Abstract
Recent progress in genomic research has raised expectations for the development of personalized preventive medicine, although genomics-related literacy of patients will be essential. Thus, enhancing genomics-related literacy is crucial, particularly for individuals with low genomics-related literacy because they might otherwise miss the opportunity to receive personalized preventive care. This should be especially emphasized when a lack of genomics-related literacy is associated with elevated disease risk, because patients could therefore be deprived of the added benefits of preventive interventions; however, whether such an association exists is unclear. Association between genomics-related literacy, calculated as the genomics literacy score (GLS), and the prevalence of non-communicable diseases was assessed using propensity score matching on 4646 participants (males: 1891; 40.7%). Notably, the low-GLS group (score below median) presented a higher risk of hypertension (relative risk (RR) 1.09, 95% confidence interval (CI) 1.03–1.16) and obesity (RR 1.11, 95% CI 1.01–1.22) than the high-GLS group. Our results suggest that a low level of genomics-related literacy could represent a risk factor for hypertension and obesity. Evaluating genomics-related literacy could be used to identify a more appropriate population for health and educational interventions.
Similar content being viewed by others
Introduction
In developed countries, aging population and low fertility levels have been causing a shortage of medical resources and an inflation of the cost of medical care.1 Thus, optimizing the quality and expense of medical care is one of the biggest challenges faced today. In this regard, personalized medicine is an attractive option: medical care is optimized for each patient according to the patient-specific information on genetics and genomics, which is more efficient and effective than the ‘mass production method’ currently used to provide medical care. Rapid progress in human genomics research has allowed the clinical validity of genomic information. For example, genomic factors alter the risk of diseases such as cardiovascular diseases and cancers, alter the sensitivity or resistance to medications, and molecular-targeted therapy against malignant tumors has been developed based on genomic factors.2, 3, 4, 5 In the near future, genomic information is expected to lead to personalized preventive medicine.6, 7
Although rapid progress has been made in genomics research, knowledge on how to deliver genomics benefits to the public remains limited.8, 9 Understanding and interpretation of genomic information are critical for appropriate delivery of genomics-based medicine: insufficient knowledge or misconceptions could lead to exaggeration of the potential of the genomic information and dissatisfaction with the information provided.10 Therefore, the public should be provided enough knowledge on genomics, have access to genomic information and be capable of critically interpreting that information,11, 12, 13, 14 as well as have the ability to regularly update this knowledge;14, 15 we refer to this as literacy related to genomics (genomic literacy). We propose that genomic literacy is a part of health literacy, which features a similar concept. Previous research has indicated that an individual genomic literacy could be predicted from their health literacy.16, 17 Health literacy can be evaluated based on the ability to read and write, collect and critically interpret information related to health, and use health or social services, and on the availability of an assistance when the individual faces health problems.18, 19 In the era of personalized medicine, education to enhance genomic literacy will be required before providing genomics-applied health care; however, educating the public and improving their genomic literacy is a daunting task. This will require a substantial amount of knowledge on genetics and genomics in addition to the knowledge necessary for health literacy (that is, basic knowledge of genes, genomes and general genomic science, as well as knowledge of science in general).
Research conducted over the past few decades has revealed that health literacy is associated with health status: a poor health status correlates with a limited knowledge of health.20, 21, 22, 23 This is used as evidence for current intervention that provides health education or counseling to individuals who, for example, present an elevated risk of developing non-communicable diseases that are screened during health checkups, and this intervention has been documented to improve lifestyles or health prognosis.24, 25 By contrast, to the best of our knowledge, such an association has not been described specifically for genomic literacy.11, 26 Thus, we aimed to evaluate our hypothesis that individuals whose genomic literacy is low face the risk of a worse health status than those whose genomic literacy is comparatively higher. We further hypothesized that if such an association were revealed by the results of this study, it would indicate that increased attention would have to be devoted to individuals who exhibit low genomic literacy or are at risk of non-communicable diseases; individuals who could benefit the most from genomics-applied preventive interventions might require added effort to improve their genomic literacy. For example, if an individual was at risk of a certain disease associated with high genetic risk, the disease risk could be reduced by utilizing genomic information. If this individual had only limited genomic literacy, the genomic information might not be understood, or not be used by the individual—extra effort will be required to help him to take advantage of the recognized genetic risk. Identifying the association between genomic literacy and health status would enable us to apply genomics into clinical practice, particularly in the field of preventive medicine, for the appropriate population.
Here, we evaluated the association between genomic literacy and health status by using the baseline data from the Yamagata Study.27 The findings that are presented herein provide suggestions for the development of efficient personalized preventive care.
Materials and methods
Design of the Yamagata study
We used data from the Yamagata study, a population-based cohort study of the general Japanese population over 40 years old. The study design has been described elsewhere.27 Briefly, the basis of this cohort study was a health checkup, and the results obtained were anthropometric traits and data from blood chemical tests. Medical history of non-communicable diseases was obtained using a self-administered questionnaire. Socioeconomic status other than income was obtained from the questionnaire at baseline survey. This cohort study was approved by the Ethics Committee of the Yamagata University Faculty of Medicine, and a written informed consent was obtained from all participants.
Questionnaire survey
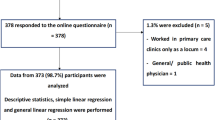
We evaluated genomic literacy by using the questionnaire developed by Professor Zentaro Yamagata (Yamanashi University) and colleagues,26 which was based on previous reports detailing how to determine scientific and genetic literacy.28, 29, 30, 31, 32, 33 We omitted questions regarding participants’ knowledge and contextual understanding of genomic terminology because the Yamagata Study was a genomic cohort study; we explained genomic terminology and research in detail to the participants and ensured every participant understanding based on their consent to join the study. Besides conducting the baseline survey, we mailed the questionnaire to 13 284 participants, all of whom were enrolled before the mailing period, between 14 December and 21 December 2012. A total of 5924 (44.6%) participants returned the questionnaire by mail, and from this group, we excluded the participants who did not answer any questions (n=10) or did not answer all questions required to evaluate genomic literacy (n=1268). Finally, data from 4646 participants recruited from 2010 to 2012 were available for the analysis. This survey was also approved by the Ethics Committee of the Yamagata University Faculty of Medicine.
Assessment of genomic literacy
The questionnaire used in the study by Ishiyama et al.26 included the following factors: (1) knowledge of genomic terminology (subjective understanding), (2) contextual understanding of genomic terminology (objective understanding), (3) awareness of the benefits and risks of genomic studies, (4) subjective understanding of genomic research, and (5) subjective understanding of the aims and applicability of genomic research. Ishiyama et al.26 used factors 1–3 to evaluate genomic literacy. As noted above, we omitted factors 1 and 2, and thus used factors 3–5. For factor 3, we asked six questions that were related to whether the participants were aware of the benefits and risks of genomic studies. For factor 4, we asked whether ‘they knew’ or ‘did not know’ the answers to each seven questions that were related to the subjective understanding of genomic research (for example, ‘Did you know that genomic studies are trying to investigate the types and mechanisms of proteins that are produced by the function of genes?’). For factor 5, we asked whether ‘they knew’ or ‘did not know’ the answers to seven questions that were related to subjective understanding of the aims and applicability of genomic research (for example, ‘Did you know that genomic research is aiming to clarify the mechanisms of disease onset?’).
Calculation of genomic literacy score
We calculated the participants’ genomic literacy score (GLS) according to the study conducted by Ishiyama et al.:26 the questions related to whether the participants were aware of the benefits and risks of genomic studies were scored as 1 for the participants who showed awareness and as 0 for the other participants, and the total score ranged from 0 to 6; subjective contextual understanding of genomic research and its aims and applicability were scored as 1 when the participants answered that ‘they knew’ about the context in the questions and as 0 when they ‘did not know,’ and the total score ranged from 0 to 7 for each factor. Each domain of the score was converted to 10 points to equalize the weights of all domains, and the final score ranged from 0 to 30. We checked the consistency of the GLS by performing the same analysis as in the original study:26 we compared the mean of the GLS among each of the demographic factors and the 10 questions ‘related to experience, sources of information, knowledge, and attitudes toward science in general and genetic testing in particular that might have an effect on decision-making and the level of genomic literacy.’26
Collection of data on non-communicable diseases
We obtained data on the prevalence of diabetes, dyslipidemia, heart disease (heart failure, angina pectoris and myocardial infarction), hypertension, obesity and stroke (intracranial hemorrhage, subarachnoid hemorrhage and cerebral infarction) based on a known diagnosis. Moreover, undiagnosed participants were included as having the aforementioned health conditions according to the following criteria: patients with diabetes were defined as patients with fasting plasma glucose ⩾6.99 mmol l−1 (126 mg dl−1), postprandial glucose ⩾11.10 mmol l−1 (200 mg dl−1), or glycosylated hemoglobin (HbA1c) ⩾0.065 (6.5%), or as patients on diabetes treatment; patients with dyslipidemia were defined as patients with triglyceride ⩾1.69 mmol l−1 (150 mg dl−1), low-density lipoprotein cholesterol ⩾3.63 mmol l−1 (140 mg dl−1), or high-density lipoprotein cholesterol <1.04 mmol l−1 (40 mg dl−1), or as patients on a treatment for dyslipidemia; hypertension patients were defined as patients with systolic blood pressure ⩾18.67 kPa (140 mm Hg) or diastolic blood pressure ⩾12.00 kPa (90 mm Hg), or as patients on a treatment for hypertension; and obesity was defined as body mass index ⩾25 kg m−2.34, 35, 36 HbA1c values were determined as Japan Diabetic Society (JDS) values and thus converted to National Glycohemoglobin Standardization Program values by adding 0.004 (0.4%) to the Japan Diabetic Society values.36
Statistical analysis
Welch’s t-test and Welch’s analysis of variance were used to compare the mean GLS values. Participants were stratified by gender, according to the previous study.26 We calculated the odds ratio for the approval of promoting genomic research and for the prevalence of non-communicable diseases per increase of 0.5 times the s.d. in GLS. We also adjusted the analysis based on factors used to calculate the propensity score: to elucidate the association between the GLS and the prevalence of non-communicable diseases, we used propensity score matching to minimize bias. We stratified the participants into high- and low-GLS groups based on their GLS being above and below the median GLS, respectively. Propensity scores were calculated for these GLS groups, using logistic regression models, and adjusted for factors based on previous studies; among those presented in Table 2, factors that could be considered pretreatment variables were used for adjustment—age, sex, educational background, parenthood, annual household income, whether or not the participants learned genetics in school, whether they enjoyed the study of science while in elementary or junior high school, whether they had ever heard about genomic research, and whether they were interested in general science and technology.37 These factors also included indicators of socioeconomic status to separate out the resultant confounding bias. We matched participants with the propensity score by using 1:1 nearest-neighbor matching without replacement, and within a caliper width of 0.05 times the s.d. of the logit.38 Matching was performed using matchit function from the MatchIt package, and balance of the covariates after matching was checked based on standardized difference by using CreateTableOne function from the tableone package in R software.39 Within the matched pairs, we used Agresti and Min’s40 method to compare the prevalence of non-communicable diseases between the GLS groups. We used an indicator variable for answer refusal, and a blank answer for categorical variables. All data were present for the factor ‘age’, which was the only continuous variable. P<0.05 was considered statistically significant. All statistical analyses were performed using R software version 3.2.5.39
Results
For this analysis, 4646 participants (1891 males; 40.7%) were available, whose mean age was 61.9 years (s.d. 8.4 years) and mean GLS was 14.2 (s.d. 7.2) (Table 1). We also performed a complete case analysis, which was conducted without the use of the indicator variable, and the age and GLS of these participants are also shown in Table 1. Demographic factors and other characteristics are shown in Table 2 and Supplementary Table S1.
When checking the consistency of the GLS, we grouped participants according to their demographic factors and responses to the questions, and we compared the mean GLS among these groups. The findings obtained were comparable between the analysis of total participants and that of complete cases in all variables except housing; results of the analysis of total participants are shown in Table 2. Mean GLS differed significantly among the groups by marital status, occupation, annual household income, and the 10 questions related to genomics (Table 2, Supplementary Table S1). However, the GLS did not differ among the groups that were stratified according to participant age or parenthood status, or based on whether the participants lived with their family.
In the adjusted logistic regression analysis, which was performed for approval of promoting genomic research and for the prevalence of non-communicable diseases, an elevated GLS was associated with the participants’ approval of promoting genomic research (odds ratio 2.11, 95% confidence interval (CI) 1.95–2.28) and a diminished risk of hypertension (odds ratio 0.91, 95% CI 0.86–0.97) (Table 3).
Table 4 shows the results of the propensity score matching. The standardized difference after matching was <10% for all matching variables in all outcomes, which showed that no imbalance existed between the groups (Supplementary Table S2). Disease risk in the lower-GLS group was increased relative to that in the higher-GLS group for hypertension (relative risk 1.09, 95% CI 1.03–1.16) and obesity (relative risk 1.11, 95% CI 1.01–1.22). The risk of diabetes, heart disease, dyslipidemia and stroke did not differ between the GLS groups (Table 4).
Discussion
The results of this study suggest that individuals possessing a low level of genomic literacy could be at risk of hypertension and obesity. This supports our hypothesis that a low level of genomic literacy is related to a poor health status. Low health literacy, which encompasses genomic literacy, has been reported to be associated with a poor health status resulting from unhealthy behavior.16, 17, 41, 42, 43 Thus, the increased risk of non-communicable diseases associated with low genomic literacy could be explained by the unfavorable health behaviors of the patients. For example, a person with low genomic literacy might exhibit an elevated intake of energy, salt or sugar, which would cause an increase in body mass index or blood pressure. However, our findings notably indicated that genomic literacy could be an individual factor related to poor health status, which was revealed by reducing confounding bias of variables such as socioeconomic status by using propensity score matching. Nevertheless, there may still be a confounding bias related to unobserved health literacy. We must also consider reverse causation, where the development of a disease might affect the level of genomic literacy and likely raise genomic literacy by increasing interest in health and disease, and this might be especially true in the case of our study population, which was composed of healthy individuals who voluntarily submitted to a health checkup. Thus, the level of genomic literacy of the participants who were diagnosed with non-communicable diseases might have been lower before the development of diseases. A follow-up study that we will conduct on this cohort will enable a more accurate interpretation. In addition, because our genomic cohort study was based on health checkups and participants were those who gave consent to the cohort study,27 genomic literacy of our study population could have been relatively higher than others. To generalize our results, we should evaluate the association between genomic literacy and health status in a more diverse population, especially of participants whose genomic literacy is lower.
Our results indicate that a person whose genomic literacy is inadequate should receive educational intervention when offered genomics-applied health care; effective and efficient intervention must be provided by using genomic information to evaluate their intrinsic risk.9, 44 This study showed that individuals of higher priority for genomics-applied health care, that is, individuals at risk of developing diseases, were likely to exhibit a lower than average level of genomic literacy, at least in the case of hypertension and obesity. Therapeutic intervention for overweight patients and diabetic patients is recognized to be more effective for patients whose health literacy is low than for patients whose health literacy is high.45, 46 This could be explained by the larger available room for improvement in health behavior and lifestyle in individuals with lower rather than higher levels of health literacy. The association between the effect of a therapeutic intervention and genomic literacy remains to be elucidated, but we speculate that genomics-applied health care could be efficiently and effectively delivered by providing intervention to individuals with a low level of genomic literacy. However, a caveat here is that these individuals might refuse genomics-applied health care due to prejudice or misconception resulting from their low level of genomic literacy. Thus, primarily targeting individuals with low genomic literacy would be crucial for efficiently and effectively providing genomics-applied health care to those who will likely benefit the most from the treatment. Genomic literacy is improved through genomic education, including genomic counseling.47, 48, 49 If genomic education is to be applied to daily health care or primary preventive intervention, primary education on terms such as gene, genome and chromosome will be required for individuals with a low level of genomic literacy. In the era of genomics-applied health care, the importance of evaluating genomic literacy and providing adequate education before interventions must be widely recognized by health-care practitioners.
The association between the GLS calculated in this study and the background factors of genomic literacy—socioeconomic status and attitudes toward genomic studies—was comparable to that in the study by Ishiyama et al.;26 for example, the GLS was higher for participants with a high socioeconomic status than for participants of a low socioeconomic status. Moreover, although the GLS did not differ among the different age groups, younger individuals were previously found to present a higher level of genomic literacy than older individuals;26 in the study by Ishiyama et al., 37% of the participants were under 40 years old, whereas all of our study participants were over 40 years old. This was also why most of the participants were married and had children. We must also take into account the difference in factors that were used to evaluate GLS. However, in terms of age composition, our study could be relatively more valuable for preventive medicine targeting non-communicable diseases: individuals over 40 years old, whose population is increasing in aging societies, face a higher risk of non-communicable diseases than younger individuals. Thus, at least for individuals over 40 years of age, we suggest that assessing genomic literacy will be useful for stratifying individuals who require educational and therapeutic interventions.
There were limitations to this study. First, the prevalence of diabetes, heart disease and stroke was low, which could have affected the statistical power of the analysis. Hypertension and obesity, the diseases for which statistically significant results were obtained, showed relatively higher prevalence. We can potentially further clarify the association between genomic literacy and the risk of non-communicable diseases by conducting either a follow-up survey for this prospective cohort study or a nested case–control study. Second, we performed a propensity score matching to reduce the effects of confounding to the maximal extent possible; we considered factors related to health literacy, such as socioeconomic status (including educational background).37 However, we could not estimate the effects of unobserved and/or unrecognized confounders. This is important because, genomic literacy could be considered to consist of multifactorial components and needs to be evaluated comprehensively. Propensity score matched analysis using scores of factors 3–5, which was used to calculate GLS alone, did not show any significant association with health status (data not shown). Thus, taking into account more related factors while removing bias of other confounders would improve the quality of the analysis. Unobserved confounders include subjective and objective understanding of genomic terminology, which was assessed in the study by Ishiyama et al.26 (factors 1 and 2). As described in the ‘Materials and Methods’, we ensured that all participants understood genomic terminology at least once when giving their consent to the study, and thus we determined that including these factors would cause a bias. Third, although the questionnaire response rate was relatively high (44.6%), a selection bias must be considered; genomic literacy of non responders might be lower than that of our population. Fourth, the period between consent to the study and questionnaire survey varied among participants. This difference might have affected both their literacy and health status.
In conclusion, we showed that a low level of genomic literacy could be a risk factor for hypertension and obesity. Genomic literacy could be used as an indicator to identify a more appropriate target population for genomics-applied preventive medicine, that is, personalized preventive medicine. Because evaluating genomic literacy will be necessary for personalized preventive medicine, future interventional studies conducted using genomics, particularly studies aiming to prevent non-communicable diseases, should consider stratifying patients according to genomic literacy.
References
National Institute on Aging, National Institutes of Health, U.S. Department of Health and Human Services, World Health Organization. Global Health and Aging. NIH publication no. 11-7737, 2011.
Ashley, E. A., Butte, A. J., Wheeler, M. T., Chen, R., Klein, T. E., Dewey, F. E. et al. Clinical assessment incorporating a personal genome. Lancet 375, 1525–1535 (2010).
Antoniou, A., Pharoah, P. D. P., Narod, S., Risch, H. A., Eyfjord, J. E., Hopper, J. L. et al. Average risks of breast and ovarian cancer associated with BRCA1 or BRCA2 mutations detected in case series unselected for family history: A combined analysis of 22 studies. Am. J. Hum. Genet. 72, 1117–1130 (2003).
Tantisira, K. G., Lasky-Su, J., Harada, M., Murphy, A., Litonjua, A. A., Himes, B. E. et al. Genomewide association between GLCCI1 and response to glucocorticoid therapy in asthma. N. Engl. J. Med. 365, 1173–1183 (2011).
McCarthy, J. J., McLeod, H. L. & Ginsburg, G. S. Genomic medicine: a decade of successes, challenges, and opportunities. Genomic medicine: a decade of successes, challenges, and opportunities. Sci. Transl. Med. 5, 189sr184 (2013).
Kaphingst, K. A., Stafford, J. D., McGowan, L. D., Seo, J., Lachance, C. R. & Goodman, M. S. Effects of racial and ethnic group and health literacy on responses to genomic risk information in a medically underserved population. Health Psychol. 34, 101–110 (2015).
Buford, T. W. & Pahor, M. Making preventive medicine more personalized: implications for exercise-related research. Prev. Med. 55, 34–36 (2012).
McBride, C. M., Bowen, D., Brody, L. C., Condit, C. M., Croyle, R. T., Gwinn, M. et al. Future health applications of genomics: priorities for communication, behavioral, and social sciences research. Am. J. Prev. Med. 38, 556–565 (2010).
Syurina, E. V., Brankovic, I., Probst-Hensch, N. & Brand, A. Genome-based health literacy: a new challenge for public health genomics. Public Health Genomics 14, 201–210 (2011).
Press, N. A., Yasui, Y., Reynolds, S., Durfy, S. J. & Burke, W. Women's interest in genetic testing for breast cancer susceptibility may be based on unrealistic expectations. Am. J. Med. Genet. 99, 99–110 (2010).
Ashida, S., Goodman, M., Pandya, C., Koehly, L. M., Lachance, C., Stafford, J. et al. Age differences in genetic knowledge, health literacy and causal beliefs for health conditions. Public Health Genomics 14, 307–316 (2011).
Vermeulen, E., Henneman, L., van El, C. G. & Cornel, M. C. Public attitudes towards preventive genomics and personal interest in genetic testing to prevent disease: a survey study. Eur. J. Public Health 24, 768–775 (2014).
Gaskell, G., Allum, N., Bauer, M., Durant, J., Allansdottir, A., Bonfadelli, H. et al. Biotechnology and the European public. Nat. Biotechnol. 18, 935–938 (2000).
O'Daniel, J. M. The prospect of genome-guided preventive medicine: a need and opportunity for genetic counselors. J. Genet. Couns. 19, 315–327 (2010).
Khoury, M. J., Gwinn, M. L., Glasgow, R. E. & Kramer, B. S. A population approach to precision medicine. Am. J. Prev. Med. 42, 639–645 (2012).
Lea, D. H., Kaphingst, K. A., Bowen, D., Lipkus, I. & Hadley, D. W. Communicating genetic and genomic information: health literacy and numeracy considerations. Public Health Genomics 14, 279–289 (2011).
Kaphingst, K. A., Blanchard, M., Milam, L., Pokharel, M., Elrick, A. & Goodman, M. S. Relationships between health literacy and genomics-related knowledge, self-efficacy, perceived importance, and communication in a medically underserved population. J. Health Commun. 21 (Suppl 1), 58–68 (2016).
Osborne, R. H., Batterham, R. W., Elsworth, G. R., Hawkins, M. & Buchbinder, R. The grounded psychometric development and initial validation of the Health Literacy Questionnaire (HLQ). BMC Public Health 13, 658 (2013).
Zoellner, J., You, W., Almeida, F., Blackman, K. C., Harden, S., Glasgow, R. E. et al. The influence of health literacy on reach, retention, and success in a worksite weight loss program. Am. J. Health Promot. 30, 279–282 (2015).
Wolf, M. S., Gazmararian, J. A. & Baker, D. W. Health literacy and functional health status among older adults. Arch. Intern. Med. 165, 1946–1952 (2005).
Baker, D. W., Wolf, M. S., Feinglass, J., Thompson, J. A., Gazmararian, J. A. & Huang, J. Health literacy and mortality among elderly persons. Arch. Intern. Med. 167, 1503–1509 (2007).
Schillinger, D., Grumbach, K., Piette, J., Wang, F., Osmond, D., Daher, C. et al. Association of health literacy with diabetes outcomes. JAMA 288, 475–482 (2002).
Geboers, B., de Winter, A. F., Luten, K. A., Jansen, C. J. & Reijneveld, S. A. The association of health literacy with physical activity and nutritional behavior in older adults, and its social cognitive mediators. J. Health Commun. 19 (Suppl 2), 61–76 (2014).
Ko, G. T., Li, J. K., Kan, E. C. & Lo, M. K. Effects of a structured health education programme by a diabetic education nurse on cardiovascular risk factors in Chinese Type 2 diabetic patients: a 1-year prospective randomized study. Diabet. Med. 21, 1274–1279 (2004).
Japanese Ministry of Health, Labour and Welfare Final summary from working-group for verification of effectiveness of specific health checkups and guidance on optimizing medical cost (in Japanese), 2015. Available at: http://www.mhlw.go.jp/file/05-Shingikai-12401000-Hokenkyoku-Soumuka/0000090330.pdf (accessed 26 January 2017).
Ishiyama, I., Nagai, A., Muto, K., Tamakoshi, A., Kokado, M., Mimura, K. et al. Relationship between public attitudes toward genomic studies related to medicine and their level of genomic literacy in Japan. Am. J. Med. Genet. A 146A, 1696–1706 (2008).
Yamagata University Genomic Cohort Consortium & Narimatsu, H. Constructing a contemporary gene-environmental cohort: study design of the Yamagata Molecular Epidemiological Cohort Study. J. Hum. Genet. 58, 54–56 (2013).
Miller, J. D. The measurement of civic scientific literacy. Public Underst. Sci. 7, 203–223 (1998).
Furr, L. A. & Kelly, S. E. The Genetic Knowledge Index: developing a standard measure of genetic knowledge. Genet. Test. 3, 193–199 (1999).
Henneman, L., Timmermans, D. R. & van der Wal, G. Public experiences, knowledge and expectations about medical genetics and the use of genetic information. Community Genet. 7, 33–43 (2004).
Jallinoja, P. & Aro, A. R. Does knowledge make a difference? The association between knowledge about genes and attitudes toward gene tests. J. Health Commun. 5, 29–39 (2000).
Shaw, J. S. 3rd & Bassi, K. L. Lay attitudes toward genetic testing for susceptibility to inherited diseases. J. Health Psychol. 6, 405–423 (2001).
Calnan, M., Montaner, D. & Horne, R. How acceptable are innovative health-care technologies? A survey of public beliefs and attitudes in England and Wales. Soc. Sci. Med. 60, 1937–1948 (2005).
Konta, T., Hao, Z., Abiko, H., Ishikawa, M., Takahashi, T., Ikeda, A. et al. Prevalence and risk factor analysis of microalbuminuria in Japanese general population: the Takahata study. Kidney Int. 70, 751–756 (2006).
Seino, Y., Nanjo, K., Tajima, N., Kadowaki, T., Kashiwagi, A., Araki, E. et al. Report of the committee on the classification and diagnostic criteria of diabetes mellitus. J. Diabetes Investig. 1, 212–228 (2010).
Japan Atherosclerosis Society. Japan Atherosclerosis Society (JAS) Guidelines for Prevention of Atherosclerotic Cardiovascular Diseases (in Japanese), 2012..
Rosenbaum, P. R. & Rubin, D. B. The central role of the propensity score in observational studies for causal effects. Biometrika 70, 41–55 (1983).
Austin, P. C. Optimal caliper widths for propensity-score matching when estimating differences in means and differences in proportions in observational studies. Pharm. Stat. 10, 150–161 (2011).
R Foundation for Statistical Computing, R Core Team R: a language and environment for statistical computing, 2016. Available at: https://www.R-project.org/.
Agresti, A. & Min, Y. Effects and non-effects of paired identical observations in comparing proportions with binary matched-pairs data. Stat. Med. 23, 65–75 (2004).
Kim, S. H. Health literacy and functional health status in Korean older adults. J. Clin. Nurs. 18, 2337–2343 (2009).
McNaughton, C. D., Kripalani, S., Cawthon, C., Mion, L. C., Wallston, K. A. & Roumie, C. L. Association of health literacy with elevated blood pressure: a cohort study of hospitalized patients. Med. Care. 52, 346–353 (2014).
Chari, R., Warsh, J., Ketterer, T., Hossain, J. & Sharif, I. Association between health literacy and child and adolescent obesity. Patient Educ. Couns. 94, 61–66 (2014).
Patel, C. J., Sivadas, A., Tabassum, R., Preeprem, T., Zhao, J., Arafat, D. et al. Whole genome sequencing in support of wellness and health maintenance. Genome Med. 5, 58 (2013).
Chinn, D. & McCarthy, C. All Aspects of Health Literacy Scale (AAHLS): developing a tool to measure functional, communicative and critical health literacy in primary healthcare settings. Patient Educ. Couns. 90, 247–253 (2013).
Rothman, R. L., DeWalt, D. A., Malone, R., Bryant, B., Shintani, A., Crigler, B. et al. Influence of patient literacy on the effectiveness of a primary care-based diabetes disease management program. JAMA 292, 1711–1716 (2004).
Nutbeam, D. The evolving concept of health literacy. Soc. Sci. Med. 67, 2072–2078 (2008).
Armstrong, J., Toscano, M., Kotchko, N., Friedman, S., Schwartz, M. D., Virgo, K. S. et al. Utilization and outcomes of BRCA genetic testing and counseling in a national commercially insured population: The ABOUT Study. JAMA Oncol. 1, 1251–1260 (2015).
Kaphingst, K. A., Facio, F. M., Cheng, M. R., Brooks, S., Eidem, H., Linn, A. et al. Effects of informed consent for individual genome sequencing on relevant knowledge. Clin. Genet. 82, 408–415 (2012).
Acknowledgements
We thank Professor Zentaro Yamagata of Yamanashi University for kindly providing us with the study questionnaire. We thank Editage (www.editage.jp) for English-language editing. This work was partially supported by JSPS KAKENHI (Grant JP 16H06277 to HN).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Nakamura, S., Narimatsu, H., Katayama, K. et al. Effect of genomics-related literacy on non-communicable diseases. J Hum Genet 62, 839–846 (2017). https://doi.org/10.1038/jhg.2017.50
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2017.50
This article is cited by
-
The impact of needs-based education on the change of knowledge and attitudes towards medical genetics in medical students
European Journal of Human Genetics (2021)
-
Awareness of paternal age effect disorders among Japanese pregnant women: implications for prenatal genetic counseling for advanced paternal age
Journal of Community Genetics (2021)