Abstract
Pathogenic germline mutations in the folliculin (FLCN) tumor suppressor gene predispose to Birt–Hogg–Dubé (BHD) syndrome, a rare disease characterized by the development of cutaneous hamartomas (fibrofolliculomas), multiple lung cysts, spontaneous pneumothoraces and renal cell cancer. In this study, we report the identification of 13 variants and three polymorphisms in the FLCN gene in 143 Danish patients or families with suspected BHD syndrome. Functional mini-gene splicing analysis revealed that two intronic variants (c.1062+2T>G and c.1177-5_1177-3del) introduced splicing aberrations. Eleven families exhibited the c.1062+2T>G mutation. Combined single nucleotide polymorphism array-haplotype analysis showed that these families share a 3-Mb genomic fragment containing the FLCN gene, revealing that the c.1062+2T>G mutation is a Danish founder mutation. On the basis of in silico prediction and functional splicing assays, we classify the 16 identified variants in the FLCN gene as follows: nine as pathogenic, one as likely pathogenic, three as likely benign and three as polymorphisms. In conclusion, the study describes the FLCN mutation spectrum in Danish BHD patients, and contributes to a better understanding of BHD syndrome and management of BHD patients.
Similar content being viewed by others
Introdution
Birt–Hogg–Dubé (BHD) syndrome (OMIM#135150) is a rare autosomal dominantly inherited condition caused by mutations in the FLCN gene encoding folliculin.1, 2, 3 BHD syndrome predisposes to the development of cutaneous hamartomas (fibrofolliculomas (FF)), multiple lung cysts (LCs), spontaneous pneumothoraces and renal cell cancer (RCC).4 The renal cancers associated with BHD are predominantly of the hybrid oncocytic/chromophobe subtype followed by the clear cell renal carcinomas.5 The phenotype is characterized in particular by multiple FF—predominantly facial and upper body—as up to 85% of all BHD syndrome patients suffer from these benign dermatological papules, which usually do not appear before the age of 20.6, 7, 8 Moreover, the dermatologic findings may be the only presenting signs in patients with BHD syndrome, thus making this recognition vital so that further systemic evaluation can be undertaken.
Currently, there is no international consensus for surveillance of BHD patients; hence the Danish recommendations consist of annually renal imaging either by MRI or ultrasound examination, depending on local competence. The onset of renal imaging is 10 years before the debut of the earliest renal cancer incidence in affected family members. The patients undergo examination by a dermatologist depending on individual requirements. Examination of the lungs is not a routine part of the Danish surveillance program.
Other clinical findings associated with BHD syndrome include a range of benign and malignant tumors, leading to a wide variability in the clinical features and consequently a high risk of underdiagnosing mutation carriers.8, 9 Although BHD syndrome was first described in 1977,1 the FLCN gene encoding folliculin was not identified until 2002.3 The FLCN gene is located on chromosome 17p11.2, contains 14 exons and encodes the 579-amino acid protein folliculin. Folliculin is widely conserved across species, indicating a crucial biological function, and recent data suggest that FLCN is involved in multiple biological pathways including metabolism, autophagy, cell-cell adhesion and apoptosis.10
So far, ~200 families with BHD syndrome caused by a pathogenic FLCN mutation have been reported worldwide,7, 8, 11, 12 and close to 150 unique DNA variants have been identified, including nonsense mutations, missense variants, frameshift mutations, intronic variants as well as large genomic rearrangements, of which ~110 are classified as pathogenic (https://grenada.lumc.nl/LOVD2/shared1/home.php?select_db=FLCN).
During the last decade, screening of patients suffering from FF, spontaneous pneumothoraces, kidney cancer as well as genetic susceptibility for BHD syndrome has been performed in Denmark. This study reports the variants identified in Danish patients with BHD syndrome or BHD syndrome-like symptoms and reveals six novel FLCN variants as well as a Danish founder mutation. Moreover, the study classifies the identified variants based on in silico analysis and functional splicing assays.
Materials and methods
Patients
Patients were referred to genetic counseling if there was a suspicion of familial BHD syndrome or if individuals had one of the following symptoms: FF, multiple LCs, spontaneous pneumothorax (SP) or RCC. In some cases, healthy individuals were included if there was a family history of any of the above symptoms. Following verbal and written consent, blood samples were collected from the proband for FLCN mutational screening. The patient/family history was verified using hospital medical records and pathology reports. The study was approved by the Capital Region Committee on Health Research Ethics (H-2-2014-088).
FLCN screening
Genomic DNA was purified from whole blood using the QIAamp DNA Mini Kit (Qiagen, Hilden, Germany) or ReliaPrep Large Volume HT gDNA Isolation Kit (Promega, Madison, WI, USA) using a Tecan Freedom EVO HSM2.0 Workstation (Tecan, Männedorf, Switzerland) according to the manufacturer’s instructions. FLCN screening was performed using intronic primers flanking each exon and the PCR products were sequenced as recently described.13 Moreover, genomic DNA was examined for large genomic rearrangements by multiplex ligation-dependent probe amplification (MLPA) analysis using the SALSA MLPA P256 FLCN Kit (MRC-Holland, Amsterdam, the Netherlands). Since 2014, the analysis has been performed using targeted next-generation sequencing and a library designed to capture all exons from the FLCN gene as previously described.14 Sequencing was performed on a MiSeq (Illumina, San Diego, CA, USA) to an average depth of at least × 100. Sequencing data were analyzed using Sequence Pilot (JSI medical systems, Ettenheim, Germany), where variants were called if the non-reference base frequency was above 25%. Sequence variants were verified in an independent blood sample by Sanger sequencing. FLCN variants are numbered according to accession number NM_144997.5 following the guidelines from the Human Genome Variation Societsy (http://www.hgvs.org/mutnomen).
In silico analysis
The integrated Alamut Visual software (v.2.6.1) (http://www.interactive-biosoftware.com) was used to predict the pathogenicity of specific protein variants using Align GVGD,15 PolyPhen-216 and SIFT,17 or to predict the effect of intronic variants on the efficiency of splicing using Splice Site Finder,18 GeneSplicer,19 Splice Site Prediction by Neural Network,20 MaxEntScan21 and Human Splicing Finder.22 The analysis was performed using default settings in all predictions. A variation in the splicing prediction of more than 10% in at least two algorithms was considered as having an effect on splicing.23 The frequency of the variants was obtained from the Exome Aggregation Consortium (ExAC) database. Classification of each variant was performed according to the five-tier scheme, where class 5 is pathogenic, class 4 is likely pathogenic, class 3 is uncertain, class 2 is likely benign and class 1 is benign.24
Mini-gene splicing assay
Wild-type FLCN exons 9 and 11 were cloned into the pSPL3 vector (Figure 1) and variations were introduced by DpnI-mediated mutagenesis performed using Phusion High-Fidelity Polymerase according to the accompanying instructions (Finnzymes, Waltham, MA, USA). Wild type and variant constructs were then transfected in duplicate into COS-7 cells and processed as recently described.25, 26, 27
Mini-gene splicing analysis of two FLCN intronic variants. COS-7 cells were transfected with wild type or mutant vectors in duplicate. Total RNA was isolated, RT-PCR analysis was performed, and PCR products were separated by agarose gel electrophoresis and visualized by ethidium bromide staining. The sizes of the DNA marker (M) are indicated to the left. (a, b) The FLCN c.1062+2T>G variant generated a strong band of 177 bp lacking exon 9 and a very weak band of 498 bp using a cryptic splice site 130 bp upstream in intron 9. (c, d) The FLCN c.1177-5_1177-3del variant resulted in one strong band of 177 bp lacking exon 11. All PCR products were verified by Sanger sequencing. M, size marker; SA, splice acceptor site; SD, splice donor site.
SNP chip and identity by descent analysis
Nine individuals carrying the FLCN c.1062+2T>G mutation were genotyped for 934 968 SNPs using the Affymetrix 6.0 SNP chip as recently described.28 To obtain estimates of allele frequencies and linkage disequilibrium, the data were merged with the 180 European individuals from HapMap 3. Only the 724 772 SNPs that overlapped between the data sets were used. In addition, SNPs with a minor allele frequency below 5% and SNPs with missing information above 5% were removed. A simple method of moment estimator from PLINK was used to estimate coefficient of relationship between all pairs of individuals.29 To infer whether the c.1062+2T>G mutation is a founder mutation, we estimated relatedness locally on the genome using RELATE.30 To accommodate linkage disequilibrium in the analysis, we conditioned on the 50 previous single-nucleotide polymorphisms (SNPs). Physical positions were used assuming a uniform recombination rate of 1.3 cM/Mb (centiMorgans per megabase). Because we were only interested in whether the individuals share at least one allele IBD, we ignored the state of sharing two alleles IBD. This approach gave us a probability across chromosome 17 (location of the FLCN gene) of two individuals sharing at least one allele IBD. If the mutation is a founder mutation then all individuals should be related in the region around the mutation. The mean probability of sharing at least one allele IBD was estimated by taking the mean posterior probability of IBD sharing between all pairs of individuals for each site. We used a cut-off of 0.95 for the mean IBD sharing to infer the region shared IBD between all pairs of individuals.
Results
Since 2006, we have performed genetic FLCN screening of potential BHD syndrome individuals. Up until 1 November, 2015, 143 individuals or families have been examined and currently we have identified 13 different FLCN variants in 31 families, of which 6 are novel (Table 1). Moreover, we have identified three common polymorphisms (Table 2).
We identified two novel frameshift mutations (c.158delA and c.233delA) as well as four known frameshift mutations (c.57_58delCT, c.584delG, c.1285delC and c.1285dupC) in the FLCN gene. The c.57_58delCT, c.233delA, c.584delG and c.1285dupC mutations were identified in one family each, the c.158delA mutation in two families, whereas the c.1285delC mutation was identified in seven distinct families. Moreover, in one family we identified a deletion of whole exon 9 (c.872-?_1062+?del). The mutations were all identified in families suffering from one or up to four of the symptoms of BHD syndrome, of which FF and spontaneous pneumothoraces were most common.
We also identified two intronic variants (c.1062+2T>G and c.1177-5_1177-3del) in the FLCN gene, present in the vicinity of exon 9 and 11, respectively. The c.1062+2T>G variant was identified in 11 putative BHD syndrome families, whereas the c.1177-5_1177-3del variant was identified in one family with RCC. FF was present in all of the 11 families with the c.1062+2T>G variant, whereas LC/SP and RCC was observed less frequently. The potential pathogenicity of the two intronic variants was investigated using five different in silico splicing prediction programs, which predict changes in splice-site strength (Table 3). The c.1062+2T>G variant was predicted to destroy the SD site by all five programs, whereas the c.1177-5_1177-3del variant was suggested by two programs to decrease the strength of the SA site by more than 10%. The functional effects of both variants were subsequently examined by mini-gene splicing assays cloning either exon 9 or 11 and surrounding intronic sequences into the pSPL3 vector (Figures 1a and c). In accordance with the in silico results, both variants revealed the presence of alternative gel bands compared with the corresponding wild types. The wild- type FLCN exon 9 construct generated a transcript comprising the expected 368 bp as well as a very weak band of 177 bp lacking exon 9. The c.1062+2T>G mutant yielded a strong band of 177 bp lacking exon 9 and a very weak band of 498 bp (Figure 1b). Sequencing revealed that the 498 bp band included an additional 130 bp of intron 9 using a cryptic splice site generating an out-of-frame mutation. Wild-type FLCN exon 11 generated two transcripts, one at the expected 301 bp and one band of 177 bp, indicating partial skipping of exon 11. The c.1177-5_1177-3del variant resulted in only one band of 177 bp lacking exon 11 (Figure 1d). We conclude that the c.1062+2T>G and c.1177-5_1177-3del mutations both result in aberrant splicing.
As the c.1062+2T>G mutation was identified in 11 seemingly unrelated Danish families, we speculated that this mutation could be a Danish founder mutation. To investigate whether this was the case, we performed SNP array and IBD analysis based on one individual from 9 of the 11 families. If the FLCN mutation is a founder mutation, we would expect the carriers to share a common haplotype around the FLCN gene that is shared IBD. We compared the pairwise IBD sharing between each pair of individuals, both for the whole genome and locally at the FLCN locus. As shown in Figure 2, the coefficient of relationship based on the genome-wide data is estimated to be very low among all nine families, indicating that they are not closely related. However, when estimating the IBD sharing locally at the FLCN locus, we see that they are all related with very high probability. Accordingly, we infer that the FLCN c.1062+2T>G mutation is a Danish founder mutation. The mean IBD sharing between all pairs of individuals across chromosome 17 is shown in Supplementary Figure 1. From this, the region shared IBD among all nine individuals is suggested to be 2.95 Mb long.
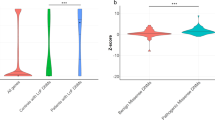
Global and local relatedness among members from nine different Danish families harboring the FLCN c.1062+2T>G intronic mutation. Each individual is indicated by their family ID (Table 1). The plot shows both the overall coefficient of relatedness between pairs of individuals based on all markers across the genome (see the upper triangle) and the local probability of relatedness at the FLCN locus (lower triangle). None of the families are closely related. However, at the FLCN gene, all of the individuals have high probability for sharing at least one allele identity by descent (IBD), which is consistent with a founder mutation.
We moreover identified a novel 21-bp deletion (c.1093_1113del) in exon 10 of the FLCN gene. The variant is an in-frame deletion leading to removal of seven amino acids (p.Ala365_Gly371del) from the FLCN protein. The deletion occurs in a highly conserved region of FLCN forming the second α-helix (Figure 3), suggesting that it is important for FLCN function. The variant was identified in two unrelated families, both of which presented with FF, as well as RCC in one family and spontaneous pneumothoraces and LCs in the other.
Conservation of the second α-helix and first β-sheet in the C-terminal part of the FLCN protein. Human, monkey, rat, mouse, dog, platypus, chicken, frog, tetraodon and zebrafish were aligned. Highly conserved amino acids are shown in purple boxes, whereas conserved amino acids are shown in blue boxes. The Ala365-Gly371 amino acids are marked in brackets. α-helices (red cylinders) and β-sheet (yellow cylinders) are shown at the bottom and is based on data from Nookala et al.42 A full color version of this figure is available at the Journal of Human Genetics journal online.
Furthermore, we identified two missense variants (p.Asp33His and p.Ala90Ser) in the FLCN gene. The p.Asp33His variant was identified in a healthy individual whose daughter died of RCC at the age of 29, whereas the p.Ala90Ser variant was identified in a patient with FF, spontaneous pneumothoraces and LCs. The p.Asp33His variant is reported in two South Asian individuals in the ExAC database, of which one is homozygous. The p.Ala90Ser variant is reported at a frequency of 0.042% in the ExAC database. In silico protein analysis predicted that neither of the variants are pathogenic. Moreover, the p.Ala90Ser variant was identified in a patient with the disease-causing FLCN mutation p.Ala365_Gly371del. Finally, we identified one synonymous variant (p.Ile426Ile) in the FLCN gene, which is reported at a frequency of 0.01% in the ExAC database. The variant was found in an individual with RCC. It is currently unknown whether the variant has any effect on the RNA level.
Discussion
Mutations in the FLCN gene predispose to FF, LCs, spontaneous pneumothoraces and RCC—a syndrome known as BHD. In this study, we report the characterization of FLCN variants identified in 143 Danish patients or families with suspected BHD syndrome.
We identified six different frameshift mutations (c.57_58delCT, c.158delA, c.233delA, c.584delG, c.1285delC and c.1285dupC) in the FLCN gene, which are all classified as pathogenic (class 5). The c.57_58delCT mutation, resulting in a premature stop at codon 35, was found in a family with FF, spontaneous pneumothoraces and RCC. The mutation has recently been described in a French patient with spontaneous pneumothoraces and RCC as well as in a Japanese patient with FF and pneumothoraces.31, 32 Two of the frameshift mutations (c.158delA and c.233delA) are novel. The c.158delA mutation, which introduces a stop codon at position 54, was identified in a large pedigree. The five mutation carriers suffered from bilateral spontaneous pneumothoraces and one family member had fatal bilateral RCC. Moreover, the mutation was identified in an apparently unrelated family with a single affected member suffering from FF, LCs and spontaneous pneumothoraces. The c.233delA mutation, which results in a frameshift and a premature stop at codon 129, was recently found in a single patient with FF. At present, no other family members have been tested. The c.584delG mutation introduces a premature stop at codon 222. This mutation was identified in a family with LCs, spontaneous pneumothoraces and FF. The mutation has previously been described in three BHD syndrome patients with FF and LCs.7, 8 Two of these patients also had spontaneous pneumothoraces or RCC, respectively.8
The deletion or insertion of cytosine at position 1285 has previously been reported to be the cause of BHD syndrome in up to 75% of a cohort of affected families.3, 8, 33 The reported phenotype included the classical symptoms of FF, spontaneous pneumothoraces and RCC. However, other skin lesions (MM, perifollicular fibroma, angiofibroma and basal cell carcinoma) and colorectal lesions were listed among the clinical characteristics of these families. In our cohort, the c.1285delC and c.1285dupC mutations were identified in ~25% of mutation positive families (8/31) and presented with a variety of the classical BHD syndrome symptoms.
We moreover identified a novel deletion of exon nine. The deletion was identified in an individual suffering from LCs and moreover had RCC in the family. The deletion results in a frameshift and a premature stop codon and is therefore classified as pathogenic (class 5). LGRs have previously been described in the literature,34, 35, 36, 37 suggesting that examinations for LGRs in the FLCN gene should be performed routinely in a diagnostic setting.
Approximately 35% of the mutation positive BHD syndrome families in our cohort (11/31) carried the c.1062+2T>G intronic variant. In silico analysis and mini-gene splicing assay in combination with co-segregation data confirmed the pathogenic effect of this intronic mutation (class 5). This mutation was first reported in two apparently unrelated families.7 In one of these families, all five affected members suffered from FF, four also had LCs, three had renal tumors and one had SP. The other family consisted of three affected members all suffering from LCs and renal tumors. On the basis of these data, the c.1062+2T>G mutation was thought to be associated with a higher frequency of renal tumors.7 However, this was not supported by a later study reporting another family with the c.1062+2T>G mutation whose affected members only suffered from FF and LCs.8 Interestingly, in a study characterizing renal tumors in BHD syndrome patients, the c.1062+2T>G mutation was reported in a patient with RCC and FF.31 In the present study, SNP analysis revealed that nine of the Danish families (two families were just recently included in the study and therefore did not participate in the analysis) share a common haplotype IBD and hence are very distant relatives. This finding suggests that the mutation is a founder mutation shared among all the Danish c.1062+2T>G carriers. Intriguingly, we are aware that two of the families have relatives (Danish immigrants) in the USA and that they have also been tested for the mutation. However, whether these families participated in the previous studies of Schmidt et al.7 and Toro et al.8 is currently unknown. The phenotype of the carriers of the Danish founder mutation is predominantly characterized by FF. Although carriers also displayed a broad spectrum of the other classical BHD syndrome symptoms (RCC, LCs and spontaneous pneumothoraces), only 3 out of the 11 families suffered renal tumors. Hence, in line with the findings of Toro et al.8, we found no evidence of an association between the Danish founder mutation and a higher frequency of renal tumors. Moreover, MM was present in one family, whereas colon cancer was observed in another family as recently described.38
We also identified another intronic variant in the FLCN gene, namely a 3-bp deletion (c.1177-5_1177-3del) in intron 10 located just before the consensus SA site. The variant was found in two sisters with bilateral RCC. In silico and mini-gene splicing analyses showed that the variant leads to complete skipping of exon 11. The variant has previously been described in six BHD syndrome families with various classical BHD syndrome symptoms (FF, LCs and spontaneous pneumothoraces),32, 36, 39, 40, 41 and has been shown to co-segregate with the disease.40 On the basis of all the data, we classify the variant as pathogenic (class 5).
A novel in-frame deletion, c.1093_1113del, leading to removal of seven amino acids (p.Ala365_Gly371del) from the FLCN protein was found in two apparently unrelated Danish families. In one family, all mutation carriers suffered from FF and two members also had bilateral RCC of the chromophobe subtype. The other family consisted of only one affected member suffering from FF, LCs and spontaneous pneumothoraces. The p.Ala365_Gly371del variant deletes seven amino acids in the second α-helix, which is essential for maintain the proper structure of the C-terminal halves of FLCN.42 The variant has currently not been examined functionally, but several in-frame deletions in FLCN have previously been shown to disrupt the protein stability in vitro.43 With this in mind and due to the clear co-segregation in one of the families, we classify this novel mutation as likely pathogenic (class 4).
Finally, we identified two novel missense mutations (p.Asp33His and p.Ala90Ser) and one synonymous variant (p.Ile426Ile) in the FLCN gene. The p.Asp33His variant was identified in an individual with no BHD syndrome symptoms. She was tested as her daughter suffered from RCC and died at the age of 29. Homozygous loss of FLCN causes embryonic lethality in mice44 and as the p.Asp33His variant has been reported in homozygote state, we classify this variant as likely benign (class 2). The p.Ala90Ser variant was identified in a patient with the likely pathogenic p.Ala365_Gly371del mutation and the p.Ile426Ile synonymous variant was identified in a patient with RCC and no other classical BHD symptoms. Both variants are classified as likely benign (class 2).
In conclusion, we have identified 13 variants and 3 polymorphism in the FLCN gene in Danish patients with suspected BHD syndrome, 6 of which are novel. Of these variants, 6 frameshift mutations, 1 large genomic rearrangement, and 2 intronic mutations are classified as pathogenic, whereas 1 in-frame deletion is classified as likely pathogenic. Two missense variants and one synonymous variant are classified as likely benign, whereas two intronic variants and one synonymous variant are classified as polymorphisms. Moreover, SNP array analysis revealed that the c.1062+2T>G intronic variant is a Danish founder mutation. The findings presented here extend our knowledge of the FLCN mutation spectrum.
References
Birt, A. R., Hogg, G. R. & Dube, W. J. Hereditary multiple fibrofolliculomas with trichodiscomas and acrochordons. Arch. Dermatol. 113, 1674–1677 (1977).
Khoo, S. K., Bradley, M., Wong, F. K., Hedblad, M. A., Nordenskjold, M. & Teh, B. T. Birt–Hogg–Dube syndrome: mapping of a novel hereditary neoplasia gene to chromosome 17p12-q11.2. Oncogene 20, 5239–5242 (2001).
Nickerson, M. L., Warren, M. B., Toro, J. R., Matrosova, V., Glenn, G., Turner, M. L. et al. Mutations in a novel gene lead to kidney tumors, lung wall defects, and benign tumors of the hair follicle in patients with the Birt–Hogg–Dube syndrome. Cancer Cell. 2, 157–164 (2002).
Schmidt, L. S. & Linehan, W. M. Molecular genetics and clinical features of Birt–Hogg–Dube syndrome. Nat. Rev. Urol. 12, 558–569 (2015).
Pavlovich, C. P., Walther, M. M., Eyler, R. A., Hewitt, S. M., Zbar, B., Linehan, W. M. et al. Renal tumors in the Birt–Hogg–Dube syndrome. Am. J. Surg. Pathol. 26, 1542–1552 (2002).
Houweling, A. C., Gijezen, L. M., Jonker, M. A., van Doorn, M. B., Oldenburg, R. A., van Spaendonck-Zwarts, K. Y. et al. Renal cancer and pneumothorax risk in Birt–Hogg–Dube syndrome; an analysis of 115 FLCN mutation carriers from 35 BHD families. Br. J. Cancer 105, 1912–1919 (2011).
Schmidt, L. S., Nickerson, M. L., Warren, M. B., Glenn, G. M., Toro, J. R., Merino, M. J. et al. Germline BHD-mutation spectrum and phenotype analysis of a large cohort of families with Birt–Hogg–Dube syndrome. Am. J. Hum. Genet. 76, 1023–1033 (2005).
Toro, J. R., Wei, M. H., Glenn, G. M., Weinreich, M., Toure, O., Vocke, C. et al. BHD mutations, clinical and molecular genetic investigations of Birt–Hogg–Dube syndrome: a new series of 50 families and a review of published reports. J. Med. Genet. 45, 321–331 (2008).
Menko, F. H., van Steensel, M. A., Giraud, S., Friis-Hansen, L., Richard, S., Ungari, S. et al. Birt–Hogg–Dube syndrome: diagnosis and management. Lancet Oncol. 10, 1199–1206 (2009).
Schmidt, L. S. & Linehan, W. M. Clinical features, genetics and potential therapeutic approaches for Birt–Hogg–Dube syndrome. Expert Opin. Orphan. Drugs 3, 15–29 (2015).
Leter, E. M., Koopmans, A. K., Gille, J. J., van Os, T. A., Vittoz, G. G., David, E. F. et al. Birt–Hogg–Dube syndrome: clinical and genetic studies of 20 families. J. Invest. Dermatol. 128, 45–49 (2008).
Woodward, E. R., Ricketts, C., Killick, P., Gad, S., Morris, M. R., Kavalier, F. et al. Familial non-VHL clear cell (conventional) renal cell carcinoma: clinical features, segregation analysis, and mutation analysis of FLCN. Clin. Cancer Res. 14, 5925–5930 (2008).
Dandanell, M., Friis-Hansen, L., Sunde, L., Nielsen, F. C. & Hansen, T. V. Identification of 3 novel VHL germ-line mutations in Danish VHL patients. BMC Med. Genet. 13, 54 (2012).
Jonson, L., Ahlborn, L. B., Steffensen, A. Y., Djursby, M., Ejlertsen, B., Timshel, S. et al. Identification of six pathogenic RAD51C mutations via mutational screening of 1228 Danish individuals with increased risk of hereditary breast and/or ovarian cancer. Breast Cancer Res. Treat. 155, 215–222 (2016).
Tavtigian, S. V., Deffenbaugh, A. M., Yin, L., Judkins, T., Scholl, T., Samollow, P. B. et al. Comprehensive statistical study of 452 BRCA1 missense substitutions with classification of eight recurrent substitutions as neutral. J. Med. Genet. 43, 295–305 (2006).
Adzhubei, I. A., Schmidt, S., Peshkin, L., Ramensky, V. E., Gerasimova, A., Bork, P. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Ng, P. C. & Henikoff, S. SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res. 31, 3812–3814 (2003).
Shapiro, M. B. & Senapathy, P. RNA splice junctions of different classes of eukaryotes: sequence statistics and functional implications in gene expression. Nucleic Acids Res. 15, 7155–7174 (1987).
Pertea, M., Lin, X. & Salzberg, S. L. GeneSplicer: a new computational method for splice site prediction. Nucleic Acids Res. 29, 1185–1190 (2001).
Reese, M. G., Eeckman, F. H., Kulp, D. & Haussler, D. Improved splice site detection in genie. J. Comput. Biol. 4, 311–323 (1997).
Yeo, G. & Burge, C. B. Maximum entropy modeling of short sequence motifs with applications to RNA splicing signals. J. Comput. Biol. 11, 377–394 (2004).
Desmet, F. O., Hamroun, D., Lalande, M., Collod-Beroud, G., Claustres, M. & Beroud, C. Human splicing finder: an online bioinformatics tool to predict splicing signals. Nucleic Acids Res. 37, e67 (2009).
Thery, J. C., Krieger, S., Gaildrat, P., Revillion, F., Buisine, M. P., Killian, A. et al. Contribution of bioinformatics predictions and functional splicing assays to the interpretation of unclassified variants of the BRCA genes. Eur. J. Hum. Genet. 19, 1052–1058 (2011).
Richards, S., Aziz, N., Bale, S., Bick, D., Das, S., Gastier-Foster, J. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–424 (2015).
Petersen, S. M., Dandanell, M., Rasmussen, L. J., Gerdes, A. M., Krogh, L. N., Bernstein, I. et al. Functional examination of MLH1, MSH2, and MSH6 intronic mutations identified in Danish colorectal cancer patients. BMC Med. Genet. 14, 103 (2013).
Steffensen, A. Y., Dandanell, M., Jonson, L., Ejlertsen, B., Gerdes, A. M., Nielsen, F. C. et al. Functional characterization of BRCA1 gene variants by mini-gene splicing assay. Eur. J. Hum. Genet. 22, 1362–1368 (2014).
Ahlborn, L. B., Dandanell, M., Steffensen, A. Y., Jonson, L., Nielsen, F. C. & Hansen, T. V. Splicing analysis of 14 BRCA1 missense variants classifies nine variants as pathogenic. Breast Cancer Res. Treat. 150, 289–298 (2015).
Hansen, T. V., Vikesaa, J., Buhl, S. S., Rossing, H. H., Timmermans-Wielenga, V. & Nielsen, F. C. High-density SNP arrays improve detection of HER2 amplification and polyploidy in breast tumors. BMC Cancer 15, 35 (2015).
Purcell, S., Neale, B., Todd-Brown, K., Thomas, L., Ferreira, M. A., Bender, D. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Albrechtsen, A., Sand, K. T., Moltke, I., van Overseem, H. T., Nielsen, F. C. & Nielsen, R. Relatedness mapping and tracts of relatedness for genome-wide data in the presence of linkage disequilibrium. Genet. Epidemiol. 33, 266–274 (2009).
Benusiglio, P. R., Giraud, S., Deveaux, S., Mejean, A., Correas, J. M., Joly, D. et al. Renal cell tumour characteristics in patients with the Birt–Hogg–Dube cancer susceptibility syndrome: a retrospective, multicentre study. Orphanet. J. Rare Dis. 9, 163 (2014).
Furuya, M., Yao, M., Tanaka, R., Nagashima, Y., Kuroda, N., Hasumi, H. et al. Genetic, epidemiologic and clinicopathologic studies of Japanese Asian patients with Birt–Hogg–Dube syndrome. Clin Genet. (e-pub ahead of print 25 May 2016. doi:10.1111/cge.12807).
Khoo, S. K., Giraud, S., Kahnoski, K., Chen, J., Motorna, O., Nickolov, R. et al. Clinical and genetic studies of Birt–Hogg–Dube syndrome. J. Med. Genet. 39, 906–912 (2002).
Benhammou, J. N., Vocke, C. D., Santani, A., Schmidt, L. S., Baba, M., Seyama, K. et al. Identification of intragenic deletions and duplication in the FLCN gene in Birt–Hogg–Dube syndrome. Genes Chromosomes Cancer 50, 466–477 (2011).
Ding, Y., Zhu, C., Zou, W., Ma, D., Min, H., Chen, B. et al. FLCN intragenic deletions in Chinese familial primary spontaneous pneumothorax. Am. J. Med. Genet. A 167A, 1125–1133 (2015).
Kunogi, M., Kurihara, M., Ikegami, T. S., Kobayashi, T., Shindo, N., Kumasaka, T. et al. Clinical and genetic spectrum of Birt–Hogg–Dube syndrome patients in whom pneumothorax and/or multiple lung cysts are the presenting feature. J. Med. Genet. 47, 281–287 (2010).
Sempau, L., Ruiz, I., Gonzalez-Moran, A., Susanna, X. & Hansen, T. V. New mutation in the Birt–Hogg–Dube gene. Actas Dermosifiliogr. 101, 637–640 (2010).
Boman, P. S., Ousager, L. B., Friis-Hansen, L., Hansen, T. V., Broesby-Olsen, S. & Gerdes, A. M. Is colorectal neoplasia part of the Birt–Hogg–Dube syndrome? J. Gastroenterol. Hepatol. Res. 3, 1039–1042 (2014).
Johannesma, P. C., van den Borne, B. E., Gille, J. J., Nagelkerke, A. F., van Waesberghe, J. T., Paul, M. A. et al. Spontaneous pneumothorax as indicator for Birt–Hogg–Dube syndrome in paediatric patients. BMC Pediatr. 14, 171 (2014).
Kunogi, O. M., Yae, T., Nagashima, O., Hirai, S., Kumasaka, T. & Iwase, A. Pneumothorax developing for the first time in a 73-year-old woman diagnosed with Birt–Hogg–Dube syndrome. Intern. Med. 52, 2453–2455 (2013).
Lim, D. H., Rehal, P. K., Nahorski, M. S., Macdonald, F., Claessens, T., Van, G. M. et al. A new locus-specific database (LSDB) for mutations in the folliculin (FLCN) gene. Hum. Mutat. 31, E1043–E1051 (2010).
Nookala, R. K., Langemeyer, L., Pacitto, A., Ochoa-Montano, B., Donaldson, J. C., Blaszczyk, B. K. et al. Crystal structure of folliculin reveals a hidDENN function in genetically inherited renal cancer. Open Biol. 2, 120071 (2012).
Nahorski, M. S., Reiman, A., Lim, D. H., Nookala, R. K., Seabra, L., Lu, X. et al. Birt–Hogg–Dube syndrome-associated FLCN mutations disrupt protein stability. Hum. Mutat. 32, 921–929 (2011).
Hasumi, Y., Baba, M., Ajima, R., Hasumi, H., Valera, V. A., Klein, M. E. et al. Homozygous loss of BHD causes early embryonic lethality and kidney tumor development with activation of mTORC1 and mTORC2. Proc. Natl Acad. Sci. USA 106, 18722–18727 (2009).
van Steensel, M. A., Verstraeten, V. L., Frank, J., Kelleners-Smeets, N. W., Poblete-Gutierrez, P., Marcus-Soekarman, D. et al. Novel mutations in the BHD gene and absence of loss of heterozygosity in fibrofolliculomas of Birt–Hogg–Dube patients. J. Invest. Dermatol. 127, 588–593 (2007).
Kawasaki, H., Sawamura, D., Nakazawa, H., Hattori, N., Goto, M., Sato-Matsumura, K. C. et al. Detection of 1733insC mutations in an Asian family with Birt–Hogg–Dube syndrome. Br. J. Dermatol. 152, 142–145 (2005).
Lamberti, C., Schweiger, N., Hartschuh, W., Schulz, T., Becker-Wegerich, P., Kuster, W. et al. Birt–Hogg–Dube syndrome: germline mutation in the (C)8 mononucleotide tract of the BHD gene in a German patient. Acta Derm. Venereol. 85, 172–173 (2005).
Acknowledgements
We thank Stine Østergaard and Susanne Smed for excellent laboratory assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Rossing, M., Albrechtsen, A., Skytte, AB. et al. Genetic screening of the FLCN gene identify six novel variants and a Danish founder mutation. J Hum Genet 62, 151–157 (2017). https://doi.org/10.1038/jhg.2016.118
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2016.118
This article is cited by
-
Exons 1–3 deletion in FLCN is associated with increased risk of pneumothorax in Chinese patients with Birt-Hogg-Dubé syndrome
Orphanet Journal of Rare Diseases (2023)
-
Systematic analysis of CNGA3 splice variants identifies different mechanisms of aberrant splicing
Scientific Reports (2023)
-
Splice-site mutation causing partial retention of intron in the FLCN gene in Birt-Hogg-Dubé syndrome: a case report
BMC Medical Genomics (2018)
-
Birt–Hogg–Dubé Syndrome: A Review of Dermatological Manifestations and Other Symptoms
American Journal of Clinical Dermatology (2018)
-
Characterization of a splice-site mutation in the tumor suppressor gene FLCN associated with renal cancer
BMC Medical Genetics (2017)