Abstract
Breast Cancer is the most common malignancy among women. Family history is the strongest single predictor of breast cancer risk, and thus great attention has been focused on BRCA1 and BRCA2 genes whose mutations lead to a high risk of developing this disease. Today, only 25% of high- and moderate-risk genes are known, suggesting the importance of the discovery of new risk modifiers. Therefore, the investigation of new polygenic alterations is of great importance, especially if considered high- and moderate-risk variants. In this study, the transmission of BRCA1-2 polymorphisms in association with the transmission of polymorphisms in the genes NUMA1, CCND1, COX11, FGFR2, TNRC9 and SLC4A7 were examined in all members of a family with the BRCA2 c.6447_6448dup mutation. This is the first study about the transmission of high-risk polygenic variants in all members of a family with a strong history of breast cancer. The results about the possible polygenic variant associations that could increase and modify the risk suggested the importance to search new variants to better manage patients and their family members.
Similar content being viewed by others
Main
Breast cancer is a hereditary disease in almost 10% of cases and is associated with mutations in BRCA1 and BRCA2 genes.
The BRCA1-2 screening has an important impact on patients and healthy family members for prognostic and prophylactic approaches because mutations in these genes increase the risk of developing breast and/or ovarian cancer. Today, only 25% of high- and moderate-risk genes are known, reflecting the existence of multiple distinct susceptibility factors and of many unknown variants of risk modifiers. Therefore, the majority of diagnostic genetic tests performed is uninformative and does not contribute to understand the familial breast cancer risk in the families.1
Our previous study2 showed that multiple single-nucleotide polymorphisms (SNPs) in BRCA1 or BRCA2, belonging to a certain haplotype, could be associated to an increased risk also in hereditary breast cancer. Specific haplotypes were transmitted in both mutated and non-mutated families, suggesting the importance of segregation analysis as the logical basis and the specific genetic model for mapping and identifying regions in genes increasing the breast cancer risk.
To our knowledge, genome-wide association (GWA) studies used a large number of common SNPs to identify associations with diseases. However, the limitation of these published studies is that only the genotype of index cases were analyzed, not considering the transmission model of the variants in all family members.3, 4
The aim of this study was to search a possible transmission association of polygenic variants that increase the familial breast cancer susceptibility. This study deals with the transmission of polymorphisms of BRCA1 and BRCA2 genes and of NUMA1, CCND1, COX11, FGFR2, TNRC9 and SLC4A7 genes in a family with the index case affected by BRCA2-mutated (c.6447_6448dup) heredo-familial breast cancer and his wife with a sporadic breast tumor wild type for BRCA1 and BRCA2 genes. This BRCA2 mutation has been reported only in non-Afrikaner breast cancer patients of the Western Cape of South Africa5 and it has never been reported in the Italian population. The polymorphisms in the genes NUMA1, CCND1, COX11, FGFR2, TNRC9 and SLC4A7 were reported in many GWA studies as breast cancer risk modifiers.6, 7, 8
The patients and all available family members were enrolled by the IRCCS National Cancer Centre “Giovanni Paolo II” of Bari after signing an informed consent. Molecular analyses were performed as previously described.2, 9, 10, 11 BRCA1/2 genes of the index case and his wife were analyzed through capillary sequencing. All available family members were screened for the BRCA2 mutation and for all polymorphic variants in BRCA1 and BRCA2 found in their parents. Moreover, all members were analyzed for the polymorphisms in the genes NUMA1, CCND1, COX11, FGFR2, TNRC9 and SLC4A7 through real-time PCR. The pedigree with family’s information was designed by the Progeny v6 Program. The BIC online database (http://www.research.nhgri.nih.gov/bic/), the LOVD-IARC database (http://brca.iarc.fr/LOVD/home.php?select_db=BRCA2), the UMD database (http://www.umd.be) and the bioinformatics tools (http://www.expasy.org/; http://www.ensembl.org) were consulted to thoroughly investigate the detected BRCA1–2 alterations.
The index case is a 64-year-old man with familial breast cancer developed when he was 58 years of age; he underwent a left nodulectomy resulting in infiltrating ductal carcinoma (IDC) of grade 3, estrogen receptor (ER), progesterone receptor (PgR) and Her2-neu fluorescent in situ hybridization (FISH) positive. He was also affected by multiple sclerosis and was treated with chemotherapy for four cycles. To date, only a single published paper suggested the association between the polymorphism rs1801406 in the BRCA2 gene and the risk to develop a secondary acute promyelocytic leukemia in patient affected by multiple sclerosis.12 During counseling, the index case mentioned other cancer cases in his family. His grandmother and his mother died because of breast cancer, whereas his uncle developed prostate cancer. The index has three daughters (one affected by breast cancer) and one son. His daughter developed breast cancer at 34 years of age, and histopathological analysis showed an IDC of grade 2, also ER, PgR and Her2-neu FISH positive. In the last years, she also developed multiple sclerosis. Another healthy daughter developed a fibroadenoma in 2002. The proband’s wife developed breast cancer too. At follow-up until June 2013, no other family members reported any other disease.
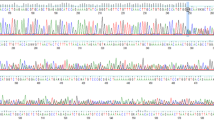
Genetic analysis of BRCA1 and BRCA2 showed a BRCA2 mutation: c.6447_6448dup and different BRCA1-2 polymorphisms as shown in Figure 1. Molecular analysis on all available relatives evidenced that the son, the affected and one of the healthy daughters are carriers of the mutation in the BRCA2 gene.
About the genotype transmission of the high-risk variants, we found that the index case, the wife and all mutated members had the same genotype AG for rs3018301 polymorphism in the NUMA1 gene, demonstrating a possible association of this SNP with an increased risk. Moreover, the intronic variant IVS16-14T>C in BRCA2 gene was ever associated with the variant in BRCA2 gene p.S2411=. The variant p.Q356R in BRCA1 was transmitted from mother to healthy BRCA2 carrier daughter, and in our previously studies9 we showed that this variant could have a possible harmful role. The healthy mutated daughter is also the only one with the genotype TT in the rs4973768 of SLC4A7 gene, suggesting a possible protective role in the development of cancer.
Regarding the polymorphism (rs2981582) in the FGFR2 gene, the affected daughter carried the genotype AA, whereas other members carried AG. Finally, the wife was the only one with the genotype AA in rs3803662 of TNRC9 gene, suggesting a possible association of the genotype AG with the familial breast cancer. All results are reported in Table 1.
This is the first study dealing with the transmission and association of BRCA1/2 genes and their modifiers among members of a BRCA2-mutated family. Thus, this can be considered a preliminary report on a possible association of polygenic variants and an increased risk of breast cancer. Because of the low number of members analyzed, no statistic model could be applied about risk modifier variants. However, this report could be an input for further studies aiming to search high-risk polygenic variants to better understand the genetic background, which could be responsible for familial breast cancer.
Further investigation is also needed to identify other polygenic and monogenic risk modifiers to better contribute to make diagnostic tests much informative.
References
Sawyer, S., Mitchell, G., McKinley, J., Chenevix-Trench, G., Beesley, J., Chen, X.Q. et al. A role for common genomic variants in the assessment of familial breast cancer. J. Clin. Oncol. 30, 4330–4336 (2012).
Pilato, B., Martinucci, M., Danza, K., Pinto, R., Petriella, D., Lacalamita, R. et al. Mutations and polymorphic BRCA variants transmission in breast cancer familial members. Breast Cancer Res. Treat 125, 651–657 (2011).
Fanale, D., Amodeo, V., Corsini, L.R., Rizzo, S., Bazan, V. & Russo, A. Breast cancer genome-wide association studies: there is strength in numbers. Oncogene 31, 2121–2128 (2012).
Turnbull, C., Ahmed, S., Morrison, J., Pernet, D., Renwick, A., Maranian, M. et al. Genome-wide association study identifies five new breast cancer susceptibility loci. Nat. Genet. 42, 504–507 (2010).
van der Merwe, N.C., Hamel, N., Schneider, S.R., Apffelstaedt, J.P., Wijnen, J.T. & Foulkes, W.D. A founder BRCA2 mutation in non-Afrikaner breast cancer patients of the Western Cape of South Africa. Clin. Genet. 81, 179–184 (2012).
Michailidou, K., Hall, P., Gonzalez-Neira, A., Ghoussaini, M., Dennis, J., Milne, R.L. et al. Large-scale genotyping identifies 41 new loci associated with breast cancer risk. Nat. Genet. 45, 353–361 361e351–352 (2013).
Rinella, E.S., Shao, Y., Yackowski, L., Pramanik, S., Oratz, R., Schnabel, F. et al. Genetic variants associated with breast cancer risk for Ashkenazi Jewish women with strong family histories but no identifiable BRCA1/2 mutation. Hum. Genet. 132, 523–536 (2013).
Antoniou, A.C., Kuchenbaecker, K.B., Soucy, P., Beesley, J., Chen, X., McGuffog, L. et al. Common variants at 12p11, 12q24, 9p21, 9q31.2 and in ZNF365 are associated with breast cancer risk for BRCA1 and/or BRCA2 mutation carriers. Breast Cancer Res. 14, R33 (2012).
Tommasi, S., Pilato, B., Pinto, R., Monaco, A., Bruno, M., Campana, M. et al. Molecular and in silico analysis of BRCA1 and BRCA2 variants. Mutat. Res. 644, 64–70 (2008).
Tommasi, S., Crapolicchio, A., Lacalamita, R., Bruno, M., Monaco, A., Petroni, S. et al. BRCA1 mutations and polymorphisms in a hospital-based consecutive series of breast cancer patients from Apulia, Italy. Mutat. Res. 578, 395–405 (2005).
Tommasi, S., Fedele, V., Lacalamita, R., Bruno, M., Schittulli, F., Ginzinger, D. et al. 655Val and 1170Pro ERBB2 SNPs in familial breast cancer risk and BRCA1 alterations. Cell Oncol. 29, 241–248 (2007).
Hasan, S.K., Buttari, F., Ottone, T., Voso, M.T., Hohaus, S., Marasco, E. et al. Risk of acute promyelocytic leukemia in multiple sclerosis: coding variants of DNA repair genes. Neurology 76, 1059–1065 (2011).
Acknowledgements
This study was partially supported by Regione Puglia (DIEF 2007), Bari. We thank Dr Caroline Oakley for language revision.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Pilato, B., De Summa, S., Danza, K. et al. Genetic risk transmission in a family affected by familial breast cancer. J Hum Genet 59, 51–53 (2014). https://doi.org/10.1038/jhg.2013.109
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2013.109
Keywords
This article is cited by
-
Distinct microbiological signatures associated with triple negative breast cancer
Scientific Reports (2015)