Abstract
The dystrophin gene, which is mutated in Duchenne muscular dystrophy (DMD), comprises 79 exons that show multiple alternative splicing events. Intron retention, a type of alternative splicing, may control gene expression. We examined intron retention in dystrophin introns by reverse-transcription PCR from skeletal muscle, focusing on the nine shortest (all <1000 bp), because these are more likely to be retained. Only one, intron 40, was retained in mRNA; sequencing revealed insertion of a complete intron 40 (851 nt) between exons 40 and 41. The intron 40 retention product accounted for 1.2% of the total product but had a premature stop codon at the fifth intronic codon. Intron 40 retention was most strongly observed in the kidney (36.6%) and was not obtained from the fetal liver, lung, spleen or placenta. This indicated that intron retention is a tissue-specific event whose level varies among tissues. In two DMD patients, intron 40 retention was observed in one patient but not in the other. Examination of splicing regulatory factors revealed that intron 40 had the highest guanine–cytosine content of all examined introns in a 30-nt segment at its 3′ end. Further studies are needed to clarify the biological role of intron 40-retained dystrophin mRNA.
Similar content being viewed by others
Introduction
The dystrophin gene is one of the largest human genes, spanning 2.4 Mb on the X chromosome. Mutations in the dystrophin gene cause dystrophin deficiency in the skeletal muscles of people with Duchenne muscular dystrophy (DMD), the most common inherited muscle disease, which shows progressive and fatal muscle wasting. The dystrophin gene encodes a 14-kb mRNA consisting of 79 exons; >99% of the gene sequence is intronic. The splicing machinery recognizes the introns properly to produce mature mRNA, even with a wide range of sizes from 107 bp (intron 14) to 248, 401 bp (intron 44). The mechanisms involved in splicing huge introns are controversial.1 Recently, multiple nested splicing was proposed as a mechanism for splicing of the huge dystrophin intron 7 (110, 119 bp).2 In this mechanism, many splice site-like sequences within the large intron are consecutively activated in the splicing of nested introns.
Alternative splicing produces different patterns of mRNA, thus generating several protein isoforms from one gene. There are three types of alternative splicing: (1) exon skipping, whereby an exon is not included in the mature mRNA, (2) alternative use of splice sites, resulting in longer or shorter exons, and (3) intron retention, whereby the full intron sequence is incorporated into the mature transcript.3, 4 Exon skipping is the most common type of alternative splicing and its physiological role has been well studied.5, 6 In the dystrophin gene, at least 20 patterns of alternative exon skipping have been identified.7, 8, 9
Intron retention is a less common type of alternative splicing and is certainly the least studied, especially because these variants are believed to be largely derived from unspliced or partially spliced pre-mRNA. Intron-retained mRNA, which can sometimes insert a premature stop codon in the mature mRNA,4 has been considered to be degraded by nonsense-mediated decay.10 However, recent reports have indicated that intron retention can have a biological role, such as in the control of gene expression.11, 12, 13, 14 For instance, multiple intron retention events were shown to regulate the differentiation of granulocytes.15 The intron-retaining transcripts were suggested to act as RNA-based signals.12, 15
We have previously analyzed the dystrophin mRNA in DMD patients to identify the responsible gene mutation. We identified splicing errors in 5% of the 442 cases.16 At the same time, we have identified the incorporation of intronic regions into the dystrophin mRNA, forming cryptic exons.17, 18, 19 So far, 15 cryptic exons have been identified, scattered among the introns of the dystrophin gene.20 In addition, we have identified circular RNA consisting of authentic dystrophin exons.8 Recently, six novel long non-coding RNAs were shown to be transcribed from the dystrophin gene.21 Taken together, these indicate the diversity of dystrophin transcripts.
The dystrophin gene possesses tissue-specific promoters that produce alternative tissue-specific mRNAs, such as the retina-specific R-dystrophin mRNA.22 In our previous study, we identified an intron retention in the 5′ untranslated region of the R-dystrophin transcript.23 This was compatible with the fact that most intron retention events occur in the 5′ untranslated region.13 It has been proposed that shorter introns have a higher chance of being retained.24 Here, we examined the possibility of retention of the nine shortest dystrophin introns in mRNA from skeletal muscle. For the first time, we identified a complete intron 40 retained in mature dystrophin mRNA.
Materials and methods
Dystrophin mRNA analysis
Human total RNAs from skeletal muscle and 19 other tissues (colon, cerebellum, whole brain, fetal brain, fetal liver, heart, kidney, liver, lung, placenta, prostate, salivary gland, small intestine, spleen, testis, thymus, thyroid gland, trachea and uterus) were obtained from a human total RNA Master Panel II (lot number 1007127A, Clontech Laboratories, Inc., Mountain View, CA, USA). Total RNA was extracted from peripheral blood cells of two DMD patients (KUCG# 434 and 449 with a mutation in the dystrophin gene of c.3347-3350AGAAdel in exon 25 and deletion of exons from 10 to 13, respectively) for clinical diagnosis as described before.16 Blood samples were obtained after obtaining the informed consent.16 Dystrophin mRNA was analyzed as described previously.25 cDNA was synthesized from 0.5 μg of each total RNA.26 To examine intron-retention events in the dystrophin mRNA, nine introns <1000-bp long (considering the efficiency of PCR amplification) were selected from the 78 dystrophin introns. The introns selected were introns 10, 14, 24, 31, 35, 40, 58, 70 and 75, and their size ranged from 107 bp (intron 14) to 991 bp (intron 24). The introns were amplified using primers within the flanking exons (Table 1). To confirm retention of introns 40, 58 and 70, the regions were amplified using a primer from a neighboring exon (Table 1).
PCR amplification was performed in a total volume of 10 μl, containing 1 μl cDNA, 1 μl 10 × ExTaq buffer (Takara Bio, Inc., Shiga, Japan), 0.25 U ExTaq polymerase (Takara Bio, Inc.), 500 nM each primer and 200 μM dNTPs (Takara Bio, Inc.). Thirty cycles of amplification were performed on a Mastercycler Gradient PCR machine (Eppendorf, Hamburg, Germany) using the following conditions: initial denaturation at 94 °C for 3 min, subsequent denaturation at 94 °C for 0.5 min, annealing at 60 °C for 0.5 min, and extension at 72 °C for 1.5 min.
Amplified products were separated and semi-quantitated with a DNA 1000 LabChip Kit on an Agilent 2100 Bioanalyzer (Agilent Technologies, Inc., Santa Clara, CA, USA). The percentages of intron-retaining mRNA were calculated using the following formula: percentage=(amount of intron-retaining product/(amount of intron-retaining product+amount of noramlly spliced product)) × 100.
The amplified products were sequenced using gel cut products by the Greiner Bio-One Co. Ltd., (Tokyo, Japan). To check the integrity and concentration of the cDNA, the mRNA of the GAPDH (glyceraldehyde 3-phosphate dehydrogenase) gene was also amplified by reverse-transcription PCR (RT–PCR) as described previously.27
Indices of splicing regulatory factors
Indices of splicing regulatory factors for each exon/intron were obtained as described before.20 Splice site strength was determined by Shapiro’s splicing probability matrix scores at the 5′ and 3′ splice sites.28 The number of exonic splicing enhancers (ESEs) was calculated using the ‘RESE’ prediction algorithm available at http://genes.mit.edu/burgelab/rescue-ese/. The number of exonic splicing silencers (ESSs) was calculated using the ‘FESS’ algorithm available at http://genes.mit.edu/fas-ess/.29 To calculate the densities of ESSs and ESEs, the RESE and FESS numbers were divided by the sequence length (in nucleotides) and this figure was multiplied by 100 to give the RESE-D and FESS-D scores.30The percentage of guanine–cytosine (GC) content for 30 nucleotides upstream and downstream of the 5′ and 3′ splice sites was also calculated.31
Results
Detection of intron 40 retention in dystrophin mRNA
The nine shortest introns were examined for intron retention. Splicing patterns of the nine introns were analyzed by RT–PCR amplification of each intron-encompassing region using total RNA from skeletal muscle. Among the nine amplifications, six regions, encompassing introns 10, 14, 24, 31, 35 and 75, were amplified as a single product with the expected size (Figure 1a). Three regions, encompassing introns 40, 58 and 70, displayed a minor larger product in addition to the major expected-size product. To confirm the larger product, the three regions were amplified again using a primer within a neighboring exon (Figure 1b). One region, extending from exons 39 to 41, was amplified as two products but the other two regions were not (Figure 1b). Sequencing of the larger product spanning exons 39 to 41 revealed that the entire 851-bp intron 40 was present between exons 40 and 41. We supposed that the intron retention would be caused by nucleotide changes that suppressed intron 40 recognition by the spliceosome, but none were identified within exon 40, intron 40 or exon 41. Therefore, the retention of intron 40 was considered to be because of alternative splicing. In other words, the genomic region from the beginning of exon 40 to the end of exon 41 was recognized as a 1187-bp exon by the spliceosome. Because a stop codon occurred at the fifth codon in intron 40 based on the dystrophin reading frame (Figure 1b,c), it is difficult to predict the role of intron 40 retention based on protein-coding ability. Overall, only one intron of the nine examined was subjected to intron retention.
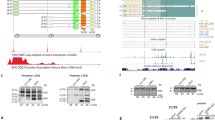
RT–PCR amplification of nine fragments of dystrophin mRNA. Nine fragments encompassing the shortest dystrophin introns were amplified from skeletal muscle total RNA. Electropherograms of the amplified products. (a) A single amplified product was obtained from six introns: a single fragment of the expected size was seen (10–11, 14–15, 24–25, 31–32, 35–36 and 75–76, containing introns 10, 14, 24, 31, 35 and 75, respectively). For the fragments encompassing exons 40–41, 58–59 and 70–71, an additional weak larger band was observed. (b) Re-amplification of these three regions using a primer in the neighboring exon revealed an additional large size product (*) from the region encompassing intron 40 (39–41) but not the other two (58–60 and 70–72). Sequencing of the large product revealed the full 851-bp sequence of intron 40 between exons 40 and 41 (*). The exon structure of the two products is described schematically on the right (* and **). Partial nucleotide sequences at the junctions between exon 40 and intron 40 and between intron 40 and exon 41 are shown under or over the boxes, respectively. The inserted sequence starts with GT and ends with AG and completely matches the full sequence of intron 40. MK refers to the size marker (DNA 1000 Marker). (c) Nucleotide sequence of the 5′ end of intron 40. A TAG stop codon appears at the fifth codon (marked with ‘bold’). A full color version of this figure is available at the Journal of Human Genetics journal online.
Tissue specificity of intron 40 retention
As tissue-specific alternative splicing has been reported in the dystrophin gene, we supposed that this intron 40 retention was a tissue-specific event. We therefore semi-quantitatively RT-RCR amplified the exon 39–41 region in 20 human tissue RNAs (Figure 2). We found that the retention levels of intron 40 varied among tissues. In the skeletal muscle, the percentage of intron 40-retaining product calculated from electropherogram density was 1.2% (Figure 2). The intron 40-retaining product was obtained most abundantly from kidney, where it represented 36.6%. The thymus showed the second strongest intron 40 retention at 9.7%. The salivary gland and testis had >8% intron 40 retention. Intron retention was not observed in the fetal liver, liver, lung, spleen or placenta (Figure 2).
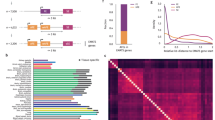
Intron 40 retention in 20 human tissues. The fragment encompassing dystrophin exons 39–41 was RT–PCR amplified from 20 human tissues and the percentage of intron 40 retention calculated. (a) Electropherograms of RT–PCR amplified products. The large fragment corresponding to intron 40 retention is marked. The exon structure of the two products is described schematically on the right (arrows). GAPDH (a 302-bp fragment) was amplified as a control. MK refers to the size marker (DNA 1000 Marker). (b) Percentage of intron 40 retention in each tissue. A full color version of this figure is available at the Journal of Human Genetics journal online.
Intron 40 retention in DMD patients
To examine whether intron 40 retention occurs in DMD patients, dystrophin mRNA of two DMD patients was RT–PCR amplified. In one patient (#449), only one band was visible, corresponding to the normally spliced product (Figure 3). In contrast, two amplified bands were visualized in the other patient (#434): one corresponded to the normally spliced product and the other larger band corresponded to the intron 40-retained product (Figure 3). Remarkably, it accounted for 52.9% of the total amplified product. There is therefore a difference in the patterns of intron 40 retention among DMD patients.
Intron 40 retention in DMD. The region from exon 39 to 41 was RT-PCR amplified from the blood cell RNA of two DMD patients. Electropherograms of RT-PCR amplified products are shown. In one patient (#449), a single, normally spliced product was observed. In the other patient (#434), two PCR products were visible: the normal fragment and the intron 40-retaining fragment. The exon structures of the two products are described schematically on the right. A full color version of this figure is available at the Journal of Human Genetics journal online.
Characterization of intron 40
Despite our supposition that many short dystrophin introns would be retained in the mRNA, only one, intron 40, was found to be retained. This suggests unique splicing regulation of intron 40. Therefore, we compared the splicing regulatory factors of intron 40 with those of the other, non-retained introns (Table 2). Although intron size has been proposed to be an important factor for intron retention,24 intron 40 ranked only the seventh shortest among the nine introns examined. The size of the generated exon (two exons plus retained intron) was the third longest. Furthermore, the sizes of the upstream and downstream introns did not highlight intron 40. Functionally, intron 40 was supposed to have weak splice site-recognition signals. We calculated scores for splice site recognition at both the 5′ and 3′ splice sites (Table 2). The average scores for the 5′ and 3′ splice sites of the nine introns were 0.81 (range: 0.74–0.87) and 0.83 (range: 0.66–0.95), respectively. The scores for intron 40 were 0.81 and 0.85, respectively, meaning that they were not notably weak. GC content and stop codon density have also been reported to contribute to intron retention,13 but once again, intron 40 displayed nothing unusual (GC content, 30.9%; stop codon density, 2.23/100 nt) (Table 2). Furthermore, there were no peculiarities in the densities of ESEs and ESSs in the intron 40 sequence (Table 2).
Recently, intron retention was shown to be related to GC content in intron subregions.15 We examined this by segmenting intron 40 into five parts (the upstream exon sequence, 30 nt at the 5′ end of the intron, 30 nt at the 3′ end of the intron, the remaining intron sequence and the downstream exon sequence) (Table 3). In four of the segments, the value for intron 40 was within one s.d. of the mean. Remarkably, however, the GC content in the 30 nt at the 3′ end of intron 40 was the highest among the nine introns examined. This was the only factor that discriminated intron 40 from the others.
Discussion
The splicing of dystrophin pre-mRNA is considered to occur with incredible fidelity, even though this gene contains huge introns. In this study, we examined intron retention in nine short dystrophin introns and determined that the full 851-nt sequence of intron 40 was inserted between exons 40 and 41 of dystrophin mRNA from the skeletal muscle (Figure 1b). This was enabled by PCR amplification using primers in flanking exons (Figure 1a). In contrast, intron retention was not identified in the other eight short dystrophin introns (Figure 1a). It was remarkable that the short intron 14, at only 107-nt long, was spliced well. With respect to the mechanism of somatic intron retention, the most intuitive explanation was that a somatic mutation had occurred at the splice sites that altered the splicing signal, therefore the splice sites were not properly recognized. However, we could not find any nucleotide change affecting splicing.
Currently, there are two ways to detect intron retention: (1) RT–PCR amplification of the target region32, 33 and (2) transcriptome analysis.14, 15, 34, 35 By whole-transcript sequencing of lung adenoma and matched normal tissues, nearly 3700 genes were found to exhibit intron retention.14 No intron retention event in the dystrophin gene was detected in this previous study, even though dystrophin has been shown to be a tumor-suppressor gene.36 Our results suggest that RT–PCR amplification of the target region is a more powerful way to identify intron retention.
In constitutive splicing, each pre-mRNA is spliced in the same way from a given gene based on the presence of consensus sequences, including the 5′ splice site, the branchpoint and the 3′ splice site, each of which is strictly required by the spliceosome for substrate recognition and catalysis.37 In alternative splicing recognition, the joining of a 5′ and 3′ splice site pair are in competition with at least one other 5′ or 3′ splice site; the competition is controlled by splicing regulatory elements that are often target sites for trans-acting factors.5 The fact that intron retention patterns can be different among various tissues suggests that other factors, such as the cellular environment, may also function in promoting intron retention.13 In our study, we demonstrated tissue-specific differences in intron 40-retention rate (Figure 2), indicating tissue-specific differences in the splicing environment. In this study, commercially available RNAs were used as source materials. RNAs of whole brain, kidney, liver and colon were from single individuals, and other tissues were from pooled populations. Therefore, it is possible that inter-individual variability in alternative splicing patterns influenced the results.
To clarify the relationship between intron retention and splicing regulatory factors, we examined the characteristics of intron 40 splicing regulatory factors. It has been suggested that cis-regulatory elements are likely to have a crucial role in regulating intron retention.24 A higher frequency of intron retention has been observed in introns with weaker splice sites and genes with shorter intron lengths, higher expression levels and lower density of both ESEs and intronic splicing enhancers.13, 24, 32, 38, 39, 40 Our findings showed that the only factor discriminating intron 40 from the non-retained introns was that it had the highest GC content in a 30-nt segment at its 3′ end (Table 3).
It has long been questioned why the dystrophin gene is so huge, with >99% of its sequence being intronic. However, transcript analysis has revealed additional active regions: 15 cryptic exons,20 12 circular RNAs8 and 6 long non-coding RNAs,21 which serve to highlight the complexity of transcripts arising from the dystrophin gene. Our study indicates yet further complexity.
Intron retention has been widely recognized to be a consequence of mis-splicing. It remains doubtful whether a significant fraction of these events have biological significance or whether they are spurious products from the splicing machinery.13 Recently, intron retention has been shown to control gene expression; it was associated with a downregulation of splicing factors and reduced mRNA and protein levels by triggering the nonsense-mediated RNA decay pathway.15 It has been hypothesized that intron retention and consequent nonsense-mediated RNA decay collectively counteract the overexpression of genes promoting cancer development.14 In the RON gene, a universally expressed tyrosine kinase receptor gene, intron retention has a role in controlling receptor function; that is, intron retention suppresses receptor activity.41 Conversely, one out of the three intron retentions in the DKC1 gene are suggested to lead to the exonizing of an intronically encoded small nucleolar RNA.33 Considering these observations, the intron 40-retaining transcript may have a biological role.
DMD is considered stereotyped in its clinical presentation, evolution and severity, with patients succumbing mostly in their twenties because of cardiac or respiratory failure.42, 43 The well-known complication of mental retardation provides some diversity in the DMD phenotype. A recent detailed examination of DMD patients revealed clinical heterogeneity, dividing DMD into four sub-phenotypes that differed in the severity of muscle and brain dysfunction.44 Our preliminary results showed a different degree of intron 40 retention in DMD patients (Figure 3); this alternative splicing event may have a role in the clinical heterogeneity. In our study, however, there was no clear clinical difference between the two patients. To clarify this, it is necessary to study intron retention in greater numbers of dystrophinopathy patients.
We identified intron 40 retention not only in skeletal muscle but also in other tissues, such as kidney, fetal brain, thymus, salivary gland and testis (Figure 2). The level of intron 40 retention differed greatly from tissue to tissue. The highest retention was observed in kidney (Figure 2). If the intron-retained transcript has a biological role, this implies that renal abnormalities would be part of the DMD phenotype, as previously suspected.45
References
Singh, J. & Padgett, R. A. Rates of in situ transcription and splicing in large human genes. Nat. Struct. Mol. Biol. 16, 1128–1133 (2009).
Suzuki, H., Kameyama, T., Ohe, K., Tsukahara, T. & Mayeda, A. Nested introns in an intron: evidence of multi-step splicing in a large intron of the human dystrophin pre-mRNA. FEBS Lett. 587, 555–561 (2013).
Black, D. L. Mechanisms of alternative pre-messenger RNA splicing. Annu. Rev. Biochem. 72, 291–336 (2003).
Matlin, A. J., Clark, F. & Smith, C. W. Understanding alternative splicing: towards a cellular code. Nat. Rev. Mol. Cell Biol. 6, 386–398 (2005).
Fu, X. D. & Ares, M. Jr. Context-dependent control of alternative splicing by RNA-binding proteins. Nat. Rev. Genet. 10, 689–701 (2014).
Keren, H., Lev-Maor, G. & Ast, G. Alternative splicing and evolution: diversification, exon definition and function. Nat. Rev. Genet. 11, 345–355 (2010).
Feener, C. A., Koenig, M. & Kunkel, L. M. Alternative splicing of human dystrophin mRNA generates isoforms at the carboxy terminus. Nature 338, 509–511 (1989).
Surono, A., Takeshima, Y., Wibawa, T., Ikezawa, M., Nonaka, I. & Matsuo, M. Circular dystrophin RNAs consisting of exons that were skipped by alternative splicing. Hum. Mol. Genet. 8, 493–500 (1999).
Sironi, M., Cagliani, R., Pozzoli, U., Bardoni, A., Comi, G. P. & Giorda, R. et al. The dystrophin gene is alternatively spliced throughout its coding sequence. FEBS Lett. 517, 163–166 (2002).
Lareau, L. F., Green, R. E., Bhatnagar, R. S. & Brenner, S. E. The evolving roles of alternative splicing. Curr. Opin. Struct. Biol. 14, 273–282 (2004).
Melhuish, T. A. & Wotton, D. The Tgif2 gene contains a retained intron within the coding sequence. BMC Mol. Biol. 7, 2 (2006).
Khaladkar, M., Buckley, P. T., Lee, M. T., Francis, C., Eghbal, M. M. & Chuong, T. et al. Subcellular RNA sequencing reveals broad presence of cytoplasmic intron-sequence retaining transcripts in mouse and rat neurons. PLoS ONE 8, e76194 (2013).
Galante, P. A., Sakabe, N. J., Kirschbaum-Slager, N. & de Souza, S. J. Detection and evaluation of intron retention events in the human transcriptome. RNA 10, 757–765 (2004).
Zhang, Q., Li, H., Jin, H., Tan, H., Zhang, J. & Sheng, S. The global landscape of intron retentions in lung adenocarcinoma. BMC Med. Genomics 7, 15 (2014).
Wong, J. J., Ritchie, W., Ebner, O. A., Selbach, M., Wong, J. W. & Huang, Y. et al. Orchestrated intron retention regulates normal granulocyte differentiation. Cell 154, 583–595 (2013).
Takeshima, Y., Yagi, M., Okizuka, Y., Awano, H., Zhang, Z. & Yamauchi, Y. et al. Mutation spectrum of the dystrophin gene in 442 Duchenne/Becker muscular dystrophy cases from one Japanese referral center. J. Hum. Genet. 55, 379–388 (2010).
Dwi Pramono, Z. A., Takeshima, Y., Surono, A., Ishida, T. & Matsuo, M. A novel cryptic exon in intron 2 of the human dystrophin gene evolved from an intron by acquiring consensus sequences for splicing at different stages of anthropoid evolution. Biochem. Biophys. Res. Commun. 267, 321–328 (2000).
Ishibashi, K., Takeshima, Y., Yagi, M., Nishiyama, A. & Matsuo, M. Novel cryptic exons identified in introns 2 and 3 of the human dystroiphin gene with duplication of exons 8-11. Kobe J. Med. Sci. 52, 61–75 (2006).
Zhang, Z., Habara, Y., Nishiyama, A., Oyazato, Y., Yagi, M. & Takeshima, Y. et al. Identification of seven novel cryptic exons embedded in the dystrophin gene and characterization of 14 cryptic dystrophin exons. J. Hum. Genet. 52, 607–617 (2007).
Malueka, R. G., Takaoka, Y., Yagi, M., Awano, H., Lee, T. & Dwianingsih, E. K. et al. Categorization of 77 dystrophin exons into 5 groups by a decision tree using indexes of splicing regulatory factors as decision markers. BMC Genet. 13, 23 (2012).
Bovolenta, M., Erriquez, D., Valli, E., Brioschi, S., Scotton, C. & Neri, M. et al. The DMD locus harbours multiple long non-coding RNAs which orchestrate and control transcription of muscle dystrophin mRNA isoforms. PLoS ONE 7, e45328 (2012).
Ahn, A. H. & Kunkel, L. M. The structural and functional diversity of dystrophin. Nat. Genet. 3, 283–291 (1993).
Kubokawa, I., Takeshima, Y., Ota, M., Enomoto, M., Okizuka, Y. & Mori, T. et al. Molecular characterization of the 5′-UTR of retinal dystrophin reveals a cryptic intron that regulates translational activity. Mol. Vis. 16, 2590–2597 (2010).
Sakabe, N. J. & de Souza, S. J. Sequence features responsible for intron retention in human. BMC Genomics 8, 59 (2007).
Matsuo, M., Masumura, T., Nishio, H., Nakajima, T., Kitoh, Y. & Takumi, T. et al. Exon skipping during splicing of dystrophin mRNA precursor due to an intraexon deletion in the dystrophin gene of Duchenne muscular dystrophy. Kobe J. Clin. Invest. 87, 2127–2131 (1991).
Nishida, A., Kataoka, N., Takeshima, Y., Yagi, M., Awano, H. & Ota, M. et al. Chemical treatment enhances skipping of a mutated exon in the dystrophin gene. Nat. Commun. 2, 308 (2011).
Tran, V. K., Zhang, Z., Yagi, M., Nishiyama, A., Habara, Y. & Takeshima, Y. et al. A novel cryptic exon identified in the 3′ region of intron 2 of the human dystrophin gene. J. Hum. Genet. 50, 425–433 (2005).
Shapiro, M. B. & Senapathy, P. RNA splice junctions of different classes of eukaryotes: sequence statistics and functional implications in gene expression. Nucleic Acids Res. 15, 7155–7174 (1987).
Fairbrother, W. G., Yeh, R. F., Sharp, P. A. & Burge, C. B. Predictive identification of exonic splicing enhancers in human genes. Science 297, 1007–1013 (2002).
Královicová, J. & Vorechovsky, I. Global control of aberrant splice-site activation by auxiliary splicing sequences: evidence for a gradient in exon and intron definition. Nucleic Acids Res. 35, 6399–6413 (2007).
Zhang, J., Kuo, C. C. & Chen, L. GC content around splice sites affects splicing through pre-mRNA secondary structures. BMC Genomics 12, 90 (2011).
Smith, L. D., Lucas, C. M. & Eperon, I. C. Intron retention in the alternatively spliced region of RON results from weak 3′ splice site recognition. PLoS ONE 8, e77208 (2013).
Turano, M., Angrisani, A., Di Maio, N. & Furia, M. Intron retention: a human DKC1 gene common splicing event. Biochem. Cell Biol. 91, 506–512 (2013).
Cho, V., Mei, Y., Sanny, A., Chan, S., Enders, A. & Bertram, E. M. et al. The RNA-binding protein hnRNPLL induces a T cell alternative splicing program delineated by differential intron retention in polyadenylated RNA. Genome Biol. 15, R26 (2014).
Pimentel, H., Parra, M., Gee, S., Ghanem, D., An, X. & Li, J. et al. A dynamic alternative splicing program regulates gene expression during terminal erythropoiesis. Nucleic Acids Res. 42, 4031–4042 (2014).
Wang, Y., Marino-Enriquez, A., Bennett, R. R., Zhu, M., Shen, Y. & Eilers, G. et al. Dystrophin is a tumor suppressor in human cancers with myogenic programs. Nat. Genet. 46, 601–606 (2014).
Wahl, M. C., Will, C. L. & Luhrmann, R. The spliceosome: design principles of a dynamic RNP machine. Cell 136, 701–718 (2009).
Talerico, M. & Berget, S. M. Intron definition in splicing of small Drosophila introns. Mol. Cell. Biol. 14, 3434–3445 (1994).
Stamm, S., Zhu, J., Nakai, K., Stoilov, P., Stoss, O. & Zhang, M. Q. An alternative-exon database and its statistical analysis. DNA Cell Biol. 19, 739–756 (2000).
Ohler, U., Shomron, N. & Burge, C. B. Recognition of unknown conserved alternatively spliced exons. PLoS Comput. Biol. 1, 113–122 (2005).
Collesi, C., Santoro, M. M., Gaudino, G. & Comoglio, P. M. A splicing variant of the RON transcript induces constitutive tyrosine kinase activity and an invasive phenotype. Mol. Cell. Biol. 16, 5518–5526 (1996).
Mendell, J. R., Province, M. A., Moxley, R. T. 3rd, Griggs, R. C., Brooke, M. H. & Fenichel, G. M. et al. Clinical investigation of Duchenne muscular dystrophy. A methodology for therapeutic trials based on natural history controls. Arch. Neurol. 44, 808–811 (1987).
Brooke, M. H., Fenichel, G. M., Griggs, R. C., Mendell, J. R., Moxley, R. & Florence, J. et al. Duchenne muscular dystrophy: patterns of clinical progression and effects of supportive therapy. Neurology 39, 475–481 (1989).
Desguerre, I., Christov, C., Mayer, M., Zeller, R., Becane, H. M. & Bastuji-Garin, S. et al. Clinical heterogeneity of duchenne muscular dystrophy (DMD): definition of sub-phenotypes and predictive criteria by long-term follow-up. PLoS ONE 4, e4347 (2009).
Lidov, H. G. & Kunkel, L. M. Dystrophin and Dp140 in the adult rodent kidney. Lab. Invest. 78, 1543–1551 (1998).
Acknowledgements
This work was supported by a Grant-in-Aid for Scientific Research (B) and a Grant-in-Aid for Exploratory Research from the Japan Society for the Promotion of Science, a Health and Labour Sciences Research Grant for Research on Psychiatric and Neurological Diseases and Mental Health and a research grant for Nervous and Mental Disorders from the Ministry of Health, Labour and Welfare, Japan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Nishida, A., Minegishi, M., Takeuchi, A. et al. Tissue- and case-specific retention of intron 40 in mature dystrophin mRNA. J Hum Genet 60, 327–333 (2015). https://doi.org/10.1038/jhg.2015.24
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2015.24
This article is cited by
-
DMD transcripts in CRL-2061 rhabdomyosarcoma cells show high levels of intron retention by intron-specific PCR amplification
Cancer Cell International (2017)
-
Normal and altered pre-mRNA processing in the DMD gene
Human Genetics (2017)
-
Neuronal SH-SY5Y cells use the C-dystrophin promoter coupled with exon 78 skipping and display multiple patterns of alternative splicing including two intronic insertion events
Human Genetics (2015)