Abstract
Islet enumeration in impure preparations by conventional dithizone staining and visual counting is inaccurate and operator dependent. We examined nuclei counting for measuring the total number of cells in islet preparations, and we combined it with morphological analysis by light microscopy (LM) for estimating the volume fraction of islets in impure preparations. Cells and islets were disrupted with lysis solution and shear, and accuracy of counting successively diluted nuclei suspensions was verified with (1) visual counting in a hemocytometer after staining with crystal violet, and automatic counting by (2) aperture electrical resistance measurement and (3) flow cytometer measurement after staining with 7-aminoactinomycin-D. DNA content averaged 6.5 and 6.9 pg of DNA per cell for rat and human islets, respectively, in agreement with literature estimates. With pure rat islet preparations, precision improved with increasing counts, and samples with about ⩾160 islets provided a coefficient of variation of about 6%. Aliquots of human islet preparations were processed for LM analysis by stereological point counting. Total nuclei counts and islet volume fraction from LM analysis were combined to obtain the number of islet equivalents (IEs). Total number of IE by the standard method of dithizone staining/manual counting was overestimated by about 90% compared with LM/nuclei counting for 12 freshly isolated human islet research preparations. Nuclei counting combined with islet volume fraction measurements from LM is a novel method for achieving accurate islet enumeration.
Similar content being viewed by others
Main
Transplantation of islets of Langerhans is an alternative to whole pancreas transplantation for a subpopulation of patients with type 1 diabetes.1, 2, 3 Improvements in pancreas procurement, preservation, islet isolation, culture, and transplantation are being pursued,2, 4, 5, 6 but progress is hindered by the absence of meaningful measures of islet dose and quality,7, 8, 9, 10 the most fundamental of which is the amount of islet tissue in a preparation. Estimation of islet volume in a pure preparation is challenging and is even more difficult with human islet preparations that typically consist of about 50% nonislet components.11, 12, 13, 14
Early measurements of islet properties reported on a per islet basis were scattered.10 The concept of an islet equivalent (IE), the volume of a sphere with a diameter of 150 μm, standardized islet enumeration on the basis of volume and reduced variability.15, 16, 17 The current standard method involves staining islets with dithizone (diphenylthiocarbazone, DTZ),18 followed by light microscopy (LM) visualization and manual counting. Usually, one dimension is estimated, such as the diameter of a circle having the same area as the islet. Islet diameters are divided into groups of 50-μm increments, and conversion factors are used to estimate total islet volume.15 This oversimplified calculation leads to inaccuracy19, 20 because the volume is calculated assuming that the islet is a sphere, and use of a single conversion factor over a large increment adds additional error. Furthermore, measurements that differed by as much as 40% between different operators occurred in one unpublished multicenter study.21 Digital image analysis applied to replace visual estimation22, 23, 24, 25 is based on a two-dimensional area estimate and on the assumption that the islet is a sphere or ellipsoid of revolution, neither of which is true.26, 27, 28
We propose an alternative approach: separate measurements are made of (1) the total amount of tissue (eg, total volume or number of cells) and (2) the fraction that is islet tissue, from which the amount of islet tissue is calculated. Methods to measure total amount of tissue include the change in electrical resistance as a tissue particle passes through an aperture and assay of DNA content, total protein, and dry weight.10, 29, 30 Measurements of the packed cell volume (PCV) are routinely made with human islet preparations31 but have not been used to estimate the volume of tissue. There remains a need for a fast, accurate, and precise method for measuring the total amount of tissue in an islet preparation.
Nuclei counting is a standard method for accurate enumeration of individual cells,32, 33 and release of nuclei from cells osmotically lysed with hypotonic citric acid occurs without nuclei breakdown or losses. With addition of a surfactant, nuclei counts can be used to quantitatively assess cell number in suspensions of cell aggregates.34 In this study, we first examine nuclei counting following cell disruption for measuring the total number of cells in rat islet preparations and cell suspensions. Nuclei are quantitated by visual counting with a hemocytometer and with two automated methods, aperture electrical resistance measurement and flow cytometry. Comparison with DNA analysis demonstrates the superiority of nuclei counting. We use nuclei counting to determine the number of IE in pure rat islet preparations and compare the data with DTZ staining/manual counting. We then combine nuclei counting and LM analysis to determine the number of IE in freshly isolated human islet preparations. We showed in a previous study31 that LM morphological analysis of 1-μm plastic sections by stereological point counting provides estimates of the islet volume fraction in impure preparations with accuracy and precision comparable to that obtained by electron microscopic (EM) ultrastructural analysis of ultrathin sections. We compare the number of IE obtained by LM/nuclei counting to the standard method of DTZ staining/manual counting and to a combination of LM analysis and PCV measurements. These data demonstrate that the current standard DTZ staining method substantially overestimates the number of IE and that nuclei counting with LM analysis provide an attractive alternative.
MATERIALS AND METHODS
Culture of Cell Lines and Islet Preparations
Mouse insulinoma βTC3 cells35 were cultured in tissue culture flasks in DMEM medium (Mediatech, Herndon, VA, USA) that contained 4.5 g/l glucose supplemented with 10% (v/v) fetal bovine serum (FBS, Mediatech), 100 U/ml penicillin (Sigma-Aldrich, St Louis, MO, USA), 100 μg/ml streptomycin (Sigma), and 10 mM HEPES (Mediatech), in a humidified 5% CO2 incubator at 37°C. Rat insulinoma INS-1 cells36 were cultured in RPMI medium with 11.1 mM D-glucose (Mediatech) supplemented with 10% FBS, 100 U/ml penicillin, 100 μg/ml streptomycin, 10 mM HEPES, 2 mM L-glutamine (Mediatech), 1 mM sodium pyruvate (Mediatech), and 50 μM 2-mercaptoethanol (Sigma). Islets, isolated from male Sprague–Dawley rats using collagenase digestion/Ficoll purification,37 were cultured in Medium 199 (Invitrogen, Carlsbad, CA, USA) supplemented with 80 mg/l ciprofloxacin (Mediatech), 5 mM sodium pyruvate, 2 mM GlutaMax (Invitrogen), 16.7 μM zinc sulfate (EM Sciences, Darmstadt, Germany), 25 mM HEPES buffer, and 10 ml/l of ITS+Premix solution (BD Biosciences, Bedford, MA, USA). Human islet preparations were cultured in CMRL 1066 (Miami formulation, Mediatech) that contained 1 g/l glucose supplemented with 10% (v/v) FBS.
Human Islet Isolation
Islets were isolated as previously described31 from cadaver pancreata, some using the two-layer perfluorocarbon (PFC) method,38 by the Islet Resource Center of the Joslin Diabetes Center using standard collagenase/protease digestion.1, 39, 40 From the final islet suspension, aliquots were taken immediately for assays, including DTZ staining for volume fraction estimation and manual counting for islet enumeration, PCV measurements, LM morphological analysis, and nuclei counting. Two 0.5 ml aliquots for morphological analysis by LM were fixed immediately. In this study, aliquots for DNA analysis were taken from islet samples received about 4–6 h after isolation was complete and were immediately frozen for subsequent measurement. Samples examined in this study by LM analysis, DTZ staining, and PCV measurement were from 12 freshly isolated preparations deemed suitable only for research because they had insufficient number of IE for transplantation.
Cells and Nuclei for Counting
Cells from cell lines were removed from flasks by incubation with 0.05% (w/v) trypsin in 0.53 mM EDTA solution (Mediatech) at 37°C for 7 min (βTC3) or 3 min (INS-1). The resulting suspension containing single cells with occasional small aggregates was stored on ice. Cell counting was performed within 30–45 min after tryspinization to avoid clumping. Islet suspensions were mixed by vortexing, and aliquots were removed for nuclei preparation using narrow-orifice (0.6-mm inside diameter (id)) pipette tips (0–200 μl, USA Scientific, Ocala, FL, USA). Selected experiments were carried out with wide-orifice (2-mm id) (0–200 μl, Thermo Fisher Scientific, Waltham, MA, USA) and medium-orifice (1-mm id) (200–1000 μl, USA Scientific, Ocala, FL, USA) pipette tips. Nuclei were prepared from cells and islets by adding equal 100-μl volumes of sample and of a lysis solution containing 0.1 M citric acid (Sigma) and 1% (v/v) Triton X-100 (Sigma) to a 1.5 ml microtube. The mixture was incubated at room temperature for 5 min with vortex mixing every 1.5–2 min. The islet mixture was additionally placed in a 1-ml syringe and rapidly forced through a 26G3/8″ needle (Becton Dickinson, Franklin Lakes, NJ, USA) 10 consecutive times. Unless otherwise specified, 105 cells and 160 IE (2.5 × 105 cells assuming 1560 cells per IE36) were used for each sample analyzed.
Counting Methods
Visual counting of cells and nuclei with hemocytometer
A cell suspension volume of 200 μl was stained with 200 μl of 0.4% (v/v) trypan blue and diluted with Dulbecco's phosphate-buffered saline (D-PBS, Invitrogen) to between 5 and 15 × 105 cells per ml. A 12-μl sample was placed on a hemocytometer (Hausser Scientific, Horsham, PA, USA) and counted using × 40 magnification.41, 42 Stained and unstained cells were counted to obtain total cell number. For each of triplicate samples, 250 cells were counted on each side of the hemocytometer (total 500 cells). For counting nuclei, crystal violet (Sigma) was added to the lysed samples to a concentration of 0.1% (w/v). Unless otherwise specified, samples were diluted with D-PBS to between 5 and 15 × 105 nuclei per ml and counted as described for cells; 500 nuclei were counted for each of triplicate samples.
Aperture electrical resistance measurement
A Coulter Multisizer II (Beckman-Coulter, Miami, FL, USA) was used to determine particle size distribution by monitoring current between two electrodes on either side of a 100-μm diameter aperture that allowed detection of diameters from 2.0 to 60 μm. The instrument was calibrated with standardized beads (CC Size Standard L 43 certified NIST traceable latex beads; Beckman-Coulter). A known volume of nuclei sample was diluted with 40% glycerol in ISOTON II diluent (Beckman-Coulter) to a volume of 20 ml. A sample volume of 500 μl was drawn through the aperture, and the resulting current pulses were analyzed (AccuComp, Beckman-Coulter) to provide a volume distribution and the sample concentration.
Flow cytometer
Unless otherwise specified, nuclei were diluted with D-PBS to no higher than 5 × 105 nuclei per ml, stained with 7-aminoactinomycin D (7-AAD, Invitrogen) at a final concentration of 0.01 mg/ml for at least 2 min at room temperature, then analyzed using a flow cytometer (Guava Personal Cell Analysis (PCA) system, Guava Technologies, Hayward, CA, USA). Instrument performance was verified with the Guava Check application and Guava Check Kit reagents (Guava Technologies). 7-AAD fluorescence was detected in photomultiplier 2 (PM2) with PM1 and PM2 voltage set at 430 and 450 V, respectively. PM2 threshold was set between 70 and 100, forward scatter (FSC) gain at × 4, and FSC gate at 100. Nuclei exhibited FSC intensities >100, and debris (events with FSC intensity <100) were excluded. Scatter plots of 7-AAD intensity and FSC were initially viewed with Guava Express software. Data from 7-AAD-stained nuclei samples were acquired using the Guava ViaCount application. The number of nuclei events appeared in the ‘total cells per ml’ section of the data table, and 1000 events were acquired per sample in triplicate.
DNA Analysis
DNA concentration in cells or islets was quantified by fluorospectrophotometry using the CyQUANT Cell Proliferation Assay Kit (Invitrogen) and the λ-DNA standard (Invitrogen), which is based on the strong fluorescence enhancement that CyQUANT GR dye undergoes when bound to cellular nucleic acids. CyQUANT dye binding to RNA was eliminated by pretreating samples with DNAse-free RNAse (Sigma). Fluorescence intensity was measured with 480 nm excitation and 520 nm emission wavelengths in a plate reader (Spectra MAX Gemini micro-plate spectrophotometer, Molecular Devices, Sunnyvale, CA, USA) and was linearly related to the nucleic acid concentration in the sample. DNA was also quantified with the PicoGreen dsDNA Quantitation Kit (Invitrogen) for comparison. The procedure, λ-DNA standard, and lysis buffer were the same as for the CyQUANT assay; only the dye was different.
Islet Volume Fraction from LM Morphological Analysis
A 5 ml aliquot was processed to produce 1-μm plastic sections stained with toluidine blue that were analyzed at × 420 by stereological point counting43 with a 90-point grid covering adjacent, nonoverlapping fields, to determine the volume fraction of islet tissue, as previously described.31 The tissue type was determined at each intercept point over tissue.
Number of IE from Nuclei Counting Data and LM Morphology
The volume fraction of islet tissue obtained from LM morphological analysis by stereological point counting was combined with the total number of all cells measured by nuclei counting to yield an estimate of the number of IE in the preparation. The calculation procedure used Equations (1)–(5) as described in the Appendix. Briefly, the first step was conversion of the volume fraction islet tissue into the volume fraction of islet cells among all cells exclusive of extracellular volume (Equation (2)), and then into the number fraction of islet cells by using estimates of the volume of each cell type (Equation (3)). Multiplication by the total number of all cells yielded the number of islet cells (Equation (4)), which was converted to number of IE following division by 1560 cells per IE31 with Equation (5). The total volume of islets could be calculated once the number of IE was known.
Volume Fraction Islets and Number of IE by DTZ Staining
DTZ was used to discriminate islet from nonislet tissue by staining islet cells.18 Volume fraction islets was estimated by LM visualization, and the number of IE in a sample was determined by size estimation using a micrometer disc calibrated reticle and use of a standard formula,44 as previously described.31
Number of IE from PCV Measurement and LM Morphology
The PCV of the islet preparation was measured as previously described,31 from which tissue volume was determined by using an estimate of 0.345 for the void volume fraction of the packed cell pellet. The calculation procedure used Equations (6)–(9) in the Appendix. The volume of islet tissue was determined from the total tissue volume together with the volume fraction islets obtained directly from the LM morphological analysis. Finally, the volume of an IE was used to calculate the number of IE (NIE) in the preparation.
Statistics
Straight lines were fitted to data by linear least squares regression. Unless specified otherwise, precision was reported as s.d. for sample means and error bars and as the s.e. of the estimate for fitted slopes. The significance of differences in means and slopes of straight lines was evaluated by two-sided Student t tests.
Additional details of methods are available elsewhere.45
RESULTS
Comparison of Nuclei Counting Methods
Visual and graphical output
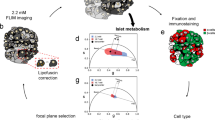
Cell disruption of high and low purity human islet preparations by osmotic lysis using hypotonic citric acid together with Triton X-100 surfactant and shear produced a suspension of individual nuclei; no clumping of two or more nuclei was observed (Figures 1a and d). Cell debris and fragments were more pronounced with islets than with cells and with low purity than with high purity preparations. Intact normal nuclei and a small fraction of nuclei shrunken during apoptosis, but not any smaller fragments, were included when counting with a hemocytometer. Aperture resistance measurements produced a size distribution with two, slightly overlapping populations that represented nuclei and smaller fragments (Figures 1b and e). The flow cytometer produced dot plots with two populations (Figures 1c and f). Only the nuclei displayed high 7-AAD fluorescence intensity, thereby permitting a clean separation of nuclei from cell fragments.
Output of each counting method for nuclei from human islets of high (about 80%, top row) and low (30%, bottom row) purity by DTZ staining). (a, d) Photomicrograph of visual image for counting nuclei from these islets. During counting with a hemocytometer, the image was defocused slightly to maximize contrast around the border of each nucleus. (b, e) Nuclei size distribution measured at a concentration of 2 × 104 nuclei/ml by the aperture resistance method (Coulter Multisizer II). For counting purposes, the position of the minimum at 3 μm is taken as the border separating larger nuclei from smaller fragments. The diameter registered by the aperture resistance measurement is smaller than the actual size (see panels a, d) because the nuclear envelope is permeable to ions,54 which carry electrical current. (c, f) Flow cytometer plot of 7-AAD fluorescence intensity vs FSC intensity using the Guava PCA system with stained nuclei. Nuclei are in the population of events with high FSC (>100) and high 7-AAD intensity (>100). Image was obtained with Guava Express software and 1000 events in the nuclei quadrant. Output of each counting method for nuclei from rat islets was essentially identical to that for high purity human islets. The higher density of incidents at low 7-AAD intensity (<100) in a, as compared with d, results from the relatively larger concentration of cellular fragments in the low purity preparation. The same trend is reflected in the size distribution of particles <3 μm in diameter from aperture resistance measurements in b, e.
Linearity, accuracy, and precision of enumeration measurements
INS-1 cells were suspended and disrupted to form a stock nuclei suspension. Aliquots diluted to cover a wide concentration range were counted by each method. Measured nuclei concentration is plotted in Figure 2a as a function of the calculated concentration over four orders of magnitude and in Figures 2b and c on linear coordinates over different concentration ranges to illustrate where each counting method departs from linearity. Concentration measured with a hemocytometer varied linearly with calculated concentration over the entire range but was uniformly about 11% higher than the line of identity (Figure 2c). Flow cytometry and aperture electrical resistance data agreed well with the line of identity up to about 5 × 105 and 7 × 107 nuclei per ml, respectively (Figure 2b), above which the data began to deviate from linearity. These deviations arose from coincident events (two nuclei registering in the sensing zone at the same time), which increased with concentration. The Coulter Multisizer II gave erroneously high readings at about <104 nuclei per ml (data not shown).
Measured vs calculated concentration of nuclei from INS-1 cells counted by all three methods. (a) All data plotted on log–log coordinates. (b) Data for concentrations up to 106 nuclei per ml plotted on linear coordinates. (c) Data for concentrations up to 2 × 105 nuclei/ml plotted on linear coordinates. Data for visual counting with a hemocytometer were fitted with a straight line through the origin and yielded a slope of 1.11±0.03. To determine calculated concentration, each measured concentration was multiplied by its dilution factor to estimate the original stock solution concentration. The average of these estimates in the range of 1–5 × 105 (flow cytometry) and 2–7 × 104 nuclei/ml (aperture resistance) agreed within 2%. The combined average was taken as the actual stock nuclei concentration, from which all other values were calculated using the known dilution factors. The dashed line is the line of identity.
The higher concentration measured with the hemocytometer was explored further using data from three different operators. Data analogous to that in Figure 2 from multiple batches of cells (n=4) and rat islets (n=3) yielded a slope of 1.16±0.02, and a single measurement with human islets yielded a ratio of 1.17±0.04 for nuclei counted by hemocytometer and flow cytometer. Lastly, nuclei and the cell suspension from which they were prepared were counted with a hemocytometer (Figure 3). The data for nuclei concentration vs cell concentration fit a straight line with a slope of 1.07±0.02. All together, nuclei concentration measured by visual counting with a hemocytometer was higher by about 7–17% (average of all data 12±5%), presumably because of the presence of large cellular fragments, particles, and debris that appeared as nuclei by visual counting but were not registered as nuclei by the two other counting methods.
Nuclei concentration vs cell concentration for INS-1 and βTC3 cells, both measured by visual counting with a hemocytometer. Different symbols represent different batches of cells. The dashed line is the line of identity. The fitted line through the origin has a slope of 1.07±0.01. Suspensions of cells were prepared to provide a wide range of concentrations. Six aliquots were taken from each suspension for counting with the hemocytometer, three for nuclei and three for cell concentration.
The precision of nuclei concentration measurement improved with an increase in the number of nuclei counted (Figure 4a). The coefficient of variation (COV) data were consistent with the prediction for data that follow a Poisson distribution, as is expected for counting data.46 The COV was lower with the medium- and wide-orifice pipette tips than with the narrow-orifice pipette tips; it decreased as the number of IE in the original sample increased (Figure 4b) and was about ⩽6% when ⩾160 IEs were sampled, which provides a reasonable balance between acceptable precision and use of a small number of islets.
(a) Coefficient of variation (COV) from triplicate counts vs number of nuclei (from INS-1 cells) counted by hemocytometer and flow cytometer. Straight line is the predicted COV for data that follow a Poisson distribution. (b) COV vs the number of rat IE (estimated assuming 1560 cells per IE)31 sampled using three different kinds of pipette tips. Samples of different volume were taken from islet stock solutions prepared from each of four batches of islets; nuclei were prepared and diluted to the same concentration within each batch, and 1000 nuclei were counted with the flow cytometer. Below about 200 islets counted, the narrow-orifice pipette tip produced a higher COV than the other two pipette tips.
All three methods were relatively rapid. A nuclei suspension sample took about 5 min to prepare. Visual counting of 500 nuclei (250 per side) took about 10 min. The Coulter Multisizer took an additional 5 min for sample preparation and about 1 min for counting 1000 nuclei. The Guava PCA flow cytometer required 2 min for sample preparation and took about 10 s for counting 1000 nuclei using a sample loaded at about 2.5 × 105 nuclei per ml. All further nuclei counting was carried out with the flow cytometer.
Comparison of Nuclei Counts with DNA Analysis
The slope of the standard curve (absorbance vs DNA concentration) of the same λ-DNA standard was constant for 3 months and increased 25% over the next 2 months. That increase led to a concomitant decrease in DNA measured with DNA controls. Subsequently, we replaced the DNA standard every 3 months. DNA concentration was linearly proportional to the nuclei concentration. With nuclei measured by flow cytometry, DNA concentration measured with the CyQUANT dye was 6.2±0.6 and 6.4±0.8 pg per cell for four batches of βTC3 cells and two batches of rat islets, respectively. Corresponding values measured with PicoGreen dye, 5.6±0.8 and 4.9±0.9 pg per cell for βTC3 cell and rat islets, respectively, were significantly lower (P<0.05). Recent measurements in a different islet center have yielded consistently higher values with PicoGreen than with CyQUANT (unpublished data, Papas K47). Measurements with CyQUANT dye over 3 years yielded an average of 6.5±1.1 pg per cell for 45 different rat islet preparations, 6.9±2.3 pg per cell for 26 preparations of freshly isolated human islets sampled 4–6 h after completion of islet isolation, and 8.5±2.3 pg per cell for 21 samples of human islet preparations shipped to our laboratory from various islet isolation centers.
Comparison of Two Methods to Determine Number of Rat Islet Preparations
The number of IE in 27 rat islet preparations was determined by the standard method of DTZ staining with microscopic visualization and by nuclei counting. All rat islet preparations were of high purity (>95%, most >98%). NIE determined by DTZ staining is plotted vs NIE determined by LM/nuclei counting in Figure 5a. The data are scattered; 23 of 27 estimates of (NIE)DTZ lie above the line of identify, which means that DTZ staining overestimates the number of NIE in comparison to LM/nuclei counting, in some cases substantially. The slope of the best-fit straight line is 1.23±0.09, and the ratio of the mean values is 1.38, corresponding to a 38% overestimate by DTZ staining.
(a) Number of IE by DTZ staining/manual counting vs number of IE by LM/nuclei counting for 27 rat islet preparations. The average NIE was 18 000±6000 for DTZ staining and 13 000±7000 for LM/nuclei counting. (b) Comparison of three methods for determination of number of IE in samples from 12 research preparations of freshly isolated human islet preparations. NIE by DTZ staining/manual counting (closed symbols) and IE by LM/PCV (open symbols) are plotted against NIE by LM/nuclei counting. On average visual counting substantially overestimated the number of IE compared with LM/nuclei counting by about 92% as shown by the slope of the straight line fit to the data. A similar trend is observed with LM/PCV with an overestimation of 67% compared with LM/nuclei counting. The mean values of NIE for all preparations were 87 000, 167 000, and 154 000 for LM/nuclei counting, DTZ staining, and LM/PCV, respectively. Solid lines are from linear regression of the data through the origin. Dashed lines are the lines of identity.
Human Islet Preparations
Data were obtained with 12 freshly isolated human islet research preparations.
Volume fraction islets
The average volume fraction of islets was 0.59±0.21 by LM morphological analysis and 0.72±0.22 by DTZ staining. The substantially higher value by DTZ staining is consistent with our previous observations.31
Comparison of three methods to determine number of IE
We compared the number of IE obtained by three different methods: (1) measurements of islet volume fraction by LM morphological analysis together with the total number of cells by nuclei counting; (2) DTZ staining with manual counting by microscopic visualization; and (3) measurements of islet volume fraction by LM together with PCV measurements. The number of IE measured by each of the three methods is compared in Figure 5b, in which NIE determined by both the DTZ staining and LM/PCV methods is plotted vs NIE determined by LM/nuclei counting. For 11 of the 12 sets of data, both DTZ staining/manual counting and LM/PCV substantially overestimate the number of IE, as reflected by the slopes of the best-fit straight lines. On average, DTZ staining overestimates the LM/nuclei counting measurement by 92%. The slope for DTZ staining is 15% higher than that for LM/PCV.
DISCUSSION
Determination of the amount of islet tissue in an islet preparation, whether by volume, protein content, DNA content, or number of cells, is important in all areas of islet investigation and is especially crucial in islet transplantation. This is the first study to demonstrate that nuclei counting provides a rapid, accurate, and precise method for enumeration of the total number of cells in an islet preparation. Nuclei counting combined with islet volume fraction measurements from morphological analysis using LM is a novel method for accurately determining the volume of islet tissue in a freshly isolated impure islet preparation.
Before this study, we evaluated the dissociation of islets with trypsin with subsequent counting of liberated cells. This approach was not quantitative, some cells were lost, and fractions of original islet cells were not recovered as single cells (data not shown), which is consistent with previous reports.48, 49, 50 Instead, in this study we disrupted the cells with a combination of (1) hypotonic citric acid, which caused osmotic lysis of the cell membrane but not the nuclear membrane,51 (2) Triton-X 100, which reduced the size of cell debris,52 and (3) application of shear force, which freed the nucleus from the cytoplasm, followed by (4) staining with crystal violet, which sharpened the nuclear image.32 The resulting suspension contained no aggregates of nuclei but did contain cellular debris and fragments. Because the nucleus swells reversibly in hypotonic solution without lysis of the nuclear membrane,51 all the nuclei from cells in the islet tissue should be recovered after the procedure, in contrast to the loss of intact cells, which occurs as a result of rupture of some cell membranes following dissociation with serine proteases such as trypsin.
We investigated three methods for counting nuclei, all of which could be used satisfactorily. They differed in the way nuclei were distinguished from fragments, the resulting accuracy, the time required for a measurement, and the range over which the measurement was linear with nuclei concentration (Figures 1, 2, 3). Visual counts with a hemocytometer were linear over the entire concentration range studied but were higher than the other two methods by about 12±5% because of the inability of the eye to consistently distinguish fragments from true nuclei. This was confirmed by visual counting of aliquots of cells and nuclei prepared from the same cell suspension. Data from flow cytometry (Guava PCA) and aperture electrical resistance measurement (Coulter Multisizer II) deviated from linearity at concentrations above about 5 × 105 and 7 × 104 nuclei per ml, respectively. Aperture resistance measurements distinguished cell fragments from nuclei purely on the basis of size. The flow cytometer dot plots with nuclei prepared from islets revealed the presence of a small population of unstained particles producing FSC (which depends on particle size and granularity) comparable to that of the stained nuclei that contained DNA. By setting the 7-AAD fluorescence intensity gate sufficiently high, it was possible to eliminate virtually all these unstained particles.
In addition to providing the most accurate measurement over a wide concentration range, flow cytometry was also the most rapid. Once nuclei samples were prepared (5 min) and stained (2 min), only seconds were required for measurement. Aperture resistance measurements required an additional 5 min for sample preparation, and visual counting took 10 min. Debris and fragments could have been eliminated by repeated centrifugation to make visual counting more accurate but with a substantially lengthened sample preparation time.32 We selected the Guava PCA flow cytometer because it had a small footprint, could be operated with minimal training, and required <20 μl sample volume (and consequently a minimal quantity of islets) because the sample was analyzed while flowing through a microcapillary without sheath fluid, which is used in flow-activated cell sorters that require a much larger sample volume.
Precision of measurements made by visual counting and by the flow cytometer approximately followed Poisson statistics (Figure 4a). Flow cytometry had better precision than visual counting because many more counts could be made in a reasonable time period. The uncertainty in counting nuclei from islets was minimized so long as ⩾160 IEs were sampled (Figure 4b). The COV increased with a smaller sample because of heterogeneity in islet sizes as well as sampling errors. The latter were minimized by using a larger pipette orifice.
We compared nuclei counts by flow cytometry with cell DNA analysis, using two commonly used fluorescent dyes. With CyQUANT, we obtained 6.4–6.5 pg of DNA per cell for rat islets and 6.9 pg of DNA per cell for human islets. These values are comparable to estimates for human cells (6.6 and 6.4 pg per cell for female and male, respectively)53 and to values of 7.3 and 5.7 pg per cell reported48 for rat islet α and β cells, respectively. This agreement between our measurements of DNA content per cell and accepted values in the literature supports the notion that the cell nuclei originally in the tissue were recovered without significant loss. It also supports the accuracy of both flow cytometer nuclei counts and DNA analysis with CyQUANT dye. We obtained lower DNA concentrations per cell with PicoGreen (P<0.05), whereas others observed opposite trends, which illustrates the difficulties of using DNA analysis for measurement of the amount of tissue in islet preparations when different dyes are used. In addition, DNA analysis requires several hours for completion; nuclei counting is much faster. The instability observed in the λ-DNA standard is another potential source of error. For these reasons, we conclude that nuclei counting is more accurate and convenient than DNA measurement for enumeration of cells in islet preparations. Furthermore, we observed values of DNA/cell that were higher than expected with human islet preparations that were received after shipment, which suggests the presence of residual, undegraded DNA released from dead cells. These high values present an additional complexity in using DNA content for measuring the total number of cells in an islet preparation; nonetheless, measurement of the ratio of DNA/cell may be informative with regard to occurrence of recent cell damage in the preparation.
With highly purified rat islet preparations, the number of IE determined with the standard counting method using DTZ stained islets correlated poorly with estimates from LM morphological analysis and nuclei counting and averaged about 38% higher (Figure 5a). Purity was in excess of 95% and thus was not an issue; the overestimation must have been due to errors in volume estimation with DTZ-stained islets and the possible presence of some dying cells in the interior of the islets. Nuclei counting provided a direct measurement of the number of cells that had intact nuclei. Dying cells that retain some structure despite no longer having intact nuclei were not counted.
We also measured the number of IE in 12 freshly isolated human islet preparations using nuclei counting combined with LM morphological analysis of plastic sections, which we have shown has accuracy and precision equivalent to that of EM analysis of ultrathin sections of human islet preparations.31 We compared these measurements with estimates obtained by the standard counting method with DTZ-stained islets and with estimates by LM analysis combined with PCV measurements. DTZ staining and LM/PCV measurements overestimated the number of IE in 12 preparations by an average of about 92 and 67%, respectively.
We attribute the large overestimation of the current standard method of DTZ staining as compared with LM/nuclei counting to three factors: (1) difficulty in identification of the stained islet tissue in impure islet preparations; (2) the presence of some dying cells that occupy volume but no longer have intact nuclei; and (3) large errors in the estimate of islet volume that result from errors in the estimate of the islet diameter. Moreover, the method is prone to systematic error because the actual shape of islets is not spherical,19, 20 which introduces an upward bias leading to overestimation of volume. The three largest dimensions in mutually perpendicular directions (obtained by rotating islets with a micropipette) have been measured in 97 freshly isolated rat islets using a microscope with a calibrated reticule,54 leading to average measured half-axis ratios of b/a=0.82 and c/a=0.60 for an ellipsoid. In the most likely stable configuration, the ellipsoid has its smallest axis c perpendicular to the surface and appears to the viewer during visual size estimation as an ellipse with half axes a and b, from which the radius of an equivalent circle can be estimated. The volume of the equivalent sphere having this radius is 50% greater than the true volume of the ellipsoid. Use of the largest measured half axis for the sphere radius leads to an overestimation of 100%. Thus, the assumption of spherical islets for the purpose of calculating islet volume could account for all or a substantial part of the error incurred by the conventional DTZ staining/manual counting technique.
Overestimation of islet volume by the LM/PCV method may also arise from the presence of dying cells that occupy volume but do not have intact nuclei. In addition, variation in centrifugation methods could result in a pellet void volume fraction substantially larger than the value of 0.3 used in estimating the total tissue volume, which would lead to a substantially larger estimated islet tissue volume than that which was actually present.
We have shown that LM morphological analysis combined with nuclei counting provides accurate, precise, and objective measurement of islet volume and number of IE in impure preparations. LM analysis of plastic sections requires 10 min, but preparation of the sections takes 3 days. Further refinement to permit use of frozen sections for LM analysis would allow the method to be used in real time. As shown by an analysis in the Appendix (Equations (10)–(14)), the number and volume of β cells is approximately linearly proportional to, and can be estimated from, the volume of islet tissue. Our findings suggest that islet volume measured with the DTZ staining/manual counting method grossly overestimates the volume of islet tissue. Compared with measurement by that standard method, the substantially lower value from LM/nuclei counting may be a more accurate indicator of functional islet mass, which is the important parameter for islet transplantation.
References
Shapiro AMJ, Lakey JRT, Ryan EA, et al. Islet transplantation in seven patients with type 1 diabetes mellitus using a glucocorticoid-free immunosuppressive regimen. N Engl J Med 2000;343:230–238.
Hering BJ, Kandaswamy R, Harmon JV, et al. Transplantation of cultured islets from two-layer preserved pancreases in type 1 diabetes with anit-CD3 antibody. Am J Transplant 2004;4:390–401.
Shapiro AMJ, Ricordi C . Unraveling the secret of single donor success in islet transplantation. Am J Transplant 2004;4:295–298.
Knazek RA . The human pancreatic islet cell resource consortium. Diab Technol Ther 2002;4:551–552.
Weber DJ, McFarland RD, Irony I . Selected Food and Drug Administration review issues for regulation of allogeneic islets of Langerhans as somatic cell therapy. Transplantation 2002;74:1816–1820.
Hirshberg B, Rother KI, Harlan DM . Islet transplantation: where do we stand now? Diab Metab Res Rev 2003;19:175–178.
Ricordi C, Lakey JRT, Hering BJ . Challenges toward standardization of islet isolation technology. Transplant Proc 2001;33:1709.
FDA Biological Response Modifier Advisory Committee summary minutes Meeting no. 26, March 20-21, http://www.fda.gov/ohrms/dockets/ac/00minutes/3604m1minutes.pdf, 2000.
Papas KK, Colton CK, Gounarides JS, et al. NMR spectroscopy in β cell engineering and islet transplantation. Ann NY Acad Sci 2001;944:96–119.
Erecinska M, Bryla J, Michalik M, et al. Energy metabolism in islets of Langerhans. Biochimica Biophysica Acta 1992.
Street CN, Lakey JRT, Shapiro AMJ, et al. Islet graft assessment in the Edmonton Protocol: implications for predicting long-term clinical outcome. Diabetes 2004;53:3107–3114.
Goto M, Eich TM, Felldin M, et al. Refinement of the automated method for human islet isolation and presentation of a closed system for in vitro islet culture. Transplantation 2004;78:1367–1375.
Keymeulen B, Ling Z, Gorus FK, et al. Implantation of standardized beta-cell grafts in a liver segment of IDDM patients: graft and recipient characteristics in two cases of insulin-independence under maintenance immunosuppression for prior kidney graft. Diabetologia 1998;41:452–459.
Sever CE, Demetris AJ, Zeng J, et al. Composition of human islet cell preparations for transplantation. Acta Diabetol 1992;28:233–238.
Ricordi C, Gray DW, Hering BJ, et al. Islet isolation assessment in man and large animals. Acta Diabetol Lat 1990;27:185–195.
Scharp DW . Islet quality control testing and the islet isolation laboratory. In: Ricordi C (eds). Pancreatic Islet Cell Transplantation. R.G. Landes Co: Austin, TX, 1992.
Bretzel RG, Alejandro R, Hering BJ, et al. Clinical islet transplantation: guidelines for islet quality control. Transplant Proc 1994;26:388–392.
Latif ZA, Noel J, Alejandro R . A simple method of staining fresh and cultured islets. Transplantation 1988;45:827–830.
Fetterhoff TJ, Wile KJ, Coffing D, et al. Quantitation of isolated pancreatic islets using imaging technology. Transplant Proc 1994;26:3351.
Van der Burg MP, Scheringa M, Basir I, et al. Assessment of isolated islet equivalents. Transplant Proc 1997;29:1971–1973.
Ansite J . Assistant Director of Islet Transplantation Program, University of Minnesota (Personal Communication) 2004.
Stegemann JP, O′Neil JJ, Nicholson DT, et al. Automated counting and sizing of isolated porcine islets using digital image analysis. Transplant Proc 1997;29:2272–2273.
Stegemann JP, O′Neil JJ, Nicholson DT, et al. Improved assessment of isolated islet volume using digital image analysis. Cell Transplant 1998;7:469–478.
Stegemann JP, Raina S, Nicholson DT, et al. Comparison of analytical methods for quantification of isolated porcine hepatocytes yields. Tissue Eng 2000;6:253–264.
O'Neil JJ, Stegemann JP, Nicholson DT, et al. The isolation and function of porcine islets from market weight pigs. Cell Transplant 2001;10:235–246.
Girman P, Kriz J, Friedmansky J, et al. Digital imaging as a possible approach in evaluation of islet yield. Cell Transplant 2003;10:235–246.
Lembert N, Wesche J, Petersen P, et al. A real density measurement is a convenient method for the determination of porcine islet equivalents without counting and sizing individual islets. Cell Transplant 2003;12:33–41.
Lehmann R, Fernandez LA, Bottino R, et al. Evaluation of islet isolation by a new automated method (Coulter Multisizer IIe) and manual counting. Transplant Proc 1998;30:373–374.
Parman AU . Quantitation of isolated rat islets of Langerhans on the basis of deoxyribonucleic acid content under metabolic conditions of altered protein synthesis. J Histochem Cytochem 1975;23:187–193.
Beckmann J, Holze S, Lenzen S, et al. Quantification of cells in islets of Langerhans using DNA determination. Acta Diabetol Lat 1981;18:51–57.
Pisania A, Weir GC, O'Neil JJ, et al. Quantitative analysis of cell composition and purity of human pancreatic islet preparations. Lab Invest 2010; published online 9 August 2010. [e-pub ahead of print].
Sanford KK, Earle WR, Evans VJ, et al. The measurement of proliferation in tissue cultures by enumeration of cell nuclei. J Nat Cancer Inst 1951;11:773–795.
Freshney RI . Culture of animal cells: A manual of basic technique. J. Wiley-Liss: New York, 2000.
Papas KK, Constantinidis I, Sambanis A . Cultivation of recombinant, insulin-secreting AtT-20 cells as free and entrapped spheroids. Cytotechnology 1993;13:1–12.
Efrat S, Linde S, Kofod H, et al. Beta cell lines derived from transgenic mice expressing a hybrid insulin gene-oncogene. Proc Natl Acad Sci USA 1988;85:9037–9041.
Asfari M, Janjic D, Meda P, et al. Establishment of 2-mercaptoethanol-dependent differentiated insulin-secreting cell lines. Endocrinology 1992;130:167–178.
Lacy PE, Kostianovsky M . Method for the isolation of intact islets of Langerhans from the rat pancreas. Diabetes 1967;16:35.
Fraker CA, Alejandro R, Ricordi C . Use of oxygenated perfluorocarbon toward making every pancreas count. Transplantation 2002;74:1811–1812.
Ricordi C, Lacy PE, Finke EH, et al. Automated method for isolation of human pancreatic islets. Diabetes 1988;37:413–420.
Linetsky E, Bottino R, Lehmann R, et al. Improved human islet isolation using a new enzyme blend, liberase. Diabetes 1997;46:1120.
Absher M . Hemocytometer counting. In: Kruse, PFJ, Patterson, MK (eds). Tissue Culture: Methods and Applications. Academic Press: New York, 1973, pp 395–397.
Patterson MK . Measurement of growth and viability of cells in culture. Methods Enzymol 1979;58:141–152.
Weibel ER . Practical Methods for Biological Morphometry. Stereological Methods. Vol. 1, Academic Press: London, 1979.
Ricordi C, Gray DW, Hering BJ, et al. Islet isolation assessment in man and large animals. Acta Diabeta Latina 1990;27:185–195.
Pisania A . Development of quantitative methods for quality assessment of islets of Langerhans, Ph.D. Thesis, Massachusetts Institute of Technology: Cambridge, MA, 2007.
Wardlaw AC . Practical Statistics for Experimental Biologists. Wiley & Sonds Ltd.: New York, 1987.
Papas KK . Comparisons of DNA content using different fluorescent dyes (Personal Communication).
Pipeleers DG, Int' Veld PA, Van De Winkel M, et al. A new in vitro model for the study of pancreatic A and B cells. Endocrinology 1985;117:806–816.
Pipeleers DG, Pipeleers-Marichal M, Hannaert JC, et al. Transplantation of purified islet cells in diabetic rats. I. Standardization of islet cell grafts. Diabetes 1991;40:908–919.
Weir GC, Halban PA, Meda P, et al. Dispersed adult rat pancreatic islet cells in culture: A, B, and D cell function. Metabolism 1984;33:447–453.
Churney L . The osmotic properties of the nucleus. Biol Bull 1942;82:52–67.
Lin AA, Nguyen T, Miller WM . A rapid method for counting cell nuclei using a particle sizer counter. Biotechnol Tech 1991;5:153–156.
Dolezel J, Bartos J, Voglmayr H, et al. Nuclear DNA content and genome size of trout and human. Cytometry Part A 2003;51A:127–128.
Avgoustiniatos ES . Oxygen diffusion limitations in pancreatic islet culture and immunoisolation, Ph.D. Thesis, Massachusetts Institute of Technology, Cambridge, MA, 2002.
Acknowledgements
The βTC3 cells were a gift from Dr Shimon Efrat (Albert Einstein College of Medicine, Bronx, NY, USA). The INS-1 cells were a gift from Dr Chris Newgard (University of Texas Southwestern Medical Center, Dallas, TX, USA). We thank the Islet Isolation Core at Joslin Diabetes Center (Ji Lei, Vaja Tchipashvilli, Gaurav Chandra, and Vassileios Kostaras) for their help in providing islet preparations as well as Chris Cahill of the Joslin DERC Advanced Microscopy Core for the preparation of the 1-μm sections. This study was supported by grants from (1) the National Institutes of Health; (2) Juvenile Diabetes Research Foundation Center for Islet Transplantation at Harvard Medical School; (3) Islet Cell Resource (ICR) at Joslin; and (4) The Diabetes Research and Wellness Foundation (Grant numbers: NIH Grants R01-DK063108-01A1, NCRR ICR U4Z 16606, NCRR ICR U42 RR0023244-01; and the Joslin Diabetes and Endocrinology Research Center (DERC), DK36836).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
APPENDIX
APPENDIX
Number of IE and Islet Volume from Nuclei Counting Data and LM Morphology
LM morphological analysis by stereological point counting produced measurements of the volume fraction ΦI of islet tissue in the preparation. To make use of this information, we employ the framework of equations and parameters we previously developed.31 In the first step, data for ΦI are converted to φIC, the volume fraction of islet cells among all cells, exclusive of extracellular space. The two quantities are related by

where ΦIEC and ΦNIEC are the volume fractions of extracellular space in islet and nonislet tissue, respectively. With values of ΦIEC and ΦNIEC equal to 0.29 and 0.19, respectively,31 Equation (1) may be rearranged to yield

In the second step, φIC is converted to the number fraction of islet cells φIC by the relation

where  =0.925 is the ratio of the average volume per cell for all cells in a human islet preparation divided by the average value for islet cells as determined for 33 clinical islet preparations.31 The total number of all cells nTC in the preparation, obtained by nuclei counting, is then used to estimate the number of islet cells from the estimate of fIC calculated with Equation (3) and the definition of fIC,
=0.925 is the ratio of the average volume per cell for all cells in a human islet preparation divided by the average value for islet cells as determined for 33 clinical islet preparations.31 The total number of all cells nTC in the preparation, obtained by nuclei counting, is then used to estimate the number of islet cells from the estimate of fIC calculated with Equation (3) and the definition of fIC,

Equations (1), (3), and (4) correspond to Equations (30), (18), and (4) in the original derivation.31 Lastly, the number of islet equivalent (IE) in the preparation NIE is determined from

where nIE=1560 cells per IE.31 NIE is also related to the volume of islets VI by

where VIE=1.77 × 10−12 cm3 is the volume of an IE. Islet volume VI can be estimated if NIE is known.
Islet Volume and Number of IE from PCV and LM Morphology
The measured total packed cell pellet volume is VPC and the volume of microscopic voids between the tissue components is VVPC. The volume fraction of voids is defined by

The total tissue volume VT in the pellet is therefore

and the total islet volume VI in the preparation is given by

from which the total number of IEs is calculated using Equation (6). The void volume fraction of the packed cell pellet ΦVPC is estimated to be 0.3 from a measurement of the distribution space of mannitol in pellets from freshly isolated rat islets.45
Relationship between Islet Volume, β-Cell Volume, and Number of β Cells
Rearrangement of Equations (24) and (20), respectively, from the original derivations31 gives expressions for islet cell volume

and β-cell volume

These volume quantities, in turn, can be used to calculate the number of islet cells and β cells in an islet preparation


By combining Equations (31) and (32) in the original derivation, one obtains

ΦIEC is a constant. Because fβIC, the volume fraction of β cells among all islet cells, varies little between different islet preparations,31 to a first approximation it is constant, and the volume (and number) of β cells is directly proportional to the islet volume (or number of IEs) in an islet preparation.
Notations
- a:
-
Half axis of a symmetrical ellipsoid
- b:
-
Half axis of a symmetrical ellipsoid
- c:
-
Half axis of a symmetrical ellipsoid
- f IC :
-
Number fraction of all islet cells
- f βIC :
-
Number fraction of β cells among all islet cells
- n IC :
-
Number of islet cells
- n IE :
-
Number of cells in an IE volume
- n β :
-
Number of β cells
- N IE :
-
Number of IEs
- (NIE)DTZ:
-
Number of IE calculated from diphenylthiocarbazone (DTZ) staining data
- (NIE)LM/Nuclei:
-
Number of IE calculated from nuclei counting and light microscopy (LM) data
- (NIE)LM/PCV:
-
Number of IE calculated from packed cell volume (PCV) and LM data
- V I :
-
Total volume of the islet domain
- V IC :
-
Volume of islet cells
- V IE :
-
Volume of an IE (1.77 × 106 μm3)
- V PC :
-
Volume of packed cell pellet
- V T :
-
Total tissue volume
- V β :
-
Volume of β cells
- φ IC :
-
Volume fraction of islet cells among all cells
- φ βIC :
-
Volume fraction of β cells among all islet cells
- ΦI:
-
Volume fraction of islets
- ΦIEC:
-
Volume fraction of islet extracellular space within the islets
- ΦVPC:
-
Void fraction of packed cell pellet
- (ΦI)DTZ:
-
ΦI calculated from DTZ staining data
- (ΦI)LM:
-
ΦI calculated from LM morphology analysis
- ν β :
-
Cell volume of β cells
-
 :
: -
Average cell volume for islet cells
-
 :
: -
Average cell volume for all the cells
Rights and permissions
About this article
Cite this article
Pisania, A., Papas, K., Powers, D. et al. Enumeration of islets by nuclei counting and light microscopic analysis. Lab Invest 90, 1676–1686 (2010). https://doi.org/10.1038/labinvest.2010.125
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2010.125
Keywords
This article is cited by
-
A replacement for islet equivalents with improved reliability and validity
Acta Diabetologica (2013)








 :
: :
: