Abstract
RASopathies or RAS/mitogen-activated protein kinase (MAPK) syndromes are a group of phenotypically overlapping syndromes caused by germline mutations that encode components of the RAS/MAPK signaling pathway. These disorders include neurofibromatosis type I, Legius syndrome, Noonan syndrome, Noonan syndrome with multiple lentigines (formerly called LEOPARD syndrome), Costello syndrome, cardiofaciocutaneous (CFC) syndrome, Noonan-like syndrome, hereditary gingival fibromatosis and capillary malformation–arteriovenous malformation. Recently, novel gene variants, including RIT1, RRAS, RASA2, A2ML1, SOS2 and LZTR1, have been shown to be associated with RASopathies, further expanding the disease entity. Although further analysis will be needed, these findings will help to better elucidate an understanding of the pathogenesis of these disorders and will aid in the development of potential therapeutic approaches. In this review, we summarize the novel genes that have been reported to be associated with RASopathies and highlight the cardiovascular abnormalities that may arise in affected individuals.
Similar content being viewed by others
Introduction
RAS is a member of small GTPases that regulate cell growth, proliferation and differentiation. RAS GTPases convey an extracellular signal to its target of effector proteins in cells. RAS cycles between the guanosine diphosphate (GDP)-bound inactive form and the guanosine triphosphate (GTP)-bound active form. GTP-bound RAS utilizes several downstream effectors, including RAF1, PI-3 kinase, PLCɛ and Ral-GDS.1
The RAS/mitogen-activated protein kinase (MAPK) pathway is an essential signaling pathway that controls cell proliferation, differentiation and survival. Numerous studies have revealed that dysregulation of the RAS/MAPK pathway causes clinically overlapping genetic disorders, termed ‘RASopathies’ or ‘RAS/MAPK syndromes’.2, 3 Although each RASopathy has a unique phenotype, these syndromes have many overlapping characteristics, including craniofacial dysmorphology, cardiovascular abnormalities, musculoskeletal abnormalities, cutaneous lesions, neurocognitive impairment and increased risk of tumor (for a review of the details of each of these disorders, see Rauen4). These disorders include the following: (1) neurofibromatosis type 1 (NF1) caused by haploinsufficiency of neurofibromin;5, 6, 7 (2) NF1-like syndrome caused by haploinsufficiency of SPRED1;8 (3) Noonan syndrome (NS) caused by mutations in PTPN11, SOS1, RAF1, KRAS, BRAF and NRAS;9, 10, 11, 12, 13, 14, 15 (4) NS with multiple lentigines (NSML) caused by mutations in PTPN11 and RAF1;9, 16 (5) Costello syndrome caused by activating mutations in HRAS;17 (6) cardiofaciocutaneous (CFC) syndrome caused by mutations in BRAF, MAP2K1/2 and KRAS;18, 19 (7) Noonan-like syndrome caused by mutations in SHOC220 or CBL;21, 22, 23 (8) hereditary gingival fibromatosis caused by a mutation in SOS1;24 and (9) capillary malformation–arteriovenous malformation caused by haploinsufficiency of RASA1 (also known as p120 Ras-GTPase activating protein (GAP)).25 Molecular analysis is beneficial for both the confirmation of clinical diagnoses and to perform follow-up according to the unique characteristics of each disorder. In this review, we summarize novel genes that have been reported to be associated with RASopathies, including RIT1, RRAS, RASA2, A2ML1, SOS2 and LZTR1, and discuss the cardiovascular abnormalities that have been associated with these syndromes.
Novel genes associated with rasopathies
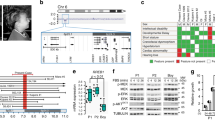
NS (MIM 163950) is an autosomal dominant disorder that is characterized by short stature, facial dysmorphism and congenital heart defects.26 The incidence of this syndrome is estimated to be between 1 in 1000 and 1 in 2500 live births.27 The distinctive craniofacial features that are observed in individuals with NS include a webbed or short neck, hypertelorism, downslanting palpebral fissures, ptosis and low-set, posteriorly rotated ears (see reviews26, 28). More than 80% of individuals with NS have cardiovascular involvement, most frequently including congenital heart diseases, pulmonary valve stenosis and hypertrophic cardiomyopathy.26 Hypertrophic cardiomyopathy is observed in ~20% of individuals.26, 28 Other clinical manifestations include cryptorchidism, bleeding disorders, mild neurocognitive delay and pectus deformity. NS is known to be associated with myeloproliferative disorders. The myeloproliferative disorders most often resolve spontaneously, although select individuals develop juvenile myelomonocytic leukemia, a myeloproliferative disorder characterized by excessive production of myelomonocytic cells.26, 28 As of 2013, seven genes have been shown to be associated with NS: PTPN11 (~50%), SOS1 (11%), RAF1 (5%), KRAS (~1.5%), NRAS (0.2%), SHOC2 (~2%) and CBL (Figure 1).26 However, it is estimated that 20–30% of the causative genes behind NS and NS-like disorders are unidentified. Recent advances in genetic analysis technologies, including whole-exome sequencing, have identified potential causes for RASopathies.
RIT1
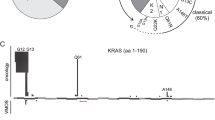
Our group performed whole-exome sequencing of 14 individuals with NS and related conditions who had no detectable mutations in known Noonan-related genes. We found four variants in RIT1 that were clustered within 14 amino acids. Combining these data with additional Sanger sequencing data revealed a total of nine missense, nonsynonymous RIT1 mutations in 17 of a group of 180 individuals (9%) (Table 1).29 The RIT1 protein shares ~50% sequence identity with RAS; comparatively, it has an additional N-terminal extension and does not possess a carboxyl-terminal CAAX motif.30, 31 Past studies have shown that a RIT1 p.Q79L mutant that corresponds to RAS p.Q61L is implicated in transforming NIH3T3 cells, modulating neurite outgrowth in neuronal cells, and activating extracellular-signal-regulated kinase (ERK) and p38 MAPK in a cell-specific manner.32, 33, 34 The mutations in RIT1 that have been identified in individuals with NS are located in its G1 domain (p.S35T) and in the switch I region that is included in its G2 domain (p.A57G). The majority of the mutations (p.E81G, p.F82V, p.F82L, p.T83P, p.Y89H, p.M90I and p.G95A) are clustered within the switch II region that corresponds to RAS. Seventy-percent of mutation-positive individuals had hypertrophic cardiomyopathy, representing a high frequency of individuals with NS. The introduction of mutant RIT1 mRNAs into one-cell stage zebrafish embryos was demonstrated to result in a significant increase of embryos with craniofacial abnormalities, incomplete looping and hypoplastic chambers in the heart, and elongated yolk sacs.29
Following the initial report, Chen et al.35 performed whole-exome sequencing of 27 individuals with NS who did not possess mutations in the genes known to be associated with NS. They identified missense mutations in RIT1 (p.A57G, p.A77P, p.F82V and p.G95A) in five individuals with NS. Bertola et al.36 and Gos et al.37 identified RIT1 mutations in 6 out of 70 individuals and 4 out of 106 individuals, respectively. In total, 10 different RIT1 mutations have been reported in 32 individuals. The most frequent mutation in RIT1 is p.G95A (10 out of 32 individuals). Out of 32 RIT1 mutation-positive individuals, 16 (50%) showed cardiac hypertrophy. Both these results and unpublished data produced by our group suggest that the frequency of RIT1 mutations can be estimated as ~5% in patients with NS, similar to the frequency of RAF1 mutations in these patients. Although somatic RIT1 mutations have previously been considered to be rare in cancer patients, recent reports have identified somatic RIT1 mutations in ~2% of lung adenocarcinomas38, 39 and myeloproliferative or mixed myelodysplastic/myeloproliferative neoplasms, particularly in chronic myelomonocytic leukemia.40
RRAS
Flex et al.41 identified two germline mutations (p.G39dup and p.V55M) in RRAS, a member of the RAS subfamily,42 in two individuals with NS. Germline mutations in RRAS are rare (2 subjects out of 504 individuals with NS and related disorders). They also identified somatic RRAS mutations in 2 out of 110 samples taken from patients with juvenile myelomonocytic leukemia. The expression of the identified RRAS mutations in Caenorhabditis elegans resulted in enhanced RAS signaling and phenotypic abnormalities, similar to what is observed in C. elegans that are expressing a NS causing SHOC2 mutant.20
RASA2
Chen et al.35 identified RASA2 variants in three individuals with NS. RASA2 is a member of the mammalian RAS-GAP family. Loss-of-function mutations in NF1 and RASA1, which are also RAS-GAPs, have been identified in individuals with NF1 and capillary malformation–arteriovenous malformation, respectively.6, 7, 25 All of the identified variations in RASA2 (p.Y326C, p.Y326N and p.R511C) affect highly conserved amino acids in the GAP domain of RASA2. The expression of these mutants in HEK293T cells did not suppress ERK after EGF treatment, unlike in cells with wild-type RASA2. It was concluded that two variants were loss-of-function mutations and one variant was a dominant negative mutation. In contrast with RASA1 (p120GAP), the functional role of RASA2 has not yet been fully elucidated. According to the COSMIC database (http://cancer.sanger.ac.uk/cosmic), somatic missense and nonsense mutations in RASA2 have been identified in various tumors, including those corresponding to colorectal, skin, lung and endometrial cancers.
A2ML1
Vissers et al.43 performed trio exome sequencing and identified a de novo variant (p.R802H) of A2ML1 in an individual with NS. Additional analyses of 155 individuals revealed missense variants (p.R592L and p.R802L) of A2ML1 in two families with NS. Introducing the identified mutations into zebrafish led to developmental defects, including a phenotype that exhibited a broad head, blunted face and cardiac malformations. The A2ML1 gene encodes the secreted protease inhibitor α-2-macroglobulin-like-1, a member of the α-macroglobulin superfamily of proteins. This family contains components of the complement system and protease inhibitors.44 The A2ML1 protein is expressed in epidermal granular keratinocytes and is secreted into extracellular space, where it demonstrates inhibitory activities toward proteases in vitro, including chymotrypsin and papain.44 Such activities suggest that it has a role in the defense mechanisms and maintenance of epidermal homeostasis. It is notable that A2ML1 autoantibodies have frequently been detected in individuals with paraneoplastic pemphigus, an autoimmune multiorgan syndrome that includes intractable stomatitis, polymorphous cutaneous lesions and lymphoproliferative tumors.45, 46 A2ML1 has been shown to bind to LPR1 (low-density lipoprotein receptor-related protein 1).47 LPR1 has been shown to interact with CBL, a causative gene of RASopathy, and it is known to control the ubiquitination of platelet-derived growth factor receptor-β.48 Both the functional properties of A2ML1 and the mechanisms by which A2ML1 regulates the RAS/ERK pathway are largely unknown. Further functional analysis will clarify the role of A2ML1 in developmental disorders.
SOS2
Yamamoto et al.49 performed whole-exome sequencing of 50 Brazilian probands who were negative for the gene mutations known to be associated with NS. They identified two missense variants in SOS2 in three families with NS. De novo occurrence was confirmed in one of three families. SOS2 is homologous to SOS1, the second most frequently mutated gene in individuals with NS. The identified variants, p.M267K and T376S, were located in the DH domain of SOS2, and this is where the SOS1 mutations that were identified in NS patients were also clustered, suggesting that these mutations could be pathogenic.
LZTR1
Yamamoto et al.49 have also identified rare variants of LZTR1, leucine-zipper-like transcription regulator 1, in individuals with NS. They concluded that five variants are predicted to cause NS; three of the variants, p.R284C, p.H287Y and p.Y119C, were confirmed to be de novo events and two of the variants, p.G248R and p.S247N, were found to be segregated in the affected individuals of two families. LZTR1 encodes a protein member of the BTB-kelch superfamily that has not been previously associated with the RAS/MAPK pathway. Somatic and germline mutations in LZTR1 have been identified in patients with glioblastoma multiforme50 and multiple schwannomas51 respectively. LZTR1 is located within the 3-Mb-long region that is most commonly deleted in patients with 22q11 deletion syndrome.52 In two individuals, Chen et al.35 identified LZTR1 p.R237Q and p.A249P variants that have not been considered to be responsible for NS phenotype. Further mutational and functional analyses will elucidate the phenotypes of individuals with LZTR1 variants and the functional consequences of these variants.
Others
Chen et al.35 identified a nonsense variant of SPRY1, a negative regulator of the RAS/ERK pathway as well as a missense variant of MAP3K8 that encodes MAP kinase kinase kinase.35 Further studies will be needed to clarify the pathogenetic significance of these variants.
Cardiovascular abnormalities in rasopathies
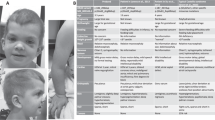
Individuals with RASopathies often have cardiovascular abnormalities (Table 2). The frequency and type of cardiac involvement is different among the different disorders. Individuals with NS, Costello syndrome, CFC syndrome or NSML frequently develop cardiac abnormalities such as hypertrophic cardiomyopathy, pulmonic valve stenosis, septal defects and arrhythmia. More than 80% of individuals with NS have cardiovascular abnormalities.26 Pulmonic valve stenosis is the most common cardiovascular abnormality in patients with NS.28 Pulmonic valve stenosis is common (~70%) in individuals with SOS1 and PTPN11 mutations and is less frequent (~20%) in individuals with RAF1 mutations.53 Individuals with RAF1 mutations and possibly also individuals with RIT1 mutations frequently develop hypertrophic cardiomyopathy (~85 and ~50%, respectively).29, 35, 36, 37, 53 In contrast, hypertrophic cardiomyopathy is less frequent in individuals with SOS1 or PTPN11 mutations.53 NSML was previously referred to as LEOPARD syndrome (an acronym for its cardinal features of multiple lentigines, electrocardiographic conduction abnormalities, ocular hypertelorism, pulmonary stenosis, abnormal genitalia, retardation of growth and sensorineural deafness).16 In contrast with the gain-of-function nature of the PTPN11 mutations that have been identified in individuals with NS, the PTPN11 mutations that have been identified in individuals with NSML have been shown to be catalytically inactive or dominant negative.54, 55, 56 Mutations in RAF1 and BRAF have less frequently been identified in individuals with NSML.9, 15 More than 80% of individuals with NSML present with heart defects; of these, hypertrophic cardiomyopathy occurs in 80%, electrocardiographic abnormalities in 73%, valvular defects in 50%, coronary abnormalities in 15% and septal defects in 1–5%.57, 58
Individuals with NS-like disorder with loose anagen hair who have a common p.S2G mutation in SHOC2 are characterized by a short stature that is associated with growth hormone deficiency, a Noonan-like facial appearance, mild neurodevelopmental delays and easily pluckable hair.20 In two of the initial studies on this disorder, cardiac defects have been observed in 27 out of 33 (~80%) individuals.20, 59 Compared with individuals with NS, septal defects (~42%) and mitral valve anomalies (~31%) were more frequent.20, 59 In following case reports on individuals with the SHOC2 p.S2G mutation, phenotypic variability was noted.60
Costello syndrome is a rare RASopathy that is characterized by distinctive facial features, including full lips, a large mouth and a full nasal tip; soft skin with deep palmer and planter creases, failure to thrive, mild to severe intellectual disability and increased risk of malignant tumors are also characteristics of these patients. Germline activating mutations in HRAS (G12S in ~80% of Costello syndrome patients) have been identified in individuals with Costello syndrome.17 The majority of individuals with Costello syndrome have cardiac abnormalities; ~60% have hypertrophic cardiomyopathy, whereas ~44% have congenital heart defects that usually include nonprogressive pulmonary stenosis and ~48% present with atrial tachycardia.61
CFC syndrome shares many overlapping features with NS and Costello syndrome. Individuals with CFC syndrome have characteristic facial features, including high cranial vault, bitemporal constriction, hypoplastic supraorbital ridges, downslanting palpebral fissures, a depressed nasal bridge and posteriorly angulated ears with prominent helices.62, 63 Other clinical features include failure to thrive, hypotonia, motor delay, moderate intellectual disability and ectodermal abnormalities, such as sparse, friable hair, hyperkeratotic skin lesions and a generalized ichthyosis-like condition.62, 63 Germline mutations in BRAF, MAP2K1/2 and KRAS have been identified in individuals with CFC syndrome.18, 19 In our previous cohort, BRAF, MAP2K1/2 and KRAS mutations were identified in 68%, 23% and 9% of individuals, respectively.64 KRAS mutations have also been identified in individuals with NS..13 In CFC syndrome, ~75% of individuals have cardiovascular involvement, including pulmonic valve stenosis, hypertrophic cardiomyopathy and atrial septal defect in ~40%, ~30% and ~20% of individuals, respectively.62
NF1 is an autosomal dominant multisystem disorder affecting ~1 in 3000 newborn.4 Clinical manifestations of NF1 include multiple café-au-lait spots, axillary and inguinal freckling, multiple cutaneous neurofibromas, iris Lisch nodules and a distinctive osseous lesion such as sphenoid dysplasia or tibial pseudarthrosis. Legius syndrome is a NF1-like disorder, characterized by multiple café-au-lait macules, intertriginous freckling, lipomas, macrocephaly and learning disabilities without neurofibromas or other tumor manifestations.8 Loss-of-function mutations in SPRED1 have been identified in individuals with Legius syndrome.8 Lin et al.65 have reported that 54 out of 2322 (2.3%) individuals with NF1 had cardiovascular malformations. Of 54 individuals with cardiovascular abnormalities, flow defects resulting from abnormal embryonic intracardiac hemodynamics were observed in 43 (80%), pulmonic stenosis in 25 (58%) and aortic coarctation in 5 (9%).65 Individuals with NF1 have been shown to have a wide range of vascular abnormalities.66 Stenosis, aneurysms and occlusions of the major arteries and of arteries in the heart, brain and kidney were observed.5, 66 Hypertension is a relatively frequent manifestation.67 Cardiac involvement is less frequent in individuals with Legius syndrome68 and NS-like syndrome with CBL mutations.21, 22, 23
Conclusions and future perspective
The identification of the causative genes that underlie the RASopathies has facilitated molecular diagnosis of these disorders, enabled the evaluation of genotype–phenotype relationships and aided in the development of possible therapeutic approaches. Recent technical advances have led to the identification of novel genes that might be associated with RASopathies. Among these, a total of 32 individuals with RIT1 mutations have been reported.29, 35, 36, 37 The clinical manifestations of RIT1 mutation-positive individuals corresponded to those of NS. Rare variants of RRAS, RASA2 and SOS2 are probably associated with RASopathies because these molecules are functionally related to the RAS/ERK pathway. Further analyses of additional cohorts and of the functional roles of A2ML1 and LZTR1 will be required to conclude that these rare variants are associated with RASopathy pathogenesis. The identification of RASopathy-related genes will also provide new insights into the biology of the RAS/MAPK signaling pathway.
A variety of cardiovascular abnormalities have been associated with individuals who are affected by RASopathies. The appropriate treatment of these cardiovascular abnormalities leads to better prognoses for patients with these disorders. Inhibitors of the RAS/MAPK signaling cascade may offer a means of therapeutically treating disorders that involve dysregulation of the RAS/MAPK pathway.69 The 3-hydroxy-3-methylglutaryl-coenzyme A reductase inhibitors have been used in clinical trials to enhance cognitive function in individuals with NF1.70 An open-label study to assess the safety, tolerability, pharmacokinetics and pharmacodynamics of a MEK inhibitor, MEK162 (Novartis), in adults with NS who also have hypertrophic cardiomyopathy is now recruiting (ClinicalTrials.gov identifier: NCT01556568). Indeed, MEK inhibitors have been shown to ameliorate the phenotype of knock-in mouse models for NS (mutations in Sos1 and Raf1)71, 72 and CFC syndrome (Braf mutation),73 suggesting that the phenotypes that are produced by RASopathies can be ameliorated by manipulating RAS/MAPK activity. An inhibitor of mammalian target of rapamycin has been shown to reverse heart defects in both a mouse model of74 and in an individual with NSML.75 Histone demethylase inhibitors that might not be directly associated with the RAS/ERK pathway have also been shown to ameliorate the phenotype of a mouse model of CFC syndrome.73 Further studies will explore the pathogenetic mechanisms behind and therapeutic approaches for RASopathies.
References
Malumbres, M. & Barbacid, M. RAS oncogenes: the first 30 years. Nat. Rev. Cancer. 3, 459–465 (2003).
Aoki, Y., Niihori, T., Narumi, Y., Kure, S. & Matsubara, Y. The RAS/MAPK syndromes: novel roles of the RAS pathway in human genetic disorders. Hum. Mutat. 29, 992–1006 (2008).
Tidyman, W. E. & Rauen, K. A. The RASopathies: developmental syndromes of Ras/MAPK pathway dysregulation. Curr. Opin. Genet. Dev. 19, 230–236 (2009).
Rauen, K. A. The RASopathies. Annu. Rev. Genomics Hum. Genet. 14, 355–369 (2013).
Friedman, J. M. Neurofibromatosis 1. GeneReviews® [Internet] (University of Washington, Seattle, WA, 2014).
Viskochil, D., Buchberg, A. M., Xu, G., Cawthon, R. M., Stevens, J., Wolff, R. K. et al. Deletions and a translocation interrupt a cloned gene at the neurofibromatosis type 1 locus. Cell 62, 187–192 (1990).
Wallace, M. R., Marchuk, D. A., Andersen, L. B., Letcher, R., Odeh, H. M., Saulino, A. M. et al. Type 1 neurofibromatosis gene: identification of a large transcript disrupted in three NF1 patients. Science 249, 181–186 (1990).
Brems, H., Chmara, M., Sahbatou, M., Denayer, E., Taniguchi, K., Kato, R. et al. Germline loss-of-function mutations in SPRED1 cause a neurofibromatosis 1-like phenotype. Nat. Genet. 39, 1120–1126 (2007).
Pandit, B., Sarkozy, A., Pennacchio, L. A., Carta, C., Oishi, K., Martinelli, S. et al. Gain-of-function RAF1 mutations cause Noonan and LEOPARD syndromes with hypertrophic cardiomyopathy. Nat. Genet. 39, 1007–1012 (2007).
Razzaque, M. A., Nishizawa, T., Komoike, Y., Yagi, H., Furutani, M., Amo, R. et al. Germline gain-of-function mutations in RAF1 cause Noonan syndrome. Nat. Genet. 39, 1013–1017 (2007).
Tartaglia, M., Mehler, E. L., Goldberg, R., Zampino, G., Brunner, H. G., Kremer, H. et al. Mutations in PTPN11, encoding the protein tyrosine phosphatase SHP-2, cause Noonan syndrome. Nat. Genet. 29, 465–468 (2001).
Tartaglia, M., Pennacchio, L. A., Zhao, C., Yadav, K. K., Fodale, V., Sarkozy, A. et al. Gain-of-function SOS1 mutations cause a distinctive form of Noonan syndrome. Nat. Genet. 39, 75–79 (2007).
Schubbert, S., Zenker, M., Rowe, S. L., Boll, S., Klein, C., Bollag, G. et al. Germline KRAS mutations cause Noonan syndrome. Nat. Genet. 38, 331–336 (2006).
Cirstea, I. C., Kutsche, K., Dvorsky, R., Gremer, L., Carta, C., Horn, D. et al. A restricted spectrum of NRAS mutations causes Noonan syndrome. Nat. Genet. 42, 27–29 (2010).
Sarkozy, A., Carta, C., Moretti, S., Zampino, G., Digilio, M. C., Pantaleoni, F. et al. Germline BRAF mutations in Noonan, LEOPARD, and cardiofaciocutaneous syndromes: molecular diversity and associated phenotypic spectrum. Hum. Mutat. 30, 695–702 (2009).
Digilio, M. C., Conti, E., Sarkozy, A., Mingarelli, R., Dottorini, T., Marino, B. et al. Grouping of multiple-lentigines/LEOPARD and Noonan syndromes on the PTPN11 gene. Am. J. Hum. Genet. 71, 389–394 (2002).
Aoki, Y., Niihori, T., Kawame, H., Kurosawa, K., Ohashi, H., Tanaka, Y. et al. Germline mutations in HRAS proto-oncogene cause Costello syndrome. Nat. Genet. 37, 1038–1040 (2005).
Niihori, T., Aoki, Y., Narumi, Y., Neri, G., Cave, H., Verloes, A. et al. Germline KRAS and BRAF mutations in cardio-facio-cutaneous syndrome. Nat. Genet. 38, 294–296 (2006).
Rodriguez-Viciana, P., Tetsu, O., Tidyman, W. E., Estep, A. L., Conger, B. A., Cruz, M. S. et al. Germline mutations in genes within the MAPK pathway cause cardio-facio-cutaneous syndrome. Science 311, 1287–1290 (2006).
Cordeddu, V., Di Schiavi, E., Pennacchio, L. A., Ma'ayan, A., Sarkozy, A., Fodale, V. et al. Mutation of SHOC2 promotes aberrant protein N-myristoylation and causes Noonan-like syndrome with loose anagen hair. Nat. Genet. 41, 1022–1026 (2009).
Martinelli, S., De Luca, A., Stellacci, E., Rossi, C., Checquolo, S., Lepri, F. et al. Heterozygous germline mutations in the CBL tumor-suppressor gene cause a Noonan syndrome-like phenotype. Am. J. Hum. Genet. 87, 250–257 (2010).
Niemeyer, C. M., Kang, M. W., Shin, D. H., Furlan, I., Erlacher, M., Bunin, N. J. et al. Germline CBL mutations cause developmental abnormalities and predispose to juvenile myelomonocytic leukemia. Nat. Genet. 42, 794–800 (2010).
Perez, B., Mechinaud, F., Galambrun, C., Ben Romdhane, N., Isidor, B., Philip, N. et al. Germline mutations of the CBL gene define a new genetic syndrome with predisposition to juvenile myelomonocytic leukaemia. J. Med. Genet. 47, 686–691 (2010).
Hart, T. C., Zhang, Y., Gorry, M. C., Hart, P. S., Cooper, M., Marazita, M. L. et al. A mutation in the SOS1 gene causes hereditary gingival fibromatosis type 1. Am. J. Hum. Genet. 70, 943–954 (2002).
Eerola, I., Boon, L. M., Mulliken, J. B., Burrows, P. E., Dompmartin, A., Watanabe, S. et al. Capillary malformation-arteriovenous malformation, a new clinical and genetic disorder caused by RASA1 mutations. Am. J. Hum. Genet. 73, 1240–1249 (2003).
Romano, A. A., Allanson, J. E., Dahlgren, J., Gelb, B. D., Hall, B., Pierpont, M. E. et al. Noonan syndrome: clinical features, diagnosis, and management guidelines. Pediatrics 126, 746–759 (2010).
Allanson, J. E., Hall, J. G., Hughes, H. E., Preus, M. & Witt, R. D. Noonan syndrome: the changing phenotype. Am. J. Med. Genet. 21, 507–514 (1985).
van der Burgt, I. Noonan syndrome. Orphanet J. Rare Dis. 2, 4 (2007).
Aoki, Y., Niihori, T., Banjo, T., Okamoto, N., Mizuno, S., Kurosawa, K. et al. Gain-of-function mutations in RIT1 cause Noonan syndrome, a RAS/MAPK pathway syndrome. Am. J. Hum. Genet. 93, 173–180 (2013).
Wes, P. D., Yu, M. & Montell, C. RIC, a calmodulin-binding Ras-like GTPase. EMBO J. 15, 5839–5848 (1996).
Lee, C. H., Della, N. G., Chew, C. E. & Zack, D. J. Rin, a neuron-specific and calmodulin-binding small G-protein, and Rit define a novel subfamily of ras proteins. J. Neurosci. 16, 6784–6794 (1996).
Rusyn, E. V., Reynolds, E. R., Shao, H., Grana, T. M., Chan, T. O., Andres, D. A. et al. Rit, a non-lipid-modified Ras-related protein, transforms NIH3T3 cells without activating the ERK, JNK, p38 MAPK or PI3K/Akt pathways. Oncogene. 19, 4685–4694 (2000).
Sakabe, K., Teramoto, H., Zohar, M., Behbahani, B., Miyazaki, H., Chikumi, H. et al. Potent transforming activity of the small GTP-binding protein Rit in NIH 3T3 cells: evidence for a role of a p38gamma-dependent signaling pathway. FEBS Lett. 511, 15–20 (2002).
Spencer, M. L. Induction of neurite extension and survival in pheochromocytoma cells by the Rit GTPase. J. Biol. Chem. 277, 20160–20168 (2002).
Chen, P. C., Yin, J., Yu, H. W., Yuan, T., Fernandez, M., Yung, C. K. et al. Next-generation sequencing identifies rare variants associated with Noonan syndrome. Proc. Natl. Acad. Sci. USA 111, 11473–11478 (2014).
Bertola, D. R., Yamamoto, G. L., Almeida, T. F., Buscarilli, M., Jorge, A. A., Malaquias, A. C. et al. Further evidence of the importance of RIT1 in Noonan syndrome. Am. J. Med. Genet. A 164A, 2952–2957 (2014).
Gos, M., Fahiminiya, S., Poznanski, J., Klapecki, J., Obersztyn, E., Piotrowicz, M. et al. Contribution of RIT1 mutations to the pathogenesis of Noonan syndrome: four new cases and further evidence of heterogeneity. Am. J. Med. Genet. A 164A, 2310–2316 (2014).
Berger, A. H., Imielinski, M., Duke, F., Wala, J., Kaplan, N., Shi, G. X. et al. Oncogenic RIT1 mutations in lung adenocarcinoma. Oncogene. 33, 4418–4423 (2014).
Cancer Genome Atlas Research Network Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550 (2014).
Gomez-Segui, I., Makishima, H., Jerez, A., Yoshida, K., Przychodzen, B., Miyano, S. et al. Novel recurrent mutations in the RAS-like GTP-binding gene RIT1 in myeloid malignancies. Leukemia 27, 1943–1946 (2013).
Flex, E., Jaiswal, M., Pantaleoni, F., Martinelli, S., Strullu, M., Fansa, E. K. et al. Activating mutations in RRAS underlie a phenotype within the RASopathy spectrum and contribute to leukaemogenesis. Hum. Mol. Genet. 23, 4315–4327 (2014).
Reuther, G. W. & Der, C. J. The Ras branch of small GTPases: Ras family members don't fall far from the tree. Curr. Opin. Cell. Biol. 12, 157–165 (2000).
Vissers, L. E., Bonetti, M., Paardekooper Overman, J., Nillesen, W. M., Frints, S. G., de Ligt, J. et al. Heterozygous germline mutations in A2ML1 are associated with a disorder clinically related to Noonan syndrome. Eur. J. Hum. Genet. 23, 317–324 (2015).
Galliano, M. F., Toulza, E., Gallinaro, H., Jonca, N., Ishida-Yamamoto, A., Serre, G. et al. A novel protease inhibitor of the alpha2-macroglobulin family expressed in the human epidermis. J. Biol. Chem. 281, 5780–5789 (2006).
Numata, S., Teye, K., Tsuruta, D., Sogame, R., Ishii, N., Koga, H. et al. Anti-alpha-2-macroglobulin-like-1 autoantibodies are detected frequently and may be pathogenic in paraneoplastic pemphigus. J. Invest. Dermatol. 133, 1785–1793 (2013).
Schepens, I., Jaunin, F., Begre, N., Laderach, U., Marcus, K., Hashimoto, T. et al. The protease inhibitor alpha-2-macroglobulin-like-1 is the p170 antigen recognized by paraneoplastic pemphigus autoantibodies in human. PLoS ONE 5, e12250 (2010).
Galliano, M. F., Toulza, E., Jonca, N., Gonias, S. L., Serre, G. & Guerrin, M. Binding of alpha2ML1 to the low density lipoprotein receptor-related protein 1 (LRP1) reveals a new role for LRP1 in the human epidermis. PLoS ONE 3, e2729 (2008).
Takayama, Y., May, P., Anderson, R. G. & Herz, J. Low density lipoprotein receptor-related protein 1 (LRP1) controls endocytosis and c-CBL-mediated ubiquitination of the platelet-derived growth factor receptor beta (PDGFR beta). J. Biol. Chem. 280, 18504–18510 (2005).
Yamamoto, G. L., Aguena, M., Gos, M., Hung, C., Pilch, J., Fahiminiya, S. et al. Rare variants in SOS2 and LZTR1 are associated with Noonan syndrome. J. Med. Genet. 52, 413–421 (2015).
Frattini, V., Trifonov, V., Chan, J. M., Castano, A., Lia, M., Abate, F. et al. The integrated landscape of driver genomic alterations in glioblastoma. Nat. Genet. 45, 1141–1149 (2013).
Piotrowski, A., Xie, J., Liu, Y. F., Poplawski, A. B., Gomes, A. R., Madanecki, P. et al. Germline loss-of-function mutations in LZTR1 predispose to an inherited disorder of multiple schwannomas. Nat. Genet. 46, 182–187 (2014).
Kurahashi, H., Akagi, K., Inazawa, J., Ohta, T., Niikawa, N., Kayatani, F. et al. Isolation and characterization of a novel gene deleted in DiGeorge syndrome. Hum. Mol. Genet. 4, 541–549 (1995).
Kobayashi, T., Aoki, Y., Niihori, T., Cavé, H., Verloes, A., Okamoto, N. et al. Molecular and clinical analysis of RAF1in Noonan syndrome and related disorders: dephosphorylation of serine 259 as the essential mechanism for mutant activation. Hum. Mutat. 31, 284–294 (2010).
Hanna, N., Montagner, A., Lee, W. H., Miteva, M., Vidal, M., Vidaud, M. et al. Reduced phosphatase activity of SHP-2 in LEOPARD syndrome: consequences for PI3K binding on Gab1. FEBS Lett. 580, 2477–2482 (2006).
Kontaridis, M. I., Swanson, K. D., David, F. S., Barford, D. & Neel, B. G. PTPN11 (Shp2) mutations in LEOPARD syndrome have dominant negative, not activating, effects. J. Biol. Chem. 281, 6785–6792 (2006).
Tartaglia, M., Martinelli, S., Stella, L., Bocchinfuso, G., Flex, E., Cordeddu, V. et al. Diversity and functional consequences of germline and somatic PTPN11 mutations in human disease. Am. J. Hum. Genet. 78, 279–290 (2006).
Martinez-Quintana, E. & Rodriguez-Gonzalez, F. LEOPARD syndrome: clinical features and gene mutations. Mol. Syndromol. 3, 145–157 (2012).
Lauriol, J. & Kontaridis, M. I. PTPN11-associated mutations in the heart: has LEOPARD changed Its RASpots? Trends Cardiovasc. Med. 21, 97–104 (2011).
Komatsuzaki, S., Aoki, Y., Niihori, T., Okamoto, N., Hennekam, R. C., Hopman, S. et al. Mutation analysis of the SHOC2 gene in Noonan-like syndrome and in hematologic malignancies. J. Hum. Genet. 55, 801–809 (2010).
Baldassarre, G., Mussa, A., Banaudi, E., Rossi, C., Tartaglia, M., Silengo, M. et al. Phenotypic variability associated with the invariant SHOC2 c.4A>G (p.Ser2Gly) missense mutation. Am. J. Med. Genet. A 164A, 3120–3125 (2014).
Lin, A. E., Alexander, M. E., Colan, S. D., Kerr, B., Rauen, K. A., Noonan, J. et al. Clinical, pathological, and molecular analyses of cardiovascular abnormalities in Costello syndrome: a Ras/MAPK pathway syndrome. Am. J. Med. Genet. A 155A, 486–507 (2011).
Allanson, J. E., Anneren, G., Aoki, Y., Armour, C. M., Bondeson, M. L., Cave, H. et al. Cardio-facio-cutaneous syndrome: does genotype predict phenotype? Am. J. Med. Genet. C Semin. Med. Genet. 157, 129–135 (2011).
Pierpont, M. E., Magoulas, P. L., Adi, S., Kavamura, M. I., Neri, G., Noonan, J. et al. Cardio-facio-cutaneous syndrome: clinical features, diagnosis, and management guidelines. Pediatrics 134, e1149–e1162 (2014).
Narumi, Y., Aoki, Y., Niihori, T., Neri, G., Cave, H., Verloes, A. et al. Molecular and clinical characterization of cardio-facio-cutaneous (CFC) syndrome: overlapping clinical manifestations with Costello syndrome. Am. J. Med. Genet. A 143A, 799–807 (2007).
Lin, A. E., Birch, P. H., Korf, B. R., Tenconi, R., Niimura, M., Poyhonen, M. et al. Cardiovascular malformations and other cardiovascular abnormalities in neurofibromatosis 1. Am. J. Med. Genet. 95, 108–117 (2000).
Oderich, G. S., Sullivan, T. M., Bower, T. C., Gloviczki, P., Miller, D. V., Babovic-Vuksanovic, D. et al. Vascular abnormalities in patients with neurofibromatosis syndrome type I: clinical spectrum, management, and results. J. Vasc. Surg. 46, 475–484 (2007).
Lama, G., Graziano, L., Calabrese, E., Grassia, C., Rambaldi, P. F., Cioce, F. et al. Blood pressure and cardiovascular involvement in children with neurofibromatosis type1. Pediatr. Nephrol. 19, 413–418 (2004).
Brems, H., Pasmant, E., Van Minkelen, R., Wimmer, K., Upadhyaya, M., Legius, E. et al. Review and update of SPRED1 mutations causing Legius syndrome. Hum. Mutat. 33, 1538–1546 (2012).
Rauen, K. A., Banerjee, A., Bishop, W. R., Lauchle, J. O., McCormick, F., McMahon, M. et al. Costello and cardio-facio-cutaneous syndromes: Moving toward clinical trials in RASopathies. Am. J. Med. Genet. C Semin. Med. Genet. 157, 136–146 (2011).
Krab, L. C., de Goede-Bolder, A., Aarsen, F. K., Pluijm, S. M., Bouman, M. J., van der Geest, J. N. et al. Effect of simvastatin on cognitive functioning in children with neurofibromatosis type 1: a randomized controlled trial. JAMA 300, 287–294 (2008).
Chen, P. C., Wakimoto, H., Conner, D., Araki, T., Yuan, T., Roberts, A. et al. Activation of multiple signaling pathways causes developmental defects in mice with a Noonan syndrome-associated Sos1 mutation. J. Clin. Invest. 120, 4353–4365 (2010).
Wu, X., Simpson, J., Hong, J. H., Kim, K. H., Thavarajah, N. K., Backx, P. H. et al. MEK-ERK pathway modulation ameliorates disease phenotypes in a mouse model of Noonan syndrome associated with the Raf1(L613V) mutation. J. Clin. Invest. 121, 1009–1025 (2011).
Inoue, S. I., Moriya, M., Watanabe, Y., Miyagawa-Tomita, S., Niihori, T., Oba, D. et al. New BRAF knockin mice provide a pathogenetic mechanism of developmental defects and a therapeutic approach in cardio-facio-cutaneous syndrome. Hum. Mol. Genet. 23, 6553–6566 (2014).
Marin, T. M., Keith, K., Davies, B., Conner, D. A., Guha, P., Kalaitzidis, D. et al. Rapamycin reverses hypertrophic cardiomyopathy in a mouse model of LEOPARD syndrome-associated PTPN11 mutation. J. Clin. Invest. 121, 1026–1043 (2011).
Hahn, A., Lauriol, J., Thul, J., Behnke-Hall, K., Logeswaran, T., Schanzer, A. et al. Rapidly progressive hypertrophic cardiomyopathy in an infant with Noonan syndrome with multiple lentigines: palliative treatment with a rapamycin analog. Am. J. Med. Genet. A 167, 744–751 (2015).
Bayrak-Toydemir, P. & Stevenson, D. RASA1-related disorders. GeneReviews® [Internet] (University of Washington, Seattle, WA, 2014).
Revencu, N., Boon, L. M., Mendola, A., Cordisco, M. R., Dubois, J., Clapuyt, P. et al. RASA1 mutations and associated phenotypes in 68 families with capillary malformation-arteriovenous malformation. Hum. Mutat. 34, 1632–1641 (2013).
Acknowledgements
We thank members of the Department of Medical Genetics, Tohoku University School of Medicine, for contributing to RASopathy diagnostics and research. We are grateful to patients with RASopathies and their families and to the doctors who participated in our studies. This work was supported in part by grants from the Ministry of Health, Labour and Welfare of Japan, from the Practical Research Project for Rare/Intractable Diseases from Japan Agency for Medical Research and development, AMED, and from the Japan Society for the Promotion of Science (a Grant-in-Aid for Scientific Research (B)).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Aoki, Y., Niihori, T., Inoue, Si. et al. Recent advances in RASopathies. J Hum Genet 61, 33–39 (2016). https://doi.org/10.1038/jhg.2015.114
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2015.114
This article is cited by
-
Neurofibromatosis-Noonan syndrome and growth deficiency in an Iranian girl due to a pathogenic variant in NF1 gene
Human Genomics (2023)
-
LZTR1 deficiency exerts high metastatic potential by enhancing sensitivity to EMT induction and controlling KLHL12-mediated collagen secretion
Cell Death & Disease (2023)
-
The exocyst complex in neurological disorders
Human Genetics (2023)
-
Refractory thrombocytopenia could be a rare initial presentation of Noonan syndrome in newborn infants: a case report and literature review
BMC Pediatrics (2022)
-
New insights on Noonan syndrome’s clinical phenotype: a single center retrospective study
BMC Pediatrics (2022)