Abstract
The presence of genetic rearrangements of bone morphogenetic protein type 2 receptor (BMPR2) was identified in pulmonary arterial hypertension (PAH) patients as the deletion or duplication of one or more exons of the gene. We recently investigated the deletion break points in exonic deletions of BMPR2 in two Japanese familial cases with PAH, and found that these were Alu-mediated via either non-allelic homologous recombination or non-homologous recombination. We herein report the third case of exonic deletion, which was in a 25-year-old female PAH patient with a deletion of BMPR2 exon 3. The break point in this case was not located in an Alu sequence. The 5′- and 3′-break point maps between the inverted Alu sequences in intron 2 and in exon 3, respectively, resulted in a 759-bp deletion. This novel exonic deletion in this PAH case may be a unique and non-recurrent rearrangement, and appears to be of a different size from that in other patients.
Similar content being viewed by others
Introduction
Pulmonary arterial hypertension (PAH) is a frequently lethal disease with a poor prognosis. The linkage analyses and positional cloning have identified the bone morphogenetic protein type 2 receptor (BMPR2) as a causative gene, and it has been reported that BMPR2 mutations account for ∼60% of familial PAH cases and 10–40% of patients with idiopathic PAH.1, 2, 3 Recent studies have clarified the presence of genome-wide structural variations (SVs) caused by genomic rearrangements such as deletion, duplication, inversion and translocation of genetic codes spanning more than 1000 base pairs.4 The presence of genetic rearrangements of BMPR2 was identified in familial PAH patients as the deletion or duplication of one or more exons of the gene.5 We recently undertook a thorough genetic analysis of BMPR2 in Japanese patients with PAH and investigated the deletion break points in exonic deletions of BMPR2 in two familial cases, and found that these were Alu-mediated by either non-allelic homologous recombination or non-homologous recombination.6 We herein describe the third case of exonic deletion, in which the Alu sequence does not appear to be involved in the deletion of an exon.
Materials and methods
Case
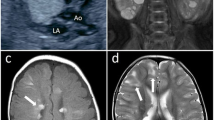
The case was a 25-year-old female PAH patient. No PAH history was reported in her family. She had dyspnea first at 12 years of age and dyspnea worsened gradually. She was diagnosed to have PAH at 18 years of age and estimated pulmonary arterial pressure was 60 mm Hg by echocardiogram and New York Heart Association functional class was II at diagnosis. After diagnosis, medical therapies such as bosentan, sildenafil and beraprost were administered. After 6 years of medical therapy (the timing of genetic analysis), mean pulmonary arterial pressure was 38 mm Hg and pulmonary vascular resistance 8.1 Wood units by right-sided heart catheterization, B-type natriuretic peptide was 14.7 pg ml−1 and New York Heart Association functional class was II. Figure 1 shows the chest X-ray and electrocardiogram. The genetic analysis of this patient was approved by the ethics committee of Kyorin University and written informed consent was obtained.
Genetic analysis
Genetic analyses were carried out as reported previously.7 In brief, DNA was isolated from the patient’s blood, and the protein coding sequence of BMPR2 was amplified by PCR. The amplified fragments were then sequenced directly using the Big Dye Terminator method. A BMPR2 exonic deletion was detected by multiplex ligation-dependent probe amplification (MLPA) using a Salsa MLPA kit for HHT/PPH1 (Cat. No. P093; FALCO Biosystems/MRC-Holland). The break points of the exonic deletion were examined using a combination of semi-quantitative PCR, long PCR and direct sequencing.
Results
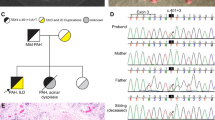
Although there was no point mutation identified in the BMPR2 gene of this patient, the MLPA analysis revealed an exon 3 deletion (Figures 2a and b). The break points had to be located in intron 2 (∼2.5 kb) and intron 3 (46 kb) of the BMPR2 gene. In order to narrow down the break points in intron 3, several semi-quantitative PCRs were carried out. These analyses indicated that the 3′-break point was within a few kb downstream of exon 3. After several long PCR trials using 5′-side primers in intron 2 and 3′-side primers in intron 3, we eventually obtained a 2.6-kb normal fragment and a 1.8-kb deleted fragment (Figure 2c). The long PCR analysis also demonstrated that the patient’s father, who has no symptoms of PAH, has the same deletion (Figure 2c).
These fragments were separately eluted from a gel, subjected to nested PCR and a sequence analysis. The 5′- and 3′-break point maps between the inverted Alu sequences in intron 2 and in exon 3, respectively, resulted in a 759-bp deletion (Figure 3a). However, a unique 9-bp sequence was inserted between the break points. The same 9 bp sequence was also located in 3′-adjacent side of exon 3. It does not appear that the rearrangement involved the Alu sequence, although the 5′-break point was found between the inverted Alu sequences (Figure 3b). Neither a variable number of tandem repeats nor transposable element insertions were involved in this rearrangement.
Discussion
We previously conducted a systematic analysis for larger gene rearrangements, together with a conventional mutation analysis in 152 PAH patients, including 43 patients diagnosed to have idiopathic PAH and 10 with familial PAH, and the analysis of the BMPR2 gene revealed four kinds of nonsense and frame-shift mutations, one missense mutation, one splice-site mutation and two types of exonic deletions.6 The penetrance of heritable PAH was calculated as 22 (2/9) and 44% (4/9) for male and female subjects, respectively, from our data. Our sample numbers are small, but the estimated penetrance is consistent with the data observed in the previous report, that is, ∼20%.8 Our previous study was the first to identify the exact break points related to PAH exonic deletions. In one case, we demonstrated an AluY-mediated non-allelic homologous recombination and a large deletion (>200 kb) involving BMPR2 exons and a neighboring gene. The second case with a medium-sized deletion (5 kb) was not due to non-allelic homologous recombination, but non-homologous recombination was taking place between the Alu-rich regions. Here, this report revealed another patient with a deletion of BMPR2 exon 3, the size of which was <1 kb.
Machado et al.9 and Pfarr et al.10 reported two and one PAH patients with the exon 3 deletions, respectively, but they did not identify the break points. Despite the presence of several highly homologous repetitive Alu sequences located adjacent to exon 3, the break point in this report was not located in such sequences. Therefore, this case may have unique and non-recurrent rearrangements of different sizes from those in other patients (Figure 3c).
Further break point analyses can clarify whether the deletions are mediated by repeated sequences with recombination hotspots.11 Mills et al.12 observed that differences in size spectra of SVs originated from the distinct mechanism of formation. There are several mechanisms that can account for a relatively small deletion: non-homologous recombination associated with the repair of DNA double-strand breaks (that is, non-homologous endjoining) and microhomology-mediated break-induced replication.13 These mechanisms can describe the etiology of non-recurrent copy number variations in human diseases.14
Over the past few years, SV has been shown to be far more common in the human genome than was previously thought. Nearly 2000 SVs have been fine-mapped at the break point level, and most have been identified from healthy donors, such as in the 1000 Genomes Project. The extensive analyses of SV break points may provide an answer to whether the deletion size is correlated to the deletion mechanism, which will be helpful not only for basic science but also for the diagnosis of familial disorders.15, 16, 17
References
Morse, J. H., Jones, A. C., Barst, R. J., Hodge, S. E., Wilhelmsen, K. C. & Nygaard, T. G. Mapping of familial primary pulmonary hypertension locus (PPH1) to chromosome 2q31-q32. Circulation 95, 2603–2606 (1997).
Deng, Z., Morse, J. H., Slager, S. L., Cuervo, N., Moore, K. J., Venetos, G. et al. Familial primary pulmonary hypertension (gene PPH1) is caused by mutations in the bone morphogenetic protein receptor-II gene. Am. J. Hum. Genet. 67, 737–744 (2000).
Elliott, C. G. Genetics of pulmonary arterial hypertension: current and future implications. Semin. Respir. Crit. Care Med. 26, 365–371 (2005).
Kidd, J. M., Graves, T., Newman, T. L., Fulton, R., Hayden, H. S., Malig, M. et al. A human genome structural variation sequencing resource reveals insights into mutational mechanisms. Cell 143, 837–847 (2010).
Cogan, J. D., Pauciulo, M. W., Batchman, A. P., Prince, M. A., Robbins, I. M., Hedges, L. K. et al. High frequency of BMPR2 exonic deletions/duplications in familial pulmonary arterial hypertension. Am. J. Respir. Crit. Care Med. 174, 590–598 (2006).
Kataoka, M., Aimi, Y., Yanagisawa, R., Ono, M., Oka, A., Fukuda, K. et al. Alu-mediated nonallelic homologous and nonhomologous recombination in the BMPR2 gene in heritable pulmonary arterial hypertension. Genet. Med. (e-pub ahead of print 11 April 2013; doi:10.1038/gim.2013.41).
Thomson, J. R., Machado, R. D., Pauciulo, M. W., Morgan, N. V., Humbert, M., Elliott, G. C. et al. Sporadic primary pulmonary hypertension is associated with germline mutations of the gene encoding BMPR-II, a receptor member of the TGF-beta family. J. Med. Genet. 37, 741–745 (2000).
Newman, J. H., Wheeler, L., Lane, K. B., Loyd, E., Gaddipati, R., Phillips, J. A. 3rd et al. Mutation in the gene for bone morphogenetic protein receptor II as a cause of primary pulmonary hypertension in a large kindred. N. Engl. J. Med. 345, 319–324 (2001).
Machado, R. D., Aldred, M. A., James, V., Harrison, R. E., Patel, B., Schwalbe, E. C. et al. Mutations of the TGF-beta type II receptor BMPR2 in pulmonary arterial hypertension. Hum. Mutat. 27, 121–132 (2006).
Pfarr, N., Szamalek-Hoegel, J., Fischer, C., Hinderhofer, K., Nagel, C., Ehlken, N. et al. Hemodynamic and clinical onset in patients with hereditary pulmonary arterial hypertension and BMPR2 mutations. Respir. Res. 12, 99 (2011).
Gu, W., Zhang, F. & Lupski, J. R. Mechanisms for human genomic rearrangements. Pathogenetics 1, 4 (2008).
Mills, R. E., Walter, K., Stewart, C., Handsaker, R. E., Chen, K., Alkan, C. et al. 1000 Genomes Project Mapping copy number variation by population-scale genome sequencing. Nature 470, 59–65 (2011).
Hastings, P. J., Ira, G. & Lupski, J. R. A microhomology-mediated break-induced replication model for the origin of human copy number variation. PLoS Genet. 5, e1000327 (2009).
Verdin, H., D'haene, B., Beysen, D., Novikova, Y., Menten, B., Sante, T. et al. Microhomology-mediated mechanisms underlie non-recurrent disease-causing microdeletions of the FOXL2 gene or its regulatory domain. PLoS Genet. 9, e1003358 (2013).
Boone, P. M., Bacino, C. A., Shaw, C. A., Eng, P. A., Hixson, P. M., Pursley, A. N. et al. Detection of clinically relevant exonic copy-number changes by array CGH. Hum. Mutat. 31, 1326–1342 (2010).
Haraksingh, R. R., Abyzov, A., Gerstein, M., Urban, A. E. & Snyder, M. Genome-wide mapping of copy number variation in humans: comparative analysis of high resolution array platforms. PLoS One 6, e27859 (2011).
Lam, H. Y., Mu, X. J., Stütz, A. M., Tanzer, A., Cayting, P. D., Snyder, M. et al. Nucleotide-resolution analysis of structural variants using BreakSeq and a breakpoint library. Nat. Biotechnol. 28, 47–55 (2010).
Acknowledgements
This work was supported in part by a Health and Labour Sciences Research Grant from the Ministry of Health, Labour and Welfare of Japan, by Grants-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology, by a Grant from the Medical Department Collaborative Project of Kyorin University and by a Fellowship from Kyorin University School of Health Sciences.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflicts of interest.
Rights and permissions
About this article
Cite this article
Aimi, Y., Hirayama, T., Kataoka, M. et al. A novel break point of the BMPR2 gene exonic deletion in a patient with pulmonary arterial hypertension. J Hum Genet 58, 815–818 (2013). https://doi.org/10.1038/jhg.2013.100
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2013.100