Abstract
The genomic regions containing PNPLA3, SAMM50 and PARVB are susceptibility loci for the development and progression of nonalcoholic fatty liver disease (NAFLD). In order to search for all common variations in this region, we amplified the genomic DNA of 28 NAFLD patients by long-range PCR, covering the entire susceptibility region and sequenced the DNA using indexed multiplex next-generation sequencing. We found 329 variations, including four novel variations. Fine mapping of variations including insertion/deletions was performed for 540 NAFLD patients (488 with nonalcoholic steatohepatitis (NASH) and 52 with simple steatosis) and 1012 control subjects. HaploView analysis showed that linkage disequilibrium (LD) block 1 and 2 occurred in PNPLA3, block 3 in SAMM50 and block 4 in PARVB. Variations in LD blocks 1–4 were significantly associated with NAFLD as compared with control subjects (P<1 × 10−8). Variations in LD block 2 were significantly associated with the NAFLD activity score (NAS), aspartate aminotransferase and alanine aminotransferase. Variations in LD block 1 were significantly associated with the fibrosis stage. The strongest associations were observed for variations in LD block 4, with NASH as compared with simple steatosis (P=7.1 × 10−6) and NAS (P=3.4 × 10−6). Our results suggested that variations, including insertion/deletions, in PARVB, as well as those in PNPLA3, are important in the progression of NAFLD.
Similar content being viewed by others
Introduction
Recently, nonalcoholic fatty liver disease (NAFLD) was recognized as an important health concern.1, 2 NAFLD includes simple steatosis, nonalcoholic steatohepatitis (NASH), fibrosis/cirrhosis and hepatocellular carcinoma. The frequency of patients presenting with NAFLD has increased gradually in Japan in proportion to the increase in the population with metabolic syndrome.3 NAFLD is observed in 20–30% of the population in Japan and ∼1–3% of these individuals are considered to have NASH, similar to American and European populations.3, 4
In addition to environmental factors, genetic factors are important in the development of NAFLD.5 In order to elucidate these genetic factors, we previously investigated candidate genes and found that variations in the peroxisome proliferator-activated receptor γ coactivator 1α (PPARGC1A), angiotensin II type 1 receptor (AGTR1), and nitric oxide synthase 2 (inducible) (NOS2) genes are associated with NAFLD in Japanese individuals.6, 7, 8 Genome-wide association studies have revealed that single-nucleotide polymorphisms (SNPs) in the patatin-like phospholipase domain containing 3 (PNPLA3) influence NAFLD and liver enzyme levels in the plasma.9, 10, 11, 12 We also reported that the risk allele (G-allele) of rs738409 in PNPLA3 is strongly associated with NAFLD, as well as with increases in aspartate aminotransferase (AST), alanine aminotransferase (ALT) and fibrosis stage in patients with NAFLD in the Japanese population.13
Recently, we performed a GWAS and found that polymorphisms in the SAMM50 sorting and assembly machinery component (SAMM50) and parvin-β (PARVB) genes, in addition to those in PNPLA3 were associated with the development and progression of NAFLD.14 Variations used for GWAS were limited and at least 10% of SNPs were monomorphic or rare SNPs in the Japanese population. Many variations in the NAFLD susceptibility genomic region have not been investigated yet. Here, we performed targeted resequencing to identify all common variations in the entire NAFLD susceptibility genomic region. We constructed a fine linkage disequilibrium (LD) map of SNPs and insertion/deletions in the region and analyzed their association with the development and severity of NAFLD.
Materials and methods
Study subjects
The entire study was conducted in accordance with the guidelines of the Declaration of Helsinki. Written informed consent was obtained from each subject, and the protocol was approved by the ethics committee of Kyoto University, Yokohama City University, Hiroshima University and Kurume University.
Control subjects (n=1012) included Japanese volunteers who had undergone medical examination for common disease screening. Japanese patients with NAFLD who underwent liver biopsy (488 with NASH and 52 with simple steatosis) were enrolled. Patients with the following diseases were excluded from the study: viral hepatitis (hepatitis B and C, Epstein–Barr virus infection), autoimmune hepatitis, primary biliary cirrhosis, sclerosing cholangitis, hemochromatosis, α1-antitrypsin deficiency, Wilson’s disease, drug-induced hepatitis and alcoholic hepatitis (present or past daily consumption of more than 20 g alcohol per day). None of the patients showed clinical evidence of hepatic decompensation, such as hepatic encephalopathy, ascites, variceal bleeding or a serum bilirubin level greater than twofold the normal upper limit.
Liver biopsy tissues were stained with hematoxylin and eosin, reticulin and Masson’s trichrome stain. The histological criterion for NAFLD diagnosis is macrovesicular fatty change in hepatocytes with displacement of the nucleus toward the cell edge.15 Patients who had more than 5% of hepatocytes affected by macrovesicular steatosis were diagnosed as having either steatosis or NASH. The minimal criteria for the diagnosis of NASH were the presence of >5% macrovesicular steatosis, inflammation and liver cell ballooning, typically with predominantly centrilobular (acinar zone 3) distribution.16, 17 The degree of steatosis was graded as follows based on the percentage of hepatocytes containing macrovesicular fat droplets: grade 0, no steatosis; grade 1, <33% hepatocytes containing macrovesicular fat droplets; grade 2, 33–66% of hepatocytes containing macrovesicular fat droplets; and grade 3, >66% of hepatocytes containing macrovesicular fat droplets.18 The hepatitis activity (necroinflammatory grade) was also determined on the basis of the composite NAFLD activity score: the unweighted sum of the scores for steatosis, lobular inflammation and hepatocellular ballooning (ranging from 0 to 8).19 Fibrosis severity was scored according to the method of Brunt18 and was expressed on a 4-point scale as follows: 0, none; 1, perivenular and/or perisinusoidal fibrosis in zone 3; 2, combined pericellular portal fibrosis; 3, septal/bridging fibrosis; and 4, cirrhosis.
Clinical and laboratory evaluation
The weight and height of the patients were measured using a calibrated scale after removing shoes and heavy clothing. Venous blood samples were obtained from subjects after overnight fasting (12 h) to measure plasma glucose, hemoglobin A1c, total cholesterol, high-density lipoprotein cholesterol, triglycerides, serum AST and ALT levels. All laboratory biochemical parameters were measured using conventional methods.
Targeted resequencing
Genomic DNA was extracted using Genomix (Talent Srl, Trieste, Italy), from blood samples collected from each subject. Genomic DNA of 28 randomly selected NAFLD patients was used for targeted resequencing. Clinical information of these 28 patients is shown in Supplementary Table 1. The genomic region from 44,317,888 to 44,425,903 on chromosome 22, which includes PNPLA3, SAMM50 and PARVB, was amplified using region-specific primers with a long-range PCR approach (Supplementary Table 2). Amplification was performed using the GeneAmp 9700 PCR System (Life Technologies, Carlsbad, CA, USA), with 50 ng of genomic DNA, 1 × PCR buffer (TOYOBO, Osaka, Japan), 0.4 mM of each dNTP, 0.3 μM of each primer and 0.02 U of KOD FX Neo (TOYOBO, Osaka, Japan) in a 14-μl reaction volume. After an initial denaturation at 94 °C for 2 min, PCR was performed with 35 cycles, each consisting of 98 °C for 10 s and 68 °C for 5–8 min, depending on the length of the amplicons. The PCR extension time is shown in Supplementary Table 2. All 16 PCR amplicons were confirmed as a single peak (Supplementary Figure 1), and the molar concentration was measured using an Agilent DNA 12000 Kit and 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA); the amplicons were then pooled at equal concentration for all individuals.
Pooled PCR products were shared using a Covaris E210 (COVARIS, Woburn, MA, USA) under the following conditions: 10% duty cycle, intensity of 5.0, 200 cycles per burst, 40-s duration and frequency sweeping mode. Each sample was dual indexed and DNA libraries were prepared using the TruSeq DNA Sample Preparation kit v2 (Illumina, San Diego, CA, USA), according to the manufacturer’s protocol. Sequencing was performed in a MiSeq system (Illumina) using 150-bp paired-end reads. We aligned the paired-end reads to the human genome (UCSC hg19) as a reference, using BWA (version 0.5.9rc1; http://www.bio-bwa.sourceforge.net/).20 For sequence data conversion, sorting and indexing, we used SAMtools (version 0.1.17; http://www.samtools.sourceforge.net/) and Picard (version 1.72; http://www.picard.sourceforge.net).21 The variants, including insertion/deletions and SNPs, were determined using GATK (version 1.6-5-g557da77; http://www.broadinstitute.org/gatk/).22 ANNOVAR (version 2012-05-25; http://www.openbioinformatics.org/annovar/) was used to appropriately annotate genetic variants obtained.23 The variations were confirmed using IGV (Supplementary Figure 2) (version 1.5.65, http://www.broadinstitute.org/igv/).24, 25
Fine LD mapping of the PNPLA3, SAMM50 and PARVB loci
Invader probes (Third Wave Technologies, Madison, WI, USA) were constructed for 192 SNPs and 9 insertion/deletions located on chromosome 22:44,321,410 to 44,417,970, which had minor allele frequencies >0.05, as estimated from the results of targeted resequencing. An invader probe for one insertion/deletion (rs67877195) could not be constructed. SNPs and insertion/deletions were genotyped using Invader assays, as previously described.26 Among the 200 variations, 173 SNPs and 7 insertion/deletions were successfully genotyped. The success rates of the 180 Invader assays were >99.0% and the concordance rate of these results with those of sequencing was >99.4%.
Statistical analysis
LD was drawn using HaploView (http://www.broad.mit.edu/mpg/haploview/).27 We categorized the genotypes as 0, 1 or 2, depending on the number of copies of risk alleles present. Odds ratios and P-values, adjusted for age, sex, body mass index (BMI) and the presence of type 2 diabetes mellitus, were calculated using multiple logistic regression analysis. Multiple linear regression analyses were performed to test the independent effect per allele of each SNP on biochemical traits and histological parameters, accounting for effects of the other variables (that is, age, sex, BMI and the presence of type 2 diabetes mellitus). BMI, AST and ALT values were logarithmically transformed before performing multiple linear and logistic regression analyses. Statistical analyses were performed using PLINK 1.07 (http://pngu.mgh.harvard.edu/purcell/plink)28 and R software (http://www.r-project.org/). P-values less than 3.0 × 10−4 (0.05/169) was considered to be significant.
Results
Targeted resequencing
GWAS indicated that a strong NAFLD susceptibility locus is present in the region from 4.3 kb downstream of the PNPLA3 (NM_025225.2) transcription start site to 7.5 kb downstream of the PARVB isoform a (NM_001003828.2) transcription start site.11, 12, 14 We amplified the ∼100-kb genomic region, from 1.7 kb upstream of PNPLA3 to 30 kb downstream of the PARVB isoform a transcription start site, using long-range PCR (3.8–13.6 kb, Supplementary Table 2). Sixteen PCR products covered the entire genomic region.
PCR amplicons prepared from 28 NAFLD patients were sequenced using indexed multiplex next-generation sequencing. The average depth of the individual sequencing ranged from 387 to 652. An example of this sequencing is shown in Supplementary Figure 1. All observed variations are listed in Supplementary Table 3. A total of 329 variations were found and most of these (325 variations) were already registered in the dbSNP database. Insertion/deletions accounted for 16 variations, and SNPs for 313. Four novel SNPs were found in this study (ss831884188, ss831884189, ss831884190 and ss831884191). No clear genomic structural variations (>50 nucleotides in length) were observed. A 13-nucleotide insertion/deletion (rs140963094) was the longest variation.
Fine variation LD mapping
According to the HapMap database, SNPs from rs58002102 (chromosome 22:44,321,410) to rs135114 (chromosome 22:44,417,970) were sufficient to produce a fine LD map in the NAFLD susceptibility genomic region. There were 280 variations in this region (Supplementary Table 3), including 192 SNPs and 9 insertion/deletions with minor allele frequencies >0.05 in the 28 NAFLD patients. The latter 201 variations with minor allele frequencies >0.05 were genotyped in 540 NAFLD patients and 1012 control subjects using Invader assays; 173 SNPs and 7 insertion/deletions (rs5845621, rs71218095, rs71313378, rs140963094, rs10656207, rs76409096 and rs34505405) were successfully identified. Genotyped variations and their allele frequencies are shown in Supplementary Tables 3 and 4.
We used 162 SNPs and 7 insertion/deletions that had minor allele frequencies >0.10 for further analysis. Fine LD mapping using 169 variation data derived from 540 NAFLD patients is shown in Figure 1 and r2 values in control and NAFLD subjects are given in Supplementary Figure 2. HaploView analysis showed that four LD blocks existed in the NAFLD susceptibility genomic region in this population: a 1-kb LD block 1 (rs738407–rs2006943), a 21-kb LD block 2 (rs139051–rs13054885), a 48-kb LD block 3 (rs7289329–rs1007863) and a 2-kb LD block 4 (rs5764455–rs6006611). LD block 1 and block 2 occurred within PNPLA3, block 3 within SAMM50 and block 4 within PARVB (Figure 1). LD blocks were the same in the NAFLD patients and control subjects (Supplementary Figure 2). Seven insertion/deletions were in LD with the SNPs.
Association of variations with severity of NAFLD
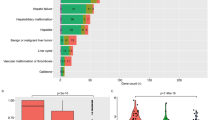
First, we examined the association of the above 169 variations with simple steatosis and NASH. Clinical characteristics of the simple steatosis and NASH patients are shown in Table 1. We performed multiple logistic regression analysis using genotypes, age, sex, BMI and the presence of type 2 diabetes as independent variables. Two SNPs, namely, rs6006610 (P=6.1 × 10−5) and rs6006611 (P=7.1 × 10−6) in LD block 4, and rs2006943 (P=5.6 × 10−5) in LD block 1, were significantly associated with NASH as compared with simple steatosis (Figure 2, Supplementary Table 4). Relatively high odds ratios were observed: rs6006611, 2.68 (1.74–4.12); rs6006610, 2.37 (1.55–3.62); rs2006943, 2.54 (1.62–4.00) (OR (95% confidence interval)). As the sample size of simple steatosis is small (n=52), we compared 540 NAFLD patients and 488 NASH patients with 1012 control subjects. As reported previously, most of the variations in LD blocks 1–4 were significantly associated with NAFLD (Figure 2, Supplementary Table 4). Stronger association and higher odds ratios between NASH and control individuals were observed in LD block 1–4, especially in block 4 (Figure 2). The risk allele frequencies of these SNPs were highest in the NASH group, and those of the simple steatosis and control group were very similar (Supplementary Table 4). No associations were observed for the simple steatosis and control groups (P>0.02). These data suggested that the variations in LD block 4 are more strongly associated with NASH, rather than with NAFLD and simple steatosis.
Regional plots of the −log10(P-value) for the association of variations. Variations are plotted by their position on chromosome 22 against their association with nonalcoholic steatohepatitis (NASH) compared with simple steatosis, nonalcoholic fatty liver disease (NAFLD) to control and NASH to control. Data were derived by logistic regression analysis, adjusted for age, sex, logarithmically transformed body mass index and the presence of type 2 diabetes. The positions of genes as well as the direction of transcription are shown above the plots. Black, red, blue and green closed circles are variations in linkage disequilibrium (LD) block 1, 2, 3 and 4, respectively. Gray closed circles are variations in other LD blocks.
Next, we examined the association of the 169 variations with histological phenotypes. Linear regression analysis, adjusted for age, sex, BMI and the presence of type 2 diabetes, revealed that no variations were significantly associated with lobular inflammation (Figure 3, Supplementary Table 5). Only one SNP (rs12484700) was significantly associated with steatosis grade (P=2.2 × 10−4). Two SNPs (rs6006610, P=1.9 × 10−4; rs6006611, P=4.4 × 10−5) in LD block 4 were significantly associated with hepatocyte ballooning. Many variations in LD blocks 2, 3 and 4 were associated with a higher NAFLD activity score (Figure 3, Supplementary Table 5). The most significant variation was rs6006611 (P=3.4 × 10−6) in LD block 4. Only rs734561 (P=2.6 × 10−4) and rs2006943 (P=1.9 × 10−4) in LD block 1 were significantly associated with a higher fibrosis stage (Figure 3, Supplementary Table 5).
Regional plot for histological scores. Data were derived by linear regression analysis for individual histological scores for steatosis grade, lobular inflammation, hepatocyte ballooning, nonalcoholic fatty liver disease activity score (NAS) and fibrosis stage. Each phenotype was adjusted for age, sex, logarithmically transformed body mass index and the presence of type 2 diabetes. The positions of genes as well as the direction of transcription are shown above the plots. Black, red, blue and green closed circles are variations in LD block 1, 2, 3 and 4, respectively. Gray closed circles are variations in other linkage disequilibrium (LD) blocks.
Finally, we investigated the association of these variations with AST and ALT levels. We observed that many of the variations in LD block 2 were significantly associated with both AST and ALT levels (Figure 4, Supplementary Table 5). Many variations in LD blocks 1, 3 and 4 were also significantly associated with AST and ALT levels; however, these associations were much stronger with variations in LD block 2.
Regional plot for aspartate aminotransferase (AST) and alanine aminotransferase (ALT). Data were derived by linear regression analysis. Values of AST and ALT levels were logarithmically transformed. Each phenotype was adjusted for age, gender, logarithmically transformed body mass index and the presence of type 2 diabetes. The positions of genes as well as the direction of transcription are shown above the plots. Black, red, blue and green closed circles are variations in linkage disequilibrium (LD) block 1, 2, 3 and 4, respectively. Gray closed circles are variations in other LD blocks.
Discussion
After the first report of an association of PNPLA3 rs738409 with NAFLD,9 many replication studies were performed in various populations,10, 11, 12, 13, 14, 29, 30 verifying the importance of rs738409 in the development of NAFLD. The LD block containing rs738409 spans ∼80 kb in the Japanese population. Many variations in the LD block containing rs738409 are registered in the dbSNP database; however, many of these were invalid, and no genomic structural variations were reported. Therefore, we performed targeted resequencing to identify and confirm variations in this region. We did not find any genomic structural variations in this region. For sequencing, only 28 NAFLD subjects were used; it is therefore possible that genomic structural variations may occur in this region at a low frequency. Using a combination of targeted resequencing and genotyping methods, we determined the frequencies and the associations of most common variations in this region, including insertion/deletions, with the severity of NAFLD. This approach would be useful for investigating other common disease susceptibility loci in detail.
Fine LD mapping revealed that 4 LD blocks covered the NAFLD susceptibility locus and that PNPLA3 was contained in LD blocks 1 and 2, SAMM50 in LD block 3 and PARVB in LD block 4. As LD blocks were separated clearly by genes, the genetic effect of each gene on the development and progression of NAFLD could be analyzed. In this study, we showed that many variations in PNPLA3 (LD blocks 1 and 2), as well as rs738409 in LD block 2, were significantly associated with NASH, and with the severity of NAFLD (NAFLD activity score, AST and ALT). These associations were stronger with the 5′-region of LD block 2. Fibrosis was significantly associated with variations in LD block 1, which did not contain rs738409. The variant, rs738409, is a C (non-risk allele) to G (risk allele) substitution that changes codon 148 from isoleucine (I) to methionine (M). Interestingly, PNPLA3-deficient mice did not show a fatty liver,31, 32 but hepatic overexpression of PNPLA3I148M led to development of a fatty liver, although overexpression of wild-type PNPLA3 in mouse liver did not.33 Our study and other studies in mice suggest that variations in LD with rs738409 and/or variations in LD block 1 of PNPLA3 could increase the transcriptional level of mRNA containing the risk allele of rs738409, leading to the development of NAFLD. LD block 2 contains the insertion/deletion rs71218095 that is located in intron 5 of PNPLA3, near exon 6 (pairwise LD statistics r2=0.93 with rs738409). The effect of the insertion/deletion would affect transcription of mRNA more strongly.
Overexpression of PNPLA3I148M in mice led to the development of fatty liver, but not to NASH and metabolic disorders,33 suggesting the contribution of other genes to the latter conditions. In our study, the strongest associations were observed between variations in PARVB (LD block 4) and NASH, and histological severity (NAFLD activity score). PARVB has at least four transcript variants; LD block 4 is located in the 2nd intron of the longest transcript variant of PARVB, corresponding to the 5′-flanking region of the shorter transcript variants. PARVB encodes parvin-β, which forms part of an integrin-linked kinase–pinch–parvin complex that transmits signals from integrin to Akt/protein kinase B.34 Overexpression of parvin-β increases mRNA expression levels, phosphorylation at serine 82 and activity of the peroxisome proliferator-activated receptor γ, leading to a concomitant increase in lipogenic gene expression.35 Another report has indicated that overexpression of parvin-β promotes apoptosis.36 Increased lipogenesis and apoptosis are considered to underlie fibrosis in NAFLD;37, 38 therefore, variations in PARVB (LD block 4) could increase the transcriptional level of parvin-β, leading to increased lipogenesis and apoptosis, resulting in the progression of fibrosis.
In conclusion, both our previous GWAS and the present study indicate that PNPLA3 is most important in the development of simple steatosis. We also showed here that PARVB, in addition to PNPLA3, has an important role in the progression of simple steatosis to NASH. The limitation of this study was a small sample size of individuals with simple steatosis; therefore, further studies would be necessary. Nevertheless, the approach used here of a combination of targeted resequencing and genotyping methods could be applied for the detailed investigation of other common disease susceptibility loci.
References
Angulo, P Nonalcoholic fatty liver disease. New Engl. J. Med. 18, 1221–1231 (2002).
Farrell, GC Non-alcoholic steatohepatitis: what is it, and why is it important in the Asia-Pacific region? J. Gastroenterol. Hepatol. 18, 124–138 (2003).
Okanoue, T, Umemura, A, Yasui, K & Itoh, Y Nonalcoholic fatty liver disease and nonalcoholic steatohepatitis in Japan. J. Gastroenterol. Hepatol. Suppl 1, 153–162 (2011).
Ludwig, J, Viggiano, TR, McGill, DB & Oh, BJ Nonalcoholic steatohepatitis: Mayo Clinic experiences with a hitherto unnamed disease. Mayo Clin. Proc. 55, 434–438 (1980).
Wilfred de Alwis, NM & Day, CP Genes and nonalcoholic fatty liver disease. Curr. Diab. Rep. 8, 156–163 (2008).
Yoneda, M, Hotta, K, Nozaki, Y, Endo, H, Uchiyama, T, Mawatari, H et al. Association between PPARGC1A polymorphisms and the occurrence of nonalcoholic fatty liver disease (NAFLD). BMC Gastroenterol. 8, 27 (2008).
Yoneda, M, Hotta, K, Nozaki, Y, Endo, H, Uchiyama, T, Mawatari, H et al. Association between angiotensin II type 1 receptor polymorphisms and the occurrence of nonalcoholic fatty liver disease. Liver Int. 29, 1078–1085 (2009).
Yoneda, M, Hotta, K, Nozaki, Y, Endo, H, Tomeno, W, Watanabe, S et al. Influence of inducible nitric oxide synthase polymorphisms in Japanese patients with non-alcoholic fatty liver disease. Hepatol. Res. 39, 963–971 (2009).
Romeo, S, Kozlitina, J, Xing, C, Pertsemlidis, A, Cox, D, Pennacchio, LA et al. Genetic variation in PNPLA3 confers susceptibility to nonalcoholic fatty liver disease. Nat. Genet. 40, 1461–1465 (2008).
Chalasani, N, Guo, X, Loomba, R, Goodarzi, MO, Haritunians, T, Kwon, S et al. Genome-wide association study identifies variants associated with histologic features of nonalcoholic fatty liver disease. Gastroenterology 139, 1567–1576 (2010).
Speliotes, EK, Yerges-Armstrong, LM, Wu, J, Hernaez, R, Kim, LJ, Palmer, CD et al. Genome-wide association analysis identifies variants associated with nonalcoholic fatty liver disease that have distinct effects on metabolic traits. PLoS Genet. 7, e1001324 (2011).
Kawaguchi, T, Sumida, Y, Umemura, A, Matsuo, K, Takahashi, M, Takamura, T et al. Genetic polymorphisms of the human PNPLA3 gene are strongly associated with severity of non-alcoholic fatty liver disease in Japanese. PLoS One 7, e38322 (2012).
Hotta, K, Yoneda, M, Hyogo, H, Ochi, H, Mizusawa, S, Ueno, T et al. Association of the rs738409 polymorphism in PNPLA3 with liver damage and the development of nonalcoholic fatty liver disease. BMC Med. Genet. 11, 172 (2010).
Kitamoto, T, Kitamoto, A, Yoneda, M, Hyogo, H, Ochi, H, Nakamura, T et al. Genome-wide scan revealed that polymorphisms in the PNPLA3, SAMM50, and PARVB genes are associated with development and progression of nonalcoholic fatty liver disease in Japan. Hum. Genet. 132, 783–792 (2013).
Sanyal, AJ American Gastroenterological Association: AGA technical review on nonalcoholic fatty liver disease. Gastroenterology 123, 1705–1725 (2002).
Matteoni, CA, Younossi, ZM, Gramlich, T, Boparai, N, Liu, YC & McCullough, AJ Nonalcoholic fatty liver disease: a spectrum of clinical and pathological severity. Gastroenterology 116, 1413–1419 (1999).
Teli, MR, James, OF, Burt, AD, Bennett, MK & Day, CP The natural history of nonalcoholic fatty liver: a follow-up study. Hepatology 22, 1714–1719 (1995).
Brunt, EM Nonalcoholic steatohepatitis: definition and pathology. Semin. Liver Dis. 21, 3–16 (2001).
Kleiner, DE, Brunt, EM, Van Natta, M, Behling, C, Contos, MJ, Cummings, OW et al. Design and validation of a histological scoring system for nonalcoholic fatty liver disease. Hepatology 41, 1313–1321 (2005).
Li, H & Durbin, R Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H, Handsaker, B, Wysoker, A, Fennell, T, Ruan, J, Homer, N et al. The sequence alignment/map (SAM) format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
McKenna, A, Hanna, M, Banks, E, Sivachenko, A, Cibulskis, K, Kernytsky, A et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Wang, K, Li, M & Hakonarson, H ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Robinson, JT, Thorvaldsdóttir, H, Winckler, W, Guttman, M, Lander, ES, Getz, G et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Thorvaldsdóttir, H, Robinson, JT & Mesirov, JP Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform. 14, 178–192 (2013).
Ohnishi, Y, Tanaka, T, Ozaki, K, Yamada, R, Suzuki, H & Nakamura, Y A high-throughput SNP typing system for genome-wide association studies. J. Hum. Genet. 46, 471–477 (2001).
Barrett, JC, Fry, B, Maller, J & Daly, MJ Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21, 263–265 (2005).
Purcell, S, Neale, B, Todd-Brown, K, Thomas, L, Ferreira, MA, Bender, D et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Sookoian, S & Pirola, CJ Meta-analysis of the influence of I148M variant of patatin-like phospholipase domain containing 3 gene (PNPLA3) on the susceptibility and histological severity of nonalcoholic fatty liver disease. Hepatology 53, 1883–1894 (2011).
Zain, SM, Mohamed, R, Mahadeva, S, Cheah, PL, Rampal, S, Basu, RC et al. A multi-ethnic study of a PNPLA3 gene variant and its association with disease severity in non-alcoholic fatty liver disease. Hum. Genet. 131, 1145–1152 (2012).
Chen, W, Chang, B, Li, L & Chan, L Patatin-like phospholipase domain-containing 3/adiponutrin deficiency in mice is not associated with fatty liver disease. Hepatology 52, 1134–1142 (2010).
Basantani, MK, Sitnick, MT, Cai, L, Brenner, DS, Gardner, NP, Li, JZ et al. Pnpla3/Adiponutrin deficiency in mice does not contribute to fatty liver disease or metabolic syndrome. J. Lipid Res. 52, 318–329 (2011).
Li, JZ, Huang, Y, Karaman, R, Ivanova, PT, Brown, HA, Roddy, T et al. Chronic overexpression of PNPLA3I148M in mouse liver causes hepatic steatosis. J. Clin. Invest. 122, 4130–4144 (2012).
Kimura, M, Murakami, T, Kizaka-Kondoh, S, Itoh, M, Yamamoto, K, Hojo, Y et al. Functional molecular imaging of ILK-mediated Akt/PKB signaling cascades and the associated role of beta-parvin. J. Cell Sci. 123, 747–755 (2010).
Johnstone, CN, Mongroo, PS, Rich, AS, Schupp, M, Bowser, MJ, Delemos, AS et al. Parvin-beta inhibits breast cancer tumorigenicity and promotes CDK9-mediated peroxisome proliferator-activated receptor gamma 1 phosphorylation. Mol. Cell Biol. 28, 687–704 (2008).
Zhang, Y, Chen, K, Tu, Y & Wu, C Distinct roles of two structurally closely related focal adhesion proteins, alpha-parvins and beta-parvins, in regulation of cell morphology and survival. J. Biol. Chem. 279, 41695–41705 (2004).
Chakraborty, JB, Oakley, F & Walsh, MJ Mechanisms and biomarkers of apoptosis in liver disease and fibrosis. Int. J. Hepatol. 2012, 648915 (2012).
Fujii, H & Kawada, N Inflammation and fibrogenesis in steatohepatitis. J. Gastroenterol. 47, 215–225 (2012).
Acknowledgements
This work was supported by a Grant-in-Aid from the Ministry of Education, Science, Sports, and Culture of Japan (25461343 to KH, 23791027 to AK and 23701082 to TK).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Rights and permissions
About this article
Cite this article
Kitamoto, T., Kitamoto, A., Yoneda, M. et al. Targeted next-generation sequencing and fine linkage disequilibrium mapping reveals association of PNPLA3 and PARVB with the severity of nonalcoholic fatty liver disease. J Hum Genet 59, 241–246 (2014). https://doi.org/10.1038/jhg.2014.17
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2014.17
Keywords
This article is cited by
-
Two differentially methylated region networks in nonalcoholic fatty liver disease, viral hepatitis, and hepatocellular carcinoma
BMC Gastroenterology (2022)
-
Identification of differentially methylated region (DMR) networks associated with progression of nonalcoholic fatty liver disease
Scientific Reports (2018)
-
Copy number variation in exportin-4 (XPO4) gene and its association with histological severity of non-alcoholic fatty liver disease
Scientific Reports (2015)