as mean±range of two independent experiments. (c) Detroit 562 cells were injected alone or in combination with normal human dermal fibroblasts (HDFs) or normal human epidermal keratinocytes (HEKs). The total monitoring period was 90 days. Data are presented as the number of mice bearing tumors/total number of mice injected.
Given that the majority of cells remain viable after implantation, we examined whether we could identify sub-populations of cells, within six HNSCC cell lines, that expressed markers or activities commonly associated with a stem cell-like phenotype. For example, expressions of CD4430 and CD13331 have been used previously to enrich tumor stem cells in HNSCCs and other solid tumors26, 30, 31 that possess stem cell-like activity. The presence of a side population32 or a slow cycling label retaining population of cells33 has been reported in normal keratinocyte stem cells and has also been reported in hematopoetic34, 35 and epithelial malignancies.36, 37 Therefore, we measured these markers in HNSCC cell lines (Figure 2). All cell lines expressed a SP, a CD44+ve sub-population and a label retaining sub-population of cells (Figure 2a). In addition, all cell lines, except SCC 9 and SCC15, contained a small CD133+ve sub-population of cells (Figure 2). These data indicate that cell lines contain sub-populations of cells that can be distinguished on the basis of their expression of markers previously associated with stem cell-like activity. However, there was no correlation between the relative abundance of cells expressing these markers and the minimum number of cells, of each cell line, required to initiate tumors (Figure 2b). In addition, there was no correlation between the expression of these markers and the time taken by the individual cell lines to form a palpable tumor (not shown). Taken together, these data suggest that the differential expression of these markers may be unrelated to tumor-initiating activity.
Expression of putative cancer stem cell markers in HNSCC cell lines. Cell surface marker expression (CD44 and CD133) and functional attributes (side population and label retention) of cancer stem cells were used to identify the sub-population of cells within the six HNSCC cell lines. (a) The percentage of cells expressing each of the markers (±s.e.m.) was determined in at least three independent experiments. (b) Plot of the correlation between the expression (expressed as a percentage of the total cell population) of each putative stem cell marker vs the minimum number of cells, of each cell line, sufficient to initiate a tumor. Each individual point represents data from an individual cell line. Data were derived from at least three independent experiments and are expressed as mean±s.e.m.
To formally test whether tumor-initiating activity resided in a sub-population of tumor-initiating cells or whether all cells retained an ability for self-renewal and TI, we randomly established single-cell clones from the most (Detroit 562), an intermediate (Cal 27) and the least tumorigenic cell lines (SCC15). Colonies were established from randomly isolated individual cells with 20–60% efficiency (ie, minimal selection pressure). Isolated clones were expanded and injected into mice within 2–5 passages. The tumor-initiating activity of the individual clones was then assessed in NOD/SCID mice (Table 2). All clones were capable of initiating tumors with varying efficiency (Table 2). For example, clone 1 of Detroit 562 cells was capable of forming tumors with 3 × 103 cells, whereas clone 5 required 105 cells to initiate a tumor (ie, 33-fold difference) and took the longest period of time to initiate a tumor (Table 2). These data indicate that the ability to initiate tumors is shared by most, if not all, cells within individual HNSCC cell lines and that the ability to initiate tumors can vary considerably between individual clones within a tumor. These data also indicate that clonal variants persist within established HNSCC cell lines.
The most tumorigenic clone of Detroit 562 cells (clone 1) or SCC15 (clone 2) were 3- and 10-fold more tumorigenic than their parental derivative, respectively (compare Table 1 with Table 2). As clones were established and injected within 2–5 passages, it is unlikely that there has been selection, thus raising the issue that individual clonal variants could interact with one another to modulate TI activity. We examined whether mixing Detroit clones 1 and 5 together at varying cell numbers could modulate the overall TI (Figure 3). Figure 3a indicates that 103 clone 1 cells or 3 × 104 clone 5 cells were unable to form tumors when injected alone (Figure 3a). However, 103 clone 1 and 3 × 104 clone 5 cells, when mixed, could initiate tumors (Figure 3a). These data indicate that interactions between clonal variants, within a tumor, have the capacity to modulate the overall TI activity. In addition, qualitative analysis of the histomorphology of tumors derived from parental cells revealed considerable heterotypia. In contrast, tumors derived from clone 1 and clone 5 were homogeneous within a tumor but differed significantly from one another, and parental cells, with respect to cell size, basophilia and the appearance of the invading stroma (Figure 3b). To examine these differences at a molecular level, and to quantitate the contribution of each clone to the tumor, we compared the transcriptomes of clones 1 and 5 and the parental cell line (GSE19994). The expression of CEACAM6 was highest in parental Detroit 562 cells (23.09-fold change) and in clone 1 (16.22-fold change) than in clone 5. In contrast, MMP9 was poorly expressed in parental Detroit 562 cells (−4.481-fold change) and in clone 1 (−4.534-fold change) when compared with clone 5. A detailed account of this analysis is being prepared for publication elsewhere (Cameron et al, in preparation). These data were confirmed immunohistochemically in vitro (Figure 4). CEACAM6 was most highly expressed in the majority of Detroit 562 cells, followed by expression in clone 1 with little evidence of staining in clone 5 cultures (Figure 4a). In contrast, MMP9 was expressed in only a few cells within cultures of parental Detroit 562 cells, not at all in clone 1, and was highly expressed in clone 5 in vitro (Figure 4a). These data, confirm that MMP9+ve and CEACAM6+ve clonal variants exist within parental Detroit 562 cell cultures, indicating that these clones were not generated through selection pressure incurred during the derivation of the clones. Similarly, CEACAM6 is abundantly expressed in discrete foci within Detroit 562-derived tumors and clone 1-derived tumors, whereas it is poorly expressed in clone 5-derived tumors (Figure 4b). Conversely, MMP9 is only present focally in Detroit 562-derived tumors and not at all in clone 1-derived tumors (Figure 4b). Clone 5-derived tumors displayed abundant MMP9 expression (Figure 4b). Significantly, tumors derived from an admixture of clone 1 and clone 5 resulted in CEACAM6+ve and MMP9+ve tumors (Figure 4b), indicating that both clone 1 and clone 5 cells contribute to the growth of tumors. In addition, examination of serial sections indicated that CEACAM6 and MMP9 were expressed in nonoverlapping foci within tumors derived from clone 1+clone 5 or tumors derived from parental Detroit 562 cells (Figure 4c). Finally, analysis of 4 human HNSCCs demonstrated that 4 of 4 and 3 of 4 HNSCCs were CEACAM6+ve or MMP9+ve, respectively. In all instances, CEACAM6 or MMP9 positivity occurred in nonoverlapping foci. This indicates that our analysis of the Detroit 562 clonal variants is relevant to human SCCs.
HNSCC clonal variants modulate each others’ tumor-initiating activity. An admix study was performed using varying numbers of the Detroit 562 cells or the most (clone 1) and least (clone5) tumorigenic clonal variants of the Detroit 562 cell line. (a) Tumor-initiating activity presented as numbers of mice bearing tumors/total number of mice injected. The average time (days) to onset of tumors is also shown. (b) Hematoxylin and eosin-stained histological sections of the Detroit 562 parental cell line, clone1, clone 5 and an admix of clone 1 (103 cells)+clone 5 (3 × 104 cells). The different morphology of cells and the tumor organization in the parental cell line (arrowheads) vs clonal variants must be noted. The differences in the structure of the stromal elements within clone 5 and clone 1 (arrowheads) must also be noted. Finally, the relative basophilia of clone 5 tumor cells vs clone 1 cells and the presence of lacunae-like structures (asterisks) within clone 5 tumors must be noted.
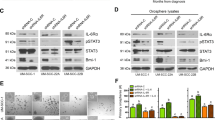
Expression of CEACAM6 or MMP9 in clonal variants in vitro and in vivo. Markers for clone 1 (CEACAM6) and clone 5 (MMP9) of the Detroit 562 parental cells were used to identify the different clones either in vitro or in vivo. (a) Detroit 562 parental cells or clone 1 or clone 5 variants were cultured on chamber slides and stained for CEACAM6 or MMP9 expression immunohistochemically ( × 40 magnification). A representative slide from at least three independent experiments is shown. (b) CEACAM6 and MMP9 expression was determined, immunohistochemically, in tumors derived from the Detroit 562 parental cell line, clone 1, clone 5, clone 1 (1000)+clone 5 (3 × 104) or in a patient oral SCC ( × 20 magnification). In some instances, focal expressions of CEACAM6 or MMP9 are highlighted by arrowheads. (c) Hematoxylin and eosin staining or CEACAM6 or MMP9 expression on serial sections of two regions of a tumor derived from clone 1+5 as described in panel b ( × 20 and × 40 magnification). Inset panels depict IgG-negative controls.
DISCUSSION
In this study, we used a xenotransplant model of HNSCC to identify biological determinants of TI. The data demonstrate that interactions between clonal cell variants within tumors can modulate the tumor-initiating activity of HNSCCs. This is significant for two reasons, first, these data show that tumors are composed of functionally distinct clonal variants that differ in their ability to initiate a tumor, differ in their gene signatures and differ in their histomorphology. As these differences are function based, this suggests that treatment strategies targeting HNSCC would need to target all functionally distinct variants within a tumor. Second, this study suggests that interactions between clonal variants within a tumor should be included in a growing list of factors that can modulate TI, such as the microenvironment, angiogenesis, the immune system and/or the presence of cancer cells with stem-like properties.
The presence of clonal variants within tumors is not novel. However, the demonstration that clonal variants vary in their inherent tumor-initiating activity and can modulate the inherent tumor-initiating activity of one another is novel. The existence of inherently tumorigenic clonal variants provides several key insights into TI. First, these data are consistent with the clonal evolution model of TI and inconsistent with the cancer stem cell model. In particular, random isolation of clonal variants indicated that TI (although variable) was a property shared by most, if not all, cells within the HNSCC tumor cell lines. Although we did find evidence for expression of putative cancer stem cell markers within the HNSCC cell lines, we found no correlation between their expression and the TI activity of the cell lines. In particular, CD44 has previously been reported to be enriched in HNSCC cancer stem cells;11 yet we found it to be abundantly expressed (>90% of cells expressing) in 3 of 6 cell lines, indicating that it is not associated with a sub-population of specialist tumor-initiating cells. Given that, TI required thousands of cells, this indicates that the presence of a CD44+ cell was not sufficient to initiate a tumor. Thus, the observation that randomly isolated individual clones gave rise to colonies that could initiate tumors indicates that TI occurs only if sufficient cells are present. However, this cell-number dependence is not dependent on a sub-population of tumor-initiating cells. Rather, TI activity is an attribute of most, if not all cells, within the cell lines. Moreover, we provide evidence that tumors and tumor cell lines were composed of clonal variants that differed in their inherent TI activity, histomorphology and gene signatures (eg, MMP9+ve CEACAM6−ve vs CEACAM6+ve MMP9−ve). This is consistent with a stochastic basis for TI and development as provided by the clonal evolution model.3 The observation that most, if not all, HNSCC cells can form tumors has not been reported for HNSCC but is similar to the findings of a recent study in melanoma.7 Although it could be argued that variant phenotypes reflect a hierarchy arising from an asymmetric division of cancer stem cells, the TI activity of the randomly isolated clones would argue against this. Similarly, it is possible that no single marker1, 11 may be sufficient to enrich stem cell-like tumor-initiating cells. However, this also remains unlikely given that randomly isolated cells give rise to colonies capable of initiating tumors in vivo.
The heterogeneity within HNSCC cell lines with respect to histomorphology and CEACAM6/MMP9 expression is significant because it indicates that functionally distinct populations are also genetically distinct. The identification of MMP9+ve and CEACAM6+ve clonal variants suggests that the inherent TI activity may be modulated by differential expression of genes involved in tumorigenesis or in adapting tumor cells to the microenvironment. Significantly, we were able to use MMP9 and CEACAM6 as ‘model’ probes for two clonal variants within the HNSCC cell population and show that the differential expression pattern was maintained in the parental cell line in vitro and in tumors derived from the parental cell line. More significantly, we found that human HNSCCs also displayed foci of CEACAM6+ve or MMP9+ve cells within the tumor. These data not only highlight the clinical relevance of our findings but also indicate that clonal variants remain in discrete foci within a tumor, which is consistent with them being clone-specific progeny. The presence of functionally distinct clonal variants also indicates that for therapies to be curative they will need to (1) target all clonal variants and (2) reduce tumor cell numbers below a ‘TI threshold’ (discussed below). The concept that significant variability in response to chemotherapeutics exists within tumor cell lines was recently shown in multiple tumor cell lines.38
Although it is clear that most tumor cells within the HNSCC cell lines retained TI activity, it is also clear that the process of TI is an inefficient one. Given that NOD/SCID mice lack B cells, T cells and NK cells coupled with the observation that HNSCC cells implanted in mice retain viable tumor-initiating activity, it is apparent that other factors exist that retard TI and growth and contribute to cell-number dependence for TI. This cell-number dependency could be considered a ‘tumor initiation threshold’ above which tumors can grow and below which tumors may not develop. Such a threshold is consistent with clinical data relating to micrometastasis, in which small foci (<2 mm) have been found in patients who have undergone treatment for breast, prostate and colon cancer.39 In these instances, the presence of micrometastases did not predict recurrence or the development of clinically relevant tumors, suggesting that tumor dormancy can occur.40 Numerous explanations for tumor dormancy have been proposed. For example, tumor dormancy can be explained by the immune editing hypothesis19, 41 or by an inability to stimulate neoangiogenesis.42, 43 Whatever the explanation, our observation that TI is cell number dependent is consistent with clinical observation and predicts that a critical number of cells may be required to alter the microenvironment from tumor suppressive to tumor permissive.
Cell number dependence relating to TI varies between individual clones and cell lines. This raises the question of what prevents TI in vivo and what is responsible for the variability between clonal variants and cell lines. We have excluded postimplant cell loss and the presence of a sub-population of tumor-initiating cells. It is also unlikely that it is simply the cell number per se44 as the addition of keratinocytes or fibroblasts does not facilitate TI. Moreover, as TI could be reproducibly enhanced simply by admixing different clonal variants, it would seem unlikely that TI, in the context of our xenotransplant model, requires further mutational events. One explanation is that below the TI threshold, there are microenvironmental pressures that are tumor suppressive. The microenvironmental constraints could be due to cell lines or clonal variants not making sufficient factors to promote tumor growth or neutralize tumor-suppressive factors present in the host environment. Alternatively, tumor suppression could be due to an inability to encourage angiogenesis or to promote stromal interactions/infiltration required to support or facilitate angiogenesis or tumor development. The importance of the stroma for the development and maintenance of a tumor phenotype has received considerable attention recently. For example, the use of antiangiogenic therapies to treat tumors or prevent recurrence/metastasis is well established.45 In addition, a recent study found that myofibrobalsts surrounding ductal carcinoma in situ of the breast could promote progression to carcinoma in situ.46 Moreover, Karnoub et al22 recently showed that infiltration of breast tumors by bone marrow-derived mesenchymal stem cells could promote breast cancer growth and metastasis. Thus, there are clear precedents for the contribution of the tumor stroma to tumor progression and growth. For this reason, we believe that the TI threshold, dictated by clonal cell variant interactions within the tumor, is likely to be linked to their ability to modify the stromal environment to make it tumor permissive.
References
Prince ME, Ailles LE . Cancer stem cells in head and neck squamous cell cancer. J Clin Oncol 2008;26:2871–2875.
Marur S, Forastiere AA . Head and neck cancer: changing epidemiology, diagnosis, and treatment. Mayo Clin Proc 2008;83:489–501.
Campbell LL, Polyak K . Breast tumor heterogeneity: cancer stem cells or clonal evolution? Cell Cycle 2007;6:2332–2338.
Wang JC, Dick JE . Cancer stem cells: lessons from leukemia. Trends Cell Biol 2005;15:494–501.
Nowell PC . The clonal evolution of tumor cell populations. Science 1976;194:23–28.
Kelly PN, Dakic A, Adams JM, et al. Tumor growth need not be driven by rare cancer stem cells. Science 2007;317:337.
Quintana E, Shackleton M, Sabel MS, et al. Efficient tumor formation by single human melanoma cells. Nature 2008;456:593–598.
Lapidot T, Sirard C, Vormoor J, et al. A cell initiating human acute myeloid leukaemia after transplantation into SCID mice. Nature 1994;367:645–648.
Al-Hajj M, Wicha MS, Benito-Hernandez A, et al. Prospective identification of tumorigenic breast cancer cells. Proc Natl Acad Sci USA 2003;100:3983–3988.
Singh SK, Hawkins C, Clarke ID, et al. Identification of human brain tumor initiating cells. Nature 2004;432:396–401.
Prince ME, Sivanandan R, Kaczorowski A, et al. Identification of a subpopulation of cells with cancer stem cell properties in head and neck squamous cell carcinoma. Proc Natl Acad Sci USA 2007;104:973–978.
Hill RP . Identifying cancer stem cells in solid tumors: case not proven. Cancer Res 2006;66:1891–1895; discussion 1890.
Kern SE, Shibata D . The fuzzy math of solid tumor stem cells: a perspective. Cancer Res 2007;67:8985–8988.
Mishra PJ, Humeniuk R, Medina DJ, et al. Carcinoma-associated fibroblast-like differentiation of human mesenchymal stem cells. Cancer Res 2008;68:4331–4339.
Orimo A, Gupta PB, Sgroi DC, et al. Stromal fibroblasts present in invasive human breast carcinomas promote tumor growth and angiogenesis through elevated SDF-1/CXCL12 secretion. Cell 2005;121:335–348.
Kuperwasser C, Chavarria T, Wu M, et al. Reconstruction of functionally normal and malignant human breast tissues in mice. Proc Natl Acad Sci USA 2004;101:4966–4971.
Shankaran V, Ikeda H, Bruce AT, et al. IFNgamma and lymphocytes prevent primary tumor development and shape tumor immunogenicity. Nature 2001;410:1107–1111.
Dunn GP, Old LJ, Schreiber RD . The three Es of cancer immunoediting. Annu Rev Immunol 2004;22:329–360.
Swann JB, Vesely MD, Silva A, et al. Demonstration of inflammation-induced cancer and cancer immunoediting during primary tumorigenesis. Proc Natl Acad Sci USA 2008;105:652–656.
Ulrich C, Schmook T, Sachse MM, et al. Comparative epidemiology and pathogenic factors for nonmelanoma skin cancer in organ transplant patients. Dermatol Surg 2004;30 (4 Part 2):622–627.
Allinen M, Beroukhim R, Cai L, et al. Molecular characterization of the tumor microenvironment in breast cancer. Cancer Cell 2004;6:17–32.
Karnoub AE, Dash AB, Vo AP, et al. Mesenchymal stem cells within tumor stroma promote breast cancer metastasis. Nature 2007;449:557–563.
Coffelt SB, Hughes R, Lewis CE . Tumor-associated macrophages: effectors of angiogenesis and tumor progression. Biochim Biophys Acta 2009;1796:11–18.
Qiu W, Hu M, Sridhar A, et al. No evidence of clonal somatic genetic alterations in cancer-associated fibroblasts from human breast and ovarian carcinomas. Nat Genet 2008;40:650–655.
Redvers RP, Li A, Kaur P . Side population in adult murine epidermis exhibits phenotypic and functional characteristics of keratinocyte stem cells. Proc Natl Acad Sci USA 2006;103:13168–13173.
Harper LJ, Piper K, Common J, et al. Stem cell patterns in cell lines derived from head and neck squamous cell carcinoma. J Oral Pathol Med 2007;36:594–603.
Gu W, Putral L, Hengst K, et al. Inhibition of cervical cancer cell growth in vitro and in vivo with lentiviral-vector delivered short hairpin RNA targeting human papillomavirus E6 and E7 oncogenes. Cancer Gene Ther 2006;13:1023–1032.
Dahler AL, Rickwood D, Guminski A, et al. Indole-3-carbinol-induced growth inhibition can be converted to a cytotoxic response in the presence of TPA+Ca(2+) in squamous cell carcinoma cell lines. FEBS Lett 2007;581:3839–3847.
Duan R, Leo P, Bradbury L, et al. Gene expression profiling reveals a down-regulation in immune-associated genes in AS patients. Ann Rheum Dis; published online 29 July 2009 (e-pub ahead of print).
Patrawala L, Calhoun T, Schneider-Broussard R, et al. Highly purified CD44+ prostate cancer cells from xenograft human tumors are enriched in tumorigenic and metastatic progenitor cells. Oncogene 2006;25:1696–1708.
Shepherd CJ, Rizzo S, Ledaki I, et al. Expression profiling of CD133(+) and CD133(−) epithelial cells from human prostate. Prostate 2008;68:1007–1024.
Terunuma A, Kapoor V, Yee C, et al. Stem cell activity of human side population and alpha6 integrin-bright keratinocytes defined by a quantitative in vivo assay. Stem Cells 2007;25:664–669.
Mackenzie IC, Bickenbach JR . Label-retaining keratinocytes and Langerhans cells in mouse epithelia. Cell Tissue Res 1985;242:551–556.
Moshaver B, van Rhenen A, Kelder A, et al. Identification of a small subpopulation of candidate leukemia-initiating cells in the side population of patients with acute myeloid leukemia. Stem Cells 2008;26:3059–3067.
Holyoake T, Jiang X, Eaves C, et al. Isolation of a highly quiescent subpopulation of primitive leukemic cells in chronic myeloid leukemia. Blood 1999;94:2056–2064.
Kabashima A, Higuchi H, Takaishi H, et al. Side population of pancreatic cancer cells predominates in TGF-beta-mediated epithelial to mesenchymal transition and invasion. Int J Cancer 2009;124:2771–2779.
Zhang HB, Ren CP, Yang XY, et al. Identification of label-retaining cells in nasopharyngeal epithelia and nasopharyngeal carcinoma tissues. Histochem Cell Biol 2007;127:347–354.
Gascoigne KE, Taylor SS . Cancer cells display profound intra- and inter-line variation following prolonged exposure to antimitotic drugs. Cancer Cell 2008;14:111–122.
Wikman H, Vessella R, Pantel K . Cancer micrometastasis and tumor dormancy. APMIS 2008;116:754–770.
Funke I, Schraut W . Meta-analyses of studies on bone marrow micrometastases: an independent prognostic impact remains to be substantiated. J Clin Oncol 1998;16:557–566.
Quesnel B . Tumor dormancy and immunoescape. APMIS 2008;116:685–694.
Achilles EG, Fernandez A, Allred EN, et al. Heterogeneity of angiogenic activity in a human liposarcoma: a proposed mechanism for ‘no take’ of human tumors in mice. J Natl Cancer Inst 2001;93:1075–1081.
Udagawa T . Tumor dormancy of primary and secondary cancers. APMIS 2008;116:615–628.
Revesz L . Effect of tumor cells killed by x-rays upon the growth of admixed viable cells. Nature 1956;178:1391–1392.
Grimm D, Bauer J, Schoenberger J . Blockade of neoangiogenesis, a new and promising technique to control the growth of malignant tumors and their metastases. Curr Vasc Pharmacol 2009;7:347–357.
Hu M, Yao J, Carroll DK, et al. Regulation of in situ to invasive breast carcinoma transition. Cancer Cell 2008;13:394–406.
Acknowledgements
This work was supported by a PhD scholarship awarded to SRC and a Principal Research Fellowship awarded to NAS by the Garnett Passe & Rodney Williams Memorial Foundation. This work was also supported by a PhD scholarship to KP and research grants awarded to NAS (nos 455929 and 569689) from the Australian NHMRC, Cancer Council QLD, no. 631479 and a Practitioner fellowship to AG from the Cancer Collaborative Group.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Cameron, S., Dahler, A., Endo-Munoz, L. et al. Tumor-initiating activity and tumor morphology of HNSCC is modulated by interactions between clonal variants within the tumor. Lab Invest 90, 1594–1603 (2010). https://doi.org/10.1038/labinvest.2010.131
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2010.131
Keywords
This article is cited by
-
Cancer stem cells in human digestive tract malignancies
Tumor Biology (2016)
-
Focal overexpression of CEACAM6 contributes to enhanced tumourigenesis in head and neck cancer via suppression of apoptosis
Molecular Cancer (2012)
-
Preclinical evaluation of dual PI3K-mTOR inhibitors and histone deacetylase inhibitors in head and neck squamous cell carcinoma
British Journal of Cancer (2012)
-
Dueling models in head and neck tumor formation
Laboratory Investigation (2010)