Abstract
Many consortia and international projects have investigated the human genetic variation of a large number of ethno-geographic groups. However, populations with peculiar genetic features, such as the Turkish population, are still absent in publically available datasets. To explore the genetic predisposition to health-related traits of the Turkish population, we analyzed 34 genes associated with different health-related traits (for example, lipid metabolism, cardio-vascular diseases, hormone metabolism, cellular detoxification, aging and energy metabolism). We observed relevant differences between the Turkish population and populations with non-European ancestries (that is, Africa and East Asia) in some of the investigated genes (that is, AGT, APOE, CYP1B1, GNB3, IL10, IL6, LIPC and PON1). As most complex traits are highly polygenic, we developed polygenic scores associated with different health-related traits to explore the genetic diversity of the Turkish population with respect to other human groups. This approach showed significant differences between the Turkish population and populations with non-European ancestries, as well as between Turkish and Northern European individuals. This last finding is in agreement with the genetic structure of European and Middle East populations, and may also agree with epidemiological evidences about the health disparities of Turkish communities in Northern European countries.
Similar content being viewed by others
Introduction
The investigation of human genetic variation can facilitate an understanding of health disparities among ancestry groups and the development of effective preventive strategies.1, 2, 3 International projects about human genetic variation, such as the Human Genome Diversity Project,4 the HapMap Project5 and the 1000 Genomes Project,6 have provided a vast amount of data that have been used to explore the effects of genetic diversity in determining a population predisposition to health-related traits.7, 8, 9 However, some populations with peculiar genetic features are not included in these publically available datasets. The Turkish population is one of these missing populations. Because the Anatolian peninsula links the Middle East, Europe and Asia, and because there have been several population migrations in ancient and recent human history,10 the Turkish population presents genetic features that are not present in other human populations. Different studies have investigated the genetic diversity of the Turkish population using molecular analyses on mitochondrial DNA,11 Y-chromosome12 and autosomal loci.13, 14 The outcomes of these investigations highlighted that the genetic pool of the Turkish population comprises an admixture of European, Middle East and Central Asian components. To the best of our knowledge, no studies have explored the genetic architecture of Turkish population to compare the genetic predisposition to health-related traits of this population with respect to other ancestry groups. Recent studies demonstrated that approaches based on population genetics can be used to investigate genetic predisposition to complex traits.15, 16 They provided information about the role human history plays in shaping the genetic diversity of disease-associated loci17 and the differences in genetic predisposition to complex traits among human populations.18 Moreover, genetic studies will derive an awareness of the role of genetics in health and help to stratify disease risks and target interventions to achieve health promotion goals in a given population.19, 20
The present study was developed to analyze the genetic diversity of disease-associated loci in the Turkish population, comparing its genetic diversity with other continental groups via single-locus and multiple-locus analysis. This approach can be used to identify the disease risk in each human population and to stratify individuals according to risk groups, improving our capacity to prevent diseases and detect them early, in accordance with the aims of the GENAR Institute for Public Health and Genomics Research.21 Specifically, we investigated the genetic diversity of 34 independent genes associated with multiple health-related traits (for example, lipid metabolism, cardio-vascular diseases, hormone metabolism, cellular detoxification, aging and energy metabolism) in different human populations. We explored the genetic differences of these loci between the Turkish population and the reference samples of the 1000 Genomes Project. We verified whether these genetic differences are due to the pressures of natural selection or stochastic demographic events. We performed a clustering analysis to check whether the genetic variation of these disease-associated loci are in agreement with those reported in previous studies about the Turkish population. Finally, because most of health-related traits are highly polygenic,22 we developed different polygenic diversity scores associated with specific health-related traits and verified the significant differences of Turkish subjects with respect to individuals with African, American, East Asian and European ancestries. To the best of our knowledge, this is the first investigation that applied polygenic score analysis to explore genetic diversity among human populations. Given this study design, we observed not only significant differences between Turkish and non-European populations in single-locus analysis, but also significant differences between Turkish and Northern European populations in polygenic-score analysis.
Materials and Methods
Samples
All the procedures used in this study conformed to the tenets of the Declaration of Helsinki, and the appropriate institutional ethics committee approval was obtained. Participants were recruited in the seven largest cities (Istanbul, Ankara, Izmir, Bursa, Adana, Antalya and Samsun) within five regions of Turkey. All of the participants voluntarily applied to the GENAR Biotechnology and Molecular Genetics, Research and Application Laboratories, to enroll in a preventive health-care intervention programme20 provided via participating physicians. The study population included unrelated Turkish participants in apparently healthy conditions. Genetic tests were performed after obtaining informed consent. Genes and single nucleotide polymorphisms (SNPs) were selected according to their role in major metabolic and disease-relevant pathways (for example, lipid and glucose metabolism, single carbon metabolism, blood pressure homeostasis and inflammation response). The screened genes were grouped according to their contribution to complex disease. Accordingly, different disease-related test packages, which contain relevant combinations of SNPs, were generated. The volunteers could choose one or more of those packages for genetic testing. This comprises the main reason for the differences among the number of typed individual per variant.
Genotyping
Genomic DNAs were isolated from buccal swabs or whole-blood samples using an MN DNA isolation kit (Macherey Nagel-Nucleospin, Düren, Germany). SequenomRealSNP software was used to design sequence-specific amplification primers (Metabion, Planegg, Germany) for the multiplex level (details regarding the PCR primers, extend probes and multiplex combinations are available upon the request). The investigated variants were selected on the basis of their involvement in multiple health-related traits (for example, lipid metabolism, cardio-vascular diseases, hormone metabolism, cellular detoxification, aging and energy metabolism), developing the Gentest practice model to screen genetic predisposition to complex diseases of healthy Turkish volunteers. Details about Gentest are available in our previous article.21 Specifically, in the current analysis, we investigated 43 SNPs located in 34 different genes. In Supplementary Table 1, we reported the details about the investigated variants. The amplification conditions were based on the manufacturer’s protocol of the Sequenom MassARRAY platform (Sequenom Inc., San Diego, CA, USA). PCR, post-PCR cleanup, homogeneous mass extend reaction and purification of homogeneous mass extend products using a cation exchange resin are the main steps of the applied protocol. Extended/desalted products were arrayed onto 384-sample SpectroCHIPs, using the Nanodispencer system (Sequenom Inc.). The target chips were analyzed in the matrix-assisted laser desorption/ionization-time-of-flight mass spectrometer of the MassARRAY Compact System (Sequenom Inc.). The analysis was performed using SpectroTYPER software (Sequenom Inc.). Detailed information about genotyping pipeline is reported in our previous study.14
As mentioned in the description of the samples, the differences in the sample sizes among the investigated variants are due to the choices of the volunteer participants for the genetic testing packages. In our genetic investigation, all investigated variants showed call rates greater than 95%.
Data from the 1000 Genomes Project
To compare Turkish data with reference to worldwide populations, we used the genotype information available for the samples of Phase 1 of the 1000 Genomes Project.6 This study comprised 1092 individual samples belonging to 14 human populations with four different ancestry origins. The African group consisted of: African Ancestry in Southwest US (ASW); Luhya in Webuye, Kenya (LWK); and Yoruba in Ibadan, Nigeria (YRI). The American group consisted of: Colombian in Medellin (CLM), Colombia; Mexican Ancestry in Los Angeles (MXL), CA; and Puerto Rican in Puerto Rico (PUR). The East Asian group consisted of: Han Chinese in Beijing, China (CHB); Han Chinese South (CHS); and Japanese in Toyko, Japan (JPT). The European group consisted of: Utah residents with Northern and Western European ancestry (CEU); Finnish from Finland (FIN); British from England and Scotland (GBR); Iberian populations in Spain (IBS); and Toscani in Italia (TSI).
Statistical analysis
We calculated the minor allele frequency for each investigated variant. We then used these frequencies to estimate observed and expected heterozygosity, and to calculate pairwise FST distances between the Turkish population and the populations of the 1000 Genomes Project. To verify the presence of selection signatures in the investigated variants, the Integrated Haplotype Scores (iHS) described by Voight et al.23 were applied. Haplotter (available at http://haplotter.uchicago.edu/) was used to calculate the iHSs in three representative ancestry groups (CEU, CHB+JPT and YRI), using HapMap Phase 2 data. |iHS|>1.5 are considered to be suggestive evidence of signatures of natural selection. We used the genotype information of 43 investigated variants to perform a cluster analysis. Structure 2.3.3 was used to perform this analysis,24 and Distruct 1.1 was used to construct the correspondence plots.25 For each run, 10 000 iterations after a burn-in period of 10 000 iterations were utilized, and cluster coefficients (K) from 2 to 4 were considered. Finally, we defined different diversity scores associated with health-related traits on the basis of the investigated loci. Although all investigated loci are involved in disease-related traits, we grouped them in gene clusters involved in specific traits. DAVID Bioniformatics Resources 6.7 was used to cluster the 34 investigated loci in accordance with different diversity scores.26, 27 Specifically, we considered clustering terms related to ‘GENETIC_ASSOCIATION_DB_DISEASE’ criterion and selected terms where all risk loci (DAVID output: Pop Hit) are included among the loci investigated in the present study (DAVID output: Count). In Supplementary Table 2, we reported the details of the investigated disease-associated clusters. On the basis of the disease-associated clusters, each diversity score was calculated as the sum total of carried minor alleles weighted for the number of variants located in the same gene. We calculated the diversity scores for each Turkish subject and each individual of the 1000 Genomes Project. To verify the differences of risk scores between the Turkish population and the populations of the 1000 Genomes Project, we used Kruskal-Wallis/Dunn’s test, correcting for multiple comparisons.
Results
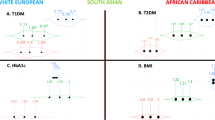
Table 1 shows the minor allele frequencies, along with the expected and observed heterozygosity of the investigated variants in the Turkish population. The genotype frequencies of all investigated variants are in agreement with the Hardy-Weinberg equilibrium. Comparing these frequencies with those reported for populations available in the 1000 Genomes Project, we observed relevant differences (pairwise FST>0.1) between the Turkish population and the non-European populations (that is, populations with African and East Asian ancestries; Supplementary Table 3). Specifically, we observed relevant FST distances between the Turkish population and other worldwide populations for: AGT rs699 (LWK, YRI, CHS and JPT), APOE rs429358 (LWK), CYP1B1 rs1056836 (LWK and YRI), GNB3 rs5443 (ASW, LWK and YRI), IL10 rs1800896 (CHB, CHS and JPT), IL6 rs1800796 (CHB, CHS and JPT), LIPC rs1800588 (LWK, YRI and JPT) and PON1 rs662 (LWK, YRI, CHS and JPT). The iHS analysis indicated cross-ancestry suggestive selection signature for MTHFR rs1801131 (CEU iHS=2.07; CHB+JPT iHS=2.62; YRI iHS=1.54). European-specific suggestive selection signatures were observed for MTHFR rs1801133 (iHS=−1.67), CYP1B1 rs1800440 (iHS=−2.13), TNF rs1800629 (iHS=−2.03). An East Asian-specific suggestive selection signature was observed for LPL rs328 (iHS=1.52). An African-specific suggestive selection signature was observed for MTR rs1805087 (iHS=−1.61). Considering the genetic variation of the investigated loci, the clustering analysis indicated three subpopulations (K) as being the most likely (Figure 1). Specifically, we observed an African cluster (that is, ASW, LWK and YRI), characterized by the ‘black’ component; a cluster comprising Turkish and European populations (CEU, FIN, GBR, IBS, TSI), characterized by the ‘white’ component; an East Asian cluster (that is, CHB, CHS and JPT), characterized by the ‘grey’ component; and an American cluster (that is, CLM, MXL and PUR), characterized by a three-component admixture. However, Turks showed a smaller percentage of the ‘white’ component (65%) than those observed in European populations (CEU: 86%; FIN: 83%; GBR: 82%; IBS: 80% and TSI: 83%). Finally, we explored the diversity between Turkish populations and the other ancestry groups by considering different scores based on the association of investigated loci with health-related traits (Supplementary Table 2). Table 2 reports the diversity scores estimated in the Turkish population and the other investigated ones. Among the 47 investigated risk scores, we observed at least one significant difference in 34 (72%) of them. Significant differences between the Turks and the other populations are observed in East Asians (CHB: 51%, JPT: 49%, CHS: 47%), Africans (YRI: 38%, LWK: 32%, ASW: 17%), Americans (MXL: 11%, CLM: 4%) and Europeans (CEU: 4%, FIN: 4%, GBR, 2%).
Population structure based on disease-associated loci. Each population is represented by a vertical block that is partitioned into 3 grayscale segments (K) that represent the population’s estimated membership fractions in three subpopulations. Black lines separate different populations. The populations are labeled below the figure, with their geographic origin groups above it.
Discussion
The present study explored the genetic diversity of disease-associated loci in the Turkish population, comparing the Turkish variation with variations present in the reference samples of the 1000 Genomes Project. Our results indicate that strong differences are present between Turkish subjects and individuals with African and East Asian ancestry. However, we developed different polygenic scores associated with health-related traits, and we observed significant differences of these diversity scores not only between Turks and non-Europeans, but also between Turkish subjects and individuals with Northern European ancestry.
Considering the pairwise FST distances, we observed strong differences between the Turkish population and subjects with African and East Asian ancestries. Specifically, we observed differences with both these ancestry groups for AGT rs699, GNB3 rs5443, LIPC rs1800588 and PON1 rs662. These relevant differences between the Turkish population and populations with African and East Asian ancestry indicated that these disease-associated loci have a strong variation across human populations. These diversities may have a strong effect on the differences in the predisposition to complex traits among human populations, as suggested by previous studies.28, 29, 30 We also observed specific ‘Turkey vs Africa’ and ‘Turkey vs East Asia’ differences. APOE rs429358 and CYP1B1 rs1056836 showed relevant FST distances only between Turks and Africans, whereas IL10 rs1800896 and IL6 rs1800796 occurred between Turks and East Asians. Regarding both African-differentiated loci (that is, APOE and CYP1B1), the variability of these loci seems to have a relevant effect on the predisposition to certain diseases in populations with African ancestry.31, 32, 33 Regarding the East Asian-differentiated loci (that is, IL10 and IL6), the low frequency of IL10 rs1800896 does not seem to contribute to the disease susceptibility of East Asians,34 whereas IL6 rs1800796 has been demonstrated to be involved in different health-related traits in East Asian populations.35, 36 No relevant differences were observed among European and American populations. While the results of the European population are due to the genetic similarity between Turks and Europeans, the outcomes of the American populations are due to the genetic admixture of European, African and Amerindian components present in the American populations of the 1000 Genomes Project.37
To check whether these allele frequency differences are due to natural selection or human demographic history, we used iHS analysis. None of the ancestry-differentiated loci showed a suggestive signature of natural selection. This outcome is in agreement with previous studies that indicated that large allele frequency differences between human populations are more likely to have occurred by drift during range expansions than by natural selection.38 However, among the investigated loci, we found different signatures of natural selection. MTHFR rs1801131 showed a cross-ancestry signature of natural selection that may be due to selective pressures related to pregnancy loss in individuals with folic acid deficiency, ultraviolet radiation or resistance to malaria.39, 40, 41 Regarding the European-specific selection signatures, we observed another MTHFR variant, along with two variants located in the CYP1B1 and TNF genes. Regarding CYP1B1, a previous study with an independent approach indicated a selection signature in this gene,42 but, to our knowledge, no studies provided a biological explanation for this evidence. For TNF, several studies indicated the protective effect of certain TNF variants against tuberculosis as an explanation of this outcome.43, 44
Our clustering analysis indicated that the genetic structure of disease-associated loci in the Turkish population is more similar to the structures observed in European populations than to those observed in African, East Asian and American populations. However, slight differences are present between Turks and Europeans. As most of the health-related traits are highly polygenic,22 we developed certain polygenic scores on the basis of the investigated loci to explore genetic differences between the Turkish population and the other ancestry groups. To the best of our knowledge, no studies used this kind of approach to explore genetic diversity among human populations. In this analysis, the Turkish population showed several significant differences with regard to the African and Asian populations, in agreement with the differences observed in our FST analysis. However, we also observed significant differences in the pairwise comparisons with Europeans and American populations. As mentioned above, the presence of slight differences between Turkish and American populations is due to the admixture of the European, African and Amerindian components of the American samples of the 1000 Genomes Project.37 Regarding the European populations, the diversity score results strongly suggest slight genetic differences of the Turkish population with respect to the European populations. Specifically, we observed significant differences in the Turkish population with FIN for the ADRB2-PPARG diversity score; GBR for the ADRB3-ADRB2 diversity score; CEU, FIN and GBR for the IL6-APOE diversity score; FIN for the NOS3-GNB3 diversity score; and CEU for the TNF-IL10 diversity score. Considering the geographic distribution of the analyzed European samples, all the significant differences are between the Turkish population and the Northern European samples. This outcome is in agreement with current knowledge about the genetic diversity of European populations: Southern European populations are more similar genetically to the populations of Mediterranean Africa and the Middle East than Northern Europe.45, 46 The strong similarity between Turkish populations and Southern European populations was also confirmed by a recent whole-genome sequencing of 16 Turkish samples.47 In addition, the significant differences observed in the diversity score analysis also indicated differences in the genetic predisposition to health-related traits of the Turkish population with respect to European and non-European populations. These findings may be supported by several epidemiological studies that indicate significant health disparities among different ethnic groups, in which Turkish communities in Northern European countries were included.48, 49, 50, 51
In conclusion, our study on several disease-associated loci indicated significant differences in the Turkish population with respect to European and non-European populations. Furthermore, our analysis, based on polygenic diversity scores, seems to be able to dissect partially the genetic background at the basis of the health disparities among the human populations. Accordingly, further studies of polygenic diversity scores may enhance understanding of the role of human genetic variation in determining health disparities among different ethnic groups. Moreover, our results will be useful in the design of future studies that investigate the contribution of genetic variation to complex diseases in Turkey.
References
Rawlings-Goss, R. A., Campbell, M. C. & Tishkoff, S. A. Global population-specific variation in miRNA associated with cancer risk and clinical biomarkers. BMC Med. Genomics 7, 53 (2014).
Gomez, F., Hirbo, J. & Tishkoff, S. A. Genetic variation and adaptation in Africa: implications for human evolution and disease. Cold Spring Harb. Perspect. Biol. 6, a008524 (2014).
Khoury, M. J., Gwinn, M. L., Glasgow, R. E. & Kramer, B. S. A population approach to precision medicine. Am. J. Prev. Med. 42, 639–645 (2012).
Li, J. Z., Absher, D. M., Tang, H., Southwick, A. M., Casto, A. M. & Ramachandran, S. et al. Worldwide human relationships inferred from genome-wide patterns of variation. Science 319, 1100–1104 (2008).
International HapMap Consortium International HapMap Consortium, Altshuler, D. M., Gibbs, R. A., Peltonen, L., Altshuler, D. M. & Gibbs, R. A. et al. Integrating common and rare genetic variation in diverse human populations. Nature 467, 52–58 (2010).
1000 Genomes Project Consortium 1000 Genomes Project Consortium, Abecasis, G. R., Altshuler, D., Auton, A., Brooks, L. D. & Durbin, R. M. et al. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Polimanti, R., Piacentini, S., Iorio, A., De Angelis, F., Kozlov, A. & Novelletto, A. et al. Haplotype differences for copy number variants in the 22q11.23 region among human populations: a pigmentation-based model for selective pressure. Eur. J. Hum. Genet 23, 116–123 (2015).
Polimanti, R., Iorio, A., Piacentini, S., Manfellotto, D. & Fuciarelli, M. Human pharmacogenomic variation of antihypertensive drugs: from population genetics to personalized medicine. Pharmacogenomics 15, 157–167 (2014).
Polimanti, R., Di Girolamo, M., Manfellotto, D. & Fuciarelli, M. Functional variation of the transthyretin gene among human populations and its correlation with amyloidosis phenotypes. Amyloid 20, 256–262 (2013).
Güvenç, B. Türk Kimliği Kültür Tarihinin Kaynakları (Turkish Identity Sources of Cultural History) 19–52 (Kültür Bakanlığı: Ankara, 1993).
Mergen, H., Oner, R. & Oner, C. Mitochondrial DNA sequence variation in the Anatolian Peninsula (Turkey). J. Genet. 83, 39–47 (2004).
Cinnioglu, C., King, R., Kivisild, T., Kalfoglu, E., Atasoy, S. & Cavalleri, G. L. et al. Excavating Y-chromosome haplotype strata in Anatolia. Hum. Genet. 114, 127–148 (2004).
Hodoglugil, U. & Mahley, R. W. Turkish population structure and genetic ancestry reveal relatedness among Eurasian populations. Ann. Hum. Genet. 76, 128–141 (2012).
Karaca, S., Karaca, M., Cesuroglu, T., Erge, S. & Polimanti, R. Analysis of GSTP1, GSTM1 and GSTT1 genetic variability in Turkish and worldwide populations. Am. J. Hum. Biol. (e-pub ahead of print 16 December 2014).
Daub, J. T., Hofer, T., Cutivet, E., Dupanloup, I., Quintana-Murci, L. & Robinson-Rechavi, M. et al. Evidence for polygenic adaptation to pathogens in the human genome. Mol. Biol. Evol. 30, 1544–1558 (2013).
Laitman, Y., Feng, B. J., Zamir, I. M., Weitzel, J. N., Duncan, P. & Port, D. et al. Haplotype analysis of the 185delAG BRCA1 mutation in ethnically diverse populations. Eur. J. Hum. Genet. 21, 212–216 (2013).
Polimanti, R., Piacentini, S. & Fuciarelli, M. HapMap-based study of human soluble glutathione S-transferase enzymes: the role of natural selection in shaping the single nucleotide polymorphism diversity of xenobiotic-metabolizing genes. Pharmacogenet. Genomics. 21, 665–672 (2011).
Goetz, L. H., Uribe-Bruce, L., Quarless, D., Libiger, O. & Schork, N. J. Admixture and clinical phenotypic variation. Hum. Hered. 77, 73–86 (2014).
Karakachoff, M., Duforet-Frebourg, N., Simonet, F., Le Scouarnec, S., Pellen, N. & Lecointe, S. et al. Fine-scale human genetic structure in Western France. Eur. J. Hum. Genet. (e-pub ahead of print 3 September 2014).
Deelen, P., Menelaou, A., van Leeuwen, E. M., Kanterakis, A., van Dijk, F. & Medina-Gomez, C. et al. Improved imputation quality of low-frequency and rare variants in European samples using the 'Genome of The Netherlands'. Eur. J. Hum. Genet. 22, 1321–1326 (2014).
Cesuroglu, T., Karaca, S. & Erge, S. A practice model for personalized healthcare with a public health genomics perspective. Personalized Medicine. 6, 567–577 (2009).
van der Sijde, M. R., Ng, A. & Fu, J. Systems genetics: From GWAS to disease pathways. Biochim. Biophys. Acta. 1842, 1903–1909 (2014).
Voight, B. F., Kudaravalli, S., Wen, X. & Pritchard, J.K. A map of recent positive selection in the human genome. PLoS Biol. 4, e72 (2006).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Rosenberg, N. A. DISTRUCT: a program for the graphical display of population structure. Molecular Ecology Notes. 4, 137–138 (2004).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13 (2009).
Maisano Delser, P. & Fuselli, S. Human loci involved in drug biotransformation: worldwide genetic variation, population structure, and pharmacogenetic implications. Hum. Genet. 132, 563–577 (2013).
Frazier-Wood, A. C., Manichaikul, A., Aslibekyan, S., Borecki, I. B., Goff, D. C. & Hopkins, P. N. et al. Genetic variants associated with VLDL, LDL and HDL particle size differ with race/ethnicity. Hum. Genet. 132, 405–413 (2013).
Huen, K., Harley, K., Beckman, K., Eskenazi, B. & Holland, N. Associations of PON1 and genetic ancestry with obesity in early childhood. PLoS ONE 8, e62565 (2013).
Atadzhanov, M., Mwaba, M. H., Mukomena, P. N., Lakhi, S., Mwaba, P. & Rayaprolu, S. et al. Frequency of APOE, MTHFR and ACE polymorphisms in the Zambian population. BMC Res. Notes 7, 194 (2014).
Chang, M. H., Yesupriya, A., Ned, R. M., Mueller, P. W. & Dowling, N. F. Genetic variants associated with fasting blood lipids in the U.S. population: Third National Health and Nutrition Examination Survey. BMC Med. Genet. 11, 62 (2010).
Reding, K. W., Chen, C., Lowe, K., Doody, D. R., Carlson, C. S. & Chen, C. T. et al. Estrogen-related genes and their contribution to racial differences in breast cancer risk. Cancer Causes Control 23, 671–681 (2012).
Guo, J., He, Y. H., Chen, F., Jiang, M. H., Gao, S. P. & Su, Y. M. et al. The A to G polymorphism at -1082 of the interleukin-10 gene is rare in the Han Chinese population. Mol. Med. Rep. 6, 894–896 (2012).
UniProt, C. Activities at the Universal Protein Resource (UniProt). Nucleic Acids Res. 42, D191–D198 (2014).
Okada, R., Wakai, K., Naito, M., Morita, E., Kawai, S. & Hamajima, N. et al. Pro-/anti-inflammatory cytokine gene polymorphisms and chronic kidney disease: a cross-sectional study. BMC Nephrol. 13, 2 (2012).
Gravel, S., Zakharia, F., Moreno-Estrada, A., Byrnes, J. K., Muzzio, M. & Rodriguez-Flores, J. L. et al. Reconstructing Native American migrations from whole-genome and whole-exome data. PLoS Genet. 9, e1004023 (2013).
Hofer, T., Ray, N., Wegmann, D. & Excoffier, L. Large allele frequency differences between human continental groups are more likely to have occurred by drift during range expansions than by selection. Ann. Hum. Genet. 73, 95–108 (2009).
Yafei, W., Lijun, P., Jinfeng, W. & Xiaoying, Z. Is the prevalence of MTHFR C677T polymorphism associated with ultraviolet radiation in Eurasia? J. Hum. Genet. 57, 780–786 (2012).
Meadows, D. N., Pyzik, M., Wu, Q., Torre, S., Gros, P. & Vidal, S. M. et al. Increased resistance to malaria in mice with methylenetetrahydrofolate reductase (Mthfr) deficiency suggests a mechanism for selection of the MTHFR 677C>T (c.665C>T) variant. Hum. Mutat. 35, 594–600 (2014).
Jennings, B. A., Willis, G. A., Skinner, J. & Relton, C. L. Genetic selection? A study of individual variation in the enzymes of folate metabolism. BMC Med. Genet. 11, 18 (2010).
Polimanti, R., Piacentini, S., Manfellotto, D. & Fuciarelli, M. Human genetic variation of CYP450 superfamily: analysis of functional diversity in worldwide populations. Pharmacogenomics 13, 1951–1960 (2012).
Correa, P. A., Gomez, L. M., Cadena, J. & Anaya, J. M. Autoimmunity and tuberculosis. Opposite association with TNF polymorphism. J. Rheumatol. 32, 219–224 (2005).
Mobley, J. L. Is rheumatoid arthritis a consequence of natural selection for enhanced tuberculosis resistance? Med. Hypotheses. 62, 839–843 (2004).
Lao, O., Lu, T. T., Nothnagel, M., Junge, O., Freitag-Wolf, S. & Caliebe, A. et al. Correlation between genetic and geographic structure in Europe. Curr. Biol. 18, 1241–1248 (2008).
Sikora, M., Carpenter, M. L., Moreno-Estrada, A., Henn, B. M., Underhill, P. A. & Sanchez-Quinto, F. et al. Population genomic analysis of ancient and modern genomes yields new insights into the genetic ancestry of the Tyrolean Iceman and the genetic structure of Europe. PLoS Genet. 10, e1004353 (2014).
Alkan, C., Kavak, P., Somel, M., Gokcumen, O., Ugurlu, S. & Saygi, C. et al. Whole genome sequencing of Turkish genomes reveals functional private alleles and impact of genetic interactions with Europe, Asia and Africa. BMC Genomics. 15, 963 (2014).
Icks, A., Razum, O., Rosenbauer, J., Bachle, C., Hungele, A. & Monkemoller, K. et al. Lower frequency of insulin pump treatment in children and adolescents of Turkish background with type 1 diabetes: analysis of 21,497 patients in Germany. Diabetes Technol. Ther. 14, 1105–1109 (2012).
de Hoog, M. L., van Eijsden, M., Stronks, K., Gemke, R. J. & Vrijkotte, T. G. Overweight at age two years in a multi-ethnic cohort (ABCD study): the role of prenatal factors, birth outcomes and postnatal factors. BMC Public Health. 11, 611 (2011).
Zubor, P., Caliskan, M., Kajo, K., Soybir, G., Topuzlu, C. & Danko, J. Ethnic disparities in breast cancer between Central Europe Caucasian women of Slavic origin and Middle East Turkish subjects. Neoplasma 61, 110–117 (2014).
Racape, J., De Spiegelaere, M., Dramaix, M., Haelterman, E. & Alexander, S. Effect of adopting host-country nationality on perinatal mortality rates and causes among immigrants in Brussels. Eur. J. Obstet. Gynecol. Reprod. Biol. 168, 145–150 (2013).
Acknowledgements
We are greatly indebted to Dr Serdar Savas, the founder of the GENAR Public Health and Genomics Research Institute, for his great contribution to the development of the Gentest Model and its components. We would also like to thank to Marco De Langen, as well as all the staff members of the GENAR Molecular Biology and Genetics Research and Application Laboratories for their contributions. This research was funded through a grant from the GENAR Institute for Public Health and Genomics Research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Karaca, S., Cesuroglu, T., Karaca, M. et al. Genetic diversity of disease-associated loci in Turkish population. J Hum Genet 60, 193–198 (2015). https://doi.org/10.1038/jhg.2015.8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2015.8
This article is cited by
-
Allergy-specific Phenome-Wide Association Study for Immunogenes in Turkish Children
Scientific Reports (2016)