Abstract
The purpose of this noninvasive prenatal testing (NIPT) study was to compare the fetal fraction of singleton gestations by gestational age, maternal characteristics and chromosome-specific aneuploidies as indicated by z-scores. This study was a multicenter prospective cohort study. Test data were collected from women who underwent NIPT by the massively parallel sequencing method. We used sequencing-based fetal fraction calculations in which we estimated fetal DNA fraction by simply counting the number of reads aligned within specific autosomal regions and applying a weighting scheme derived from a multivariate model. Relationships between fetal fractions and gestational age, maternal weight and height, and z-scores for chromosomes 21, 18 and 13 were assessed. A total of 7740 pregnant women enrolled in the study, of which 6993 met the study criteria. As expected, fetal fraction was inversely correlated with maternal weight (P<0.001). The median fetal fraction of samples with euploid result (n=6850) and trisomy 21 (n=70) were 13.7% and 13.6%, respectively. In contrast, the median fetal fraction values for samples with trisomies 18 (n=35) and 13 (n=9) were 11.0% and 8.0%, respectively. The fetal fraction of samples with trisomy 21 NIPT result is comparable to that of samples with euploid result. However, the fetal fractions of samples with trisomies 13 and 18 are significantly lower compared with that of euploid result. We conclude that it may make detecting these two trisomies more challenging.
Similar content being viewed by others
Introduction
Noninvasive prenatal testing (NIPT) by massively parallel sequencing has been reported to be highly accurate for the detection of fetal chromosomal aneuploidies.1, 2, 3, 4, 5 This has resulted in widespread adoption of this screening test. Although NIPT has a higher accuracy than conventional prenatal screening method, patients must understand the implications of the results before undergoing testing, including the likelihood of test failure, false positives, false negatives and findings of unclear significance.3
In Japan, NIPT for trisomies 21, 18 and 13 was started in April 2013, after receiving approval from the Japan Society of Obstetrics and Gynecology (JSOG) and the Japanese Association of Medical Sciences (JAMS). The initial nationwide trial was conducted by the Japan NIPT consortium.6 The JAMS has determined that NIPT should be permitted at institutes where appropriate genetic counseling is available.6, 7 The indications for NIPT included a positive maternal serum screen result for an aneuploidy, fetal ultrasound findings indicating an increased risk of aneuploidy, history of a prior pregnancy with a trisomy or maternal age of 35 years or older at the time of delivery.
The placenta releases significant levels of fetal DNA into the maternal circulation, with cell-free fetal DNA fractions reaching levels of 10–20% between 10 and 21 weeks of gestation.8, 9 The cell-free fetal DNA is derived from apoptotic trophoblastic cells in the placenta.10 Fetal fraction is an important parameter that affects the performance of cell-free fetal DNA-based prenatal tests.8 Samples with sufficient fetal fractions that pass quality control metrics can provide an accurate assessment of the chromosomes tested.3, 8 Several lines of evidence suggest that the test performance for trisomy 21 is better than trisomies 18 and 13.2, 8 In contrast, some data show the findings indicating similar detection ability for trisomies 13 and 18 relative to trisomy 21.9
Fetal fraction, a key parameter that ensures adequate fetal chromosomal representation, is affected by maternal weight, maternal body mass index (BMI), gestational age and fetal aneuploidy.3, 9, 11 Recent reports suggest that fetal fraction correlated positively with gestational age and negatively with maternal weight. Studies that compared fetal fraction among average risk pregnancies in the first trimester did not find it significantly different compared with fetal fractions in high-risk women.11, 12 The purpose of our study was to compare fetal fractions by gestational age, maternal weight and height, BMI, indication of NIPT and z-scores for chromosomes 21, 18 and 13. In addition, we examined if a relationship exists between fetal fraction and trisomy involving chromosomes 21, 18 and 13.
Materials and methods
Study population
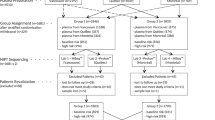
Pregnant women with high risk for fetal aneuploidy and singleton gestation were enrolled at 10 to 20 weeks of gestation. The high-risk indications included maternal age of ⩾35 years at the time of delivery, abnormal fetal ultrasound, abnormal serum screen, personal history of a child with aneuploidy or a parent carrying a balanced Robertsonian translocation with an increased risk of trisomy 13 or 21. The study design was approved by all of the hospitals’ Institutional Review Board and all women provided informed written consent to participate. NIPT for trisomies 21, 18 and 13 using cell-free DNA in maternal plasma was performed among high-risk pregnant women who requested testing at institutions authorized by the JAMS between April 2013 and March 2014.6 The details of the study protocol, including the recruitment of high-risk pregnant women who requested testing, are provided on the internet (http://www.nipt.jp/).
Sample collection and preparation
Blood samples (20 ml) were collected from the pregnant women at each institution and were sent to Sequenom Laboratories (San Diego, CA, USA) for MaterniT21 Plus tests within 7 days of collection. If the results were positive, then either amniocentesis or chorionic villus sampling was performed for conventional karyotyping. Exclusion criteria included cases with missing information about maternal characteristics, multiple gestation or fetal demise before NIPT.
Test methods
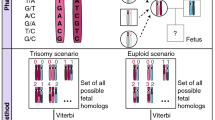
Cell-free maternal plasma DNA extracted from each sample was subjected to library preparation and massively parallel sequencing using Illumina HiSeq 2000 (Illumina, San Diego, CA, USA) as described earlier.2, 11 SeqFF method is a multivariate regression model that determines fetal DNA fraction.12 In brief, we used sequencing-based fetal fraction calculations in which we estimated fetal DNA fraction by simply counting the number of reads aligned within specific autosomal regions and applying a weighting scheme derived from a multivariate model. The response variable could be any quantitative metric that reflects fetal DNA fraction. For SeqFF, chromosome Y was chosen as this will allow for the direct comparison of fetal DNA fraction from sequence data rather than a secondary assay. The predictor variables were the aggregated normalized counts of single-end sequence reads aligned to 50 kb contiguously partitioned regions of the human reference genome (hg19). As the magnitude of copy number variation can also be used to estimate fetal DNA fraction, bins located on chromosomes 13, 18, 21, X and Y are excluded from the SeqFF method to avoid issues of model overfitting and circular evidence.12
The sequencing data were used to calculate z-score, which are robust estimates of normalized chromosomal representation compared with a euploid genome. All samples were required to meet the quality control criteria, including a minimum fetal fraction. Z-scores of 3 or above were considered to be indicative of trisomy 21, and z-scores 3.95 or above were considered to be indicative of trisomies 13 or 18.
Confirmatory invasive testing
Cases with positive result on NIPT were followed up by villus sampling or amniocentesis to confirm the finding. In cases with intrauterine fetal demise, chorionic villus sampling was performed. Following standard metaphase conversion of cultured fetal cells, conventional karyotyping was performed and at least 20 cells were analyzed. The clinical data, test results and pregnancy outcomes were collected and aggregated every month at the data center of the secretariat. This study is a part of a clinical trial registered with the University Medical Information Network clinical trials registry (UMIN000009338).
Statistical analysis
Statistical methods were used to evaluate the correlation between fetal fraction, maternal characteristics and z-scores of chromosomes 21, 18 and 13. Descriptive data of demographic information are presented as median and interquartile range. The measured fetal fraction was represented as square root (√) transformed distribution to ensure the normality as described earlier.13 The association between fetal fraction and maternal weight was calculated by Jonckheere–Terpstra trend test. The differences among levels of variables were compared pairwise using one-way analysis of variance test with post hoc Tukey's HSD (honest significant differences) test. P-value of ⩽0.05 indicated a statistically significant difference. Relationships between fetal DNA fraction and z-scores in chromosomes 21, 18 and 13 were demonstrated as scatter plots. The statistical analyses, except the trend test, were performed using statistical software package SPSS 22.0 (SPSS, Chicago, IL, USA). The Jonckheere–Terpstra trend test was performed using R version 2.13.0, EZR on R commander version 1.1 designed to add statistical functions frequently used in biostatistics.14, 15, 16
Results
Of 7740 women who participated in the study, 747 were excluded owing to the lack of details such as maternal and gestational age. Of the 6993 high-risk pregnant women tested in this study, two cases had fetal fraction over 60% and were excluded from the analysis of fetal fraction metrics. Maternal and fetal characteristics of the study population are shown in Table 1 and the frequency distribution of maternal plasma √fetal DNA fractions is presented in Figure 1. The √fetal fraction has a bell-shaped distribution that peaks between 20 and 40% at 10–20 weeks of gestation.
We examined the relationship between fetal cell-free DNA fraction and gestational age. The median fetal DNA fraction within 10–20 weeks of gestation was 13.7%, with an interquartile range of 10.7–17.9%. More than 99.8% of samples (n=6981) had fetal fraction above the lower acceptable limit for accurate interpretation of fetal aneuploidy. There was no change in fetal DNA fraction from 10 to 20 weeks (R2=0.02).
More than 95% of the tests (95.5%, 6677/6993) were performed in women 35 years of age or older with a median age of 38.0 (22–49) years. The median gestational age at the time of testing was 13.0 (10.0–20.2) weeks, the median maternal weight was 52.0 (34.0–115.0) kg and the median BMI was 20.5 (14.5–45.3) kg m−2 (Table 1). We excluded three of 6993 women because their weight and height data were missing (Table 1). Association between fetal fraction and maternal weight are presented in Figure 2. There was an overall inverse relationship between fetal fraction and maternal weight with a median fetal fraction of 18.1% and 9.6 for maternal weight of <40 and >90 kg, respectively. There was a significant correlation of fetal DNA fraction in early gestational age with maternal weight in 6990 pregnancies, with Jonckheere–Terpstra test (trend test) (P<0.001).
In 0.26% of samples (18/6993), NIPT failed because of insufficient fetal DNA or other technical reason, and a ‘not reportable’ result was issued, of which 16 were retested. Thirteen women found to be negative for fetal aneuploidy, one case had trisomy 18 and two women again received not reportable results. The number of NIPT-positive and -negative cases were 140 and 6851, respectively. Invasive testing using amniocentesis/chrionic villus sampling was performed in 126 NIPT-positive cases, whereas for remaining 14 cases, confirmatory testing could not be performed because of intrauterine fetal death or other reasons. Conventional karyotyping of amniocentesis/chorionic villus samples confirmed trisomies 21, 18 and 13 in 70, 34 and 9 cases, respectively. The positive predictive value was 95.9% (70/73) for trisomy 21, 81.0% (34/42) for trisomy 18 and 81.8% (9/11) for trisomy 13, respectively. Of the 5483 women who tested NIPT negative for birth outcome was available, only one false-negative case of non-mosaic trisomy 18 was found. This false-negative case had a fetal fraction of 6.06%, and z-scores of chromosomes 21, 18 and 13 were −1.867, 2.928 and −1.744, respectively.
Figure 3 depicts fetal cell-free DNA fractions in pregnant women carrying fetuses with different trisomies. Median fetal fraction values for trisomies 21 (n=70), 18 (n=35) and 13 (n=9) were 13.6% (10.2–18.0%), 11.0% (8.1–15.1%) and 8.0% (6.5–10.1%), respectively, although the fetal fraction of NIPT-negative cases (n=6850) was 13.7%. In the cases with trisomy 13 or 18, the fetal fractions in maternal plasma were significantly less than that of the NIPT-negative cases by Tukey's HSD (analysis of variance test) analysis (P=0.004 and P=0.04, respectively). In contrast, no significant differences between NIPT-negative and trisomy 21-positive cases were found (P=0.9993).
The relationship of z-score for trisomies 21, 18 and 13 and negative samples with fetal DNA fractions is presented in Figure 4. It shows a positive correlation between the z-score of trisomies 13, 18 and 21 and fetal fractions. Later gestational age often results in higher positivity rate because of improved classification driven by increased fetal fraction and additional risk factors. Our study, however, did not control positivity rate based on gestational age, instead all the samples between gestational age of 10 and 20 weeks were considered.
Discussion
In this multicenter prospective cohort study, a total of 6993 women among 7740 high-risk women who underwent NIPT were included. Here we show that the fetal fraction in negative and trisomy 21-positive samples by NIPT were not statistically different (R2=0.02). Trisomy 18- or 13-positive samples, by contrast, had significantly lower fetal fractions compared with aneuploidy-negative samples. Because fetal fraction is an important quality metric for aneuploidy detection by NIPT, the differential status of specific chromosome aneuploidy may affect the diagnostic accuracy of the test.
In our experience, the fetal DNA fraction between 10 and 20 weeks’ gestation showed no significant correlation with gestational age, maternal weight and height, or BMI; in contrast to an earlier report.17 Shi et al.18 observed an overall positive trend for fetal fractions between the first and second trimester, with 59% of pregnancies showing an increase, 17% showing no change and 24% showing a decrease. However, another study reported that between 10 and 22 weeks gestational age, there was no statistical difference in fetal fraction.19
Although circulating DNA in healthy women derives mainly from hematopoietic cells undergoing apoptosis,20 in obese pregnant women, it partly derives from apoptotic and necrotic cells of adipose and stromal vascular tissues.21 Our data showed an overall trend towards a slight decrease in fetal fraction in pregnant women who weighed 34 kg (fetal fraction 34.8%) to 115 kg (fetal fraction 6.0%). A similar correlation was observed by Ashoor et al.13 who reported that the median fetal fraction was 11.7% in women who weighed 60 kg, but this decreased to 3.9% in women who weighed 160 kg. They also reported that the estimated proportion with fetal fraction below 4% increased with maternal weight from 0.7% at 60 kg to 7.1% at 100 kg.13
In 0.26% of the patients (6975/6993), NIPT failed because of insufficient fetal DNA or other technical reason. This failure rate is lower compared with that previously reported,2 although all the blood samples were sent to the United States from Japan. Of the 6993 test-negative patients, birth outcome data was available in 5483 cases, and it identified one false-negative trisomy 18, indicating a false-negative rate of <0.1%.
In the samples positive for trisomies 13 and 18, the fetal fractions were significantly lower compared with that of the NIPT-negative cases (P=0.004 and 0.04, respectively), indicating that different fetal aneuploidies have varied effects on the fetal DNA fraction, depending on the affected chromosome, a finding similar to an earlier report.17 We postulate that smaller placental size and IUGR observed with trisomy 13 and 18 might be contributing to lower observed fetal fraction. It is possible that slow cell cycle speed in trophoblast cells in trisomies 13 and 18 affects the low fetal fraction. In contrast, the trisomy 21-positive samples had fetal fractions similar to that of NIPT-negative samples. Taglauer et al.9 reported that compared with euploid fetuses, those with trisomy 21 have an increased fetal fraction.9 This apparent higher fetal fractions in samples positive for trisomy 21 may be one of the reasons that test performance for trisomy 21 is better than that of trisomy 13 or 18.8
At the end of March 2015, NIPT was carried out at 50 institutions, of which 46 were participants in the Japan NIPT consortium. For further research, assessment of fetal DNA fraction at early gestational age can be important not only in screening for fetal aneuploidy but also for prediction of many pregnancy complications.22 Our prospective nationwide data of NIPT performance in Japan will be helpful to define the accuracy in a larger scale study. Such a study can detect associations between NIPT result and pregnancy complications (pregnancy-induced hypertension, preeclampsia, fetal growth restriction and preterm birth), and neonatal and long-term prognosis.
References
Chitty, L. S. & Bianchi, D. W. Noninvasive prenatal testing: the paradigm is shifting rapidly. Prenat. Diagn. 33, 511–513 (2013).
Palomaki, G. E., Deciu, C., Kloza, E. M., Lambert-Messerlian, G. M., Haddow, J. E., Neveux, L. M. et al. DNA sequencing of maternal plasma reliably identifies trisomy 18 and trisomy 13 as well as Down syndrome: an international collaborative study. Genet. Med. 14, 296–305 (2012).
Bianchi, D. W., Platt, L. D., Goldberg, J. D., Abuhamad, A. Z., Sehnert, A. J. & Rava, R. P. Genome-wide fetal aneuploidy detection by maternal plasma DNA sequencing. Maternal blood is source to accurately diagnose fetal aneuploidy (MELISSA) Study Group. Obstet. Gynecol. 119, 890–901 (2012).
Porreco, R. P., Garite, T. J., Maurel, K., Marusiak, B ., Obstetrix Collaborative Research Network, Ehrich, M. et al. Noninvasive prenatal screening for fetal trisomies 21, 18, 13 and the common sex chromosome aneuploidies from maternal blood using massively parallel genomic sequencing of DNA. Am. J. Obstet. Gynecol. 211, 365. e1–e12 (2014).
Norton, M. E., Jacobsson, B., Swamy, G. K., Laurent, L. C., Ranzini, A. C., Brar, H. et al. Cell-free DNA analysis for noninvasive examination of trisomy. N. Engl. J. Med. 372, 1589–1597 (2015).
Sago, H. & Sekizawa, A.,, Japan NIPT Consortium Nationwide demonstration project of next-generation sequencing of cell-free DNA in maternal plasma in Japan: 1-year experience. Prenat. Diagn. 35, 1–6 (2015).
Suzumori, N., Ebara, T., Kumagai, K., Goto, S., Yamada, Y., Kamijima, M. et al. Non-specific psychological distress in women undergoing noninvasive prenatal testing because of advanced maternal age. Prenat. Diagn. 34, 1055–1060 (2014).
Bianchi, D. W. & Wilkins-Haug, L. Integration of noninvasive DNA testing for aneuploidy into prenatal care: What has happened since the rubber met the road? Clin. Chem. 60, 78–87 (2014).
Taglauer, E. S., Wilkins-Haug, L. & Bianchi, D. W. Review: cell-free fetal DNA in the maternal circulation as an indication of placental health and disease. Placenta 28, S64–S68 (2014).
Hahn, S. & Huppertz, W. Fetal cells and cell free fetal nucleic acids in maternal blood: new tools to study abnormal placentation? Placenta 26, 515–526 (2005).
Hudecova, I., Sahota, D., Heung, M. M., Jin, Y., Lee, W. S., Leung, T. Y. et al. Maternal plasma fetal DNA fractions in pregnancies with low and high risks for fetal chromosomal aneuploidies. PLoS ONE 9, e88484 (2014).
Kim, S. K., Hannum, G., Geis, J., Tynan, J., Hogg, G., Zhao, C. et al. Determination of fetal DNA fraction from the plasma of pregnant women using sequence read counts. Prenat. Diagn. 35, 810–815 (2015).
Ashoor, G., Syngelaki, A., Poon, L. C. Y., Rezende, J. C. & Nicolaides, K. H. Fetal fraction in maternal plasma cell-free DNA at 11–13 weeks’ gestation: relation to maternal and fetal characteristics. Ultrasound Obstet. Gynecol. 41, 26–32 (2013).
R Development Core Team R: A Language and Environment for Statistical Computing, (R Foundation for Statistical Computing, Vienna, Austria, 2005); ISBN 3-900051-07-0; available at: http://www.R-project.org Accessed 26 February 2015.
Kanda, Y. Investigation of the freely-available easy-to-use software ‘EZR’ (Easy R) for medical statistics. Bone Marrow Transplant. 48, 452–458 (2013).
Fox, J. The R Commander: a basic-statistics graphical user interface to R. J. Stat. Softw. 19, 1–42 (2005).
Rava, R. P., Srinvasan, A., Sehnert, A. J. & Bianchi, D. W. Circulating fetal cell-free DNA fractions differ in autosomal aneuploidies and monosomy X. Clin. Chem. 60, 243–250 (2014).
Shi, X., Zhang, Z., Cram, D. S. & Liu, C. Feasibility of noninvasive prenatal testing for common fetal aneuploidies in an early gestational window. Clin. Chim. Acta 439, 24–28 (2015).
Brar, H., Wang, E., Struble, C., Musci, T. & Norton, M. E. The fetal fraction of cell-free DNA in maternal plasma is not affected by a prior risk of fetal trisomy. J. Matern. Fetal Neonatal Med. 26, 143–145 (2013).
Liu, Y. Y., Chik, K. W., Chiu, R. W., Ho, C. Y. & Lo, Y. M. Predominant hematopoietic origin of cell-free DNA in plasma and serum after sex-mismatched bone marrow transplantation. Clin. Chem. 48, 421–427 (2002).
Haghiac, M., Vora, N. L., Basu, S., Johnson, K. L., Presley, L., Bianchi, D. W. et al. Increased death of adipose cells, a path to release cell free DNA into systemic circulation of obese women. Obesity 20, 2213–2219 (2012).
Nicolaides, K. H. Turning the pyramid of prenatal care. Fetal Diagn. Ther. 29, 183–196 (2011).
Acknowledgements
This study was supported by Sequenom Inc. We thank all the members of the Japan NIPT consortium for their thoughtful cooperation on this project. We also thank Mr Susumu Ono, Makoto Inaki and Shunsuke Miyai at GeneTech (Tokyo, Japan) for their help with the data sample analysis. This report was supported by the Grant of National Center for Child Health and Development 24-3, Japan.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Members of the Japan NIPT Consortium Toshiaki Endo1, Akimune Hukushima2, Satoshi Nanba3, Hisao Osada4, Yasuyo Kasai5, Atsushi Watanabe6, Yukiko Katagiri7, Naoki Takesita7, Masaki Ogawa8, Takashi Okai9, Shun-ichiro Izumi10, Haruka Hamanoue11, Kazufumi Haino12, Naoki Hamajima13, Haruki Nishizawa14, Yoko Okamoto15, Hiroaki Nakamura16, Takeshi Kanekawa17, Jun Yoshimatsu18, Shinya Tairaku19, Katsuhiko Naruse20, Hisashi Masuyama21, Maki Hyodo22, Takashi Kaji23, Kazutoshi Maeda24, Keiichi Matsubara25, Masanobu Ogawa26, Toshiyuki Yoshizato27, Takashi Ohba28, Yukie Kawano29 1Department of Obstetrics and Gynecology, Sapporo Medical University School of Medicine, Japan; 2Departments of Obstetrics and Gynecology, Iwate Medical University School of Medicine, Japan; 3Departments of Obstetrics and Gynecology, Saitama Medical University School of Medicine, Japan; 4Department of Obstetrics and Gynecology, Chiba University Graduate School of Medicine, Japan, 5Department of Obstetrics and Gynecology, Japanese Red Cross Medical Center, Japan; 6Division of Clinical Genetics, Nippon Medical School Hospital, Japan; 7Department of Obstetrics and Gynecology, Toho University Omori Medical Center, Japan; 8Perinatal Medical Center, Tokyo Women’s Medical University Hospital, Japan; 9Maternal and Child Health Center, Aiiku Hospital, Tokyo, Japan; 10Department of Obstetrics and Gynecology, Tokai University School of Medicine, Japan; 11Department of Human Genetics, Yokohama City University Graduate School of Medicine, Japan; 12Department of Obstetrics and Gynecology, Niigata University Medical and Dental Hospital, Japan; 13Department of Pediatrics, Nagoya City West Medical Center, Japan; 14Department of Obstetrics and Gynecology, Fujita Health University, Japan; 15Department of Obstetrics, Osaka Medical Center and Research Institute for Maternal and Child Health, Japan; 16Department of Obstetrics, Osaka City General Hospital, Japan; 17Department of Obstetrics and Gynecology, Osaka University Faculty of Medicine, Japan; 18Department of Perinatology and Gynecology, National Cerebral and Cardiovascular Center, Japan; 19Department of Obstetrics and Gynecology, Kobe University Graduate School of Medicine, Japan; 20Department of Obstetrics and Gynecology, Nara Medical University, Japan; 21Okayama University Graduate School of Medicine, Japan; 22Hiroshima University Graduate School of Medicine, Japan; 23The University of Tokushima Faculty of Medicine, Japan; 24Department of Obstetrics and Gynecology, Shikoku Medical Center for Children and Adults, Japan; 25Department of Obstetrics and Gynecology, Ehime University School of Medicine, Japan; 26Department of Obstetrics and Gynecology, Clinical Research Institute, National Hospital Organization Kyushu Medical Center, Japan; 27Center for Maternal, Fetal and Neonatal Medicine, Fukuoka Univerisity Hospital, Japan; 28Department of Obstetrics and Gynecology, Kumamoto University, Kumamoto, Japan; 29Department of Molecular Pathology, Faculty of Medicine, Oita University, Japan
Rights and permissions
About this article
Cite this article
Suzumori, N., Ebara, T., Yamada, T. et al. Fetal cell-free DNA fraction in maternal plasma is affected by fetal trisomy. J Hum Genet 61, 647–652 (2016). https://doi.org/10.1038/jhg.2016.25
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2016.25
This article is cited by
-
Association between cell-free DNA fetal fraction and pregnant character: a retrospective cohort study of 27,793 maternal plasmas
Scientific Reports (2023)
-
Factors affecting low fetal fraction in fetal screening with cell-free DNA in pregnant women: a systematic review and meta-analysis
BMC Pregnancy and Childbirth (2022)
-
Chromosomal phase improves aneuploidy detection in non-invasive prenatal testing at low fetal DNA fractions
Scientific Reports (2022)
-
Fetal fraction evaluation in non-invasive prenatal screening (NIPS)
European Journal of Human Genetics (2019)
-
The Effect of Freezing on Non-invasive Prenatal Testing
Scientific Reports (2019)