Abstract
MicroRNAs (miRNAs) are a class of small, non-coding RNA molecules consisting of 19–22 nucleotides that are involved in a variety of biological processes, including development, differentiation, apoptosis and cell proliferation. In cancer research, a growing body of evidence has indicated that miRNAs are aberrantly expressed in many types of human cancers and can function either as tumor suppressors or oncogenes. Bioinformatic predictions suggest that miRNAs regulate more than 30% of protein-coding genes. Aberrant expression of miRNAs in cancer cells causes destruction of miRNA-regulated messenger RNA networks. Therefore, the identification of miRNA-regulated cancer pathways is important for understanding the molecular mechanisms of human cancer. Searching for the aberrant expression of miRNAs in cancer cells is the first step in the functional analysis of miRNAs in cancer cells. Genome-wide miRNA expression signatures can rapidly and precisely reveal aberrant expression of miRNA in cancers. The miRNA expression signatures of human cancers have revealed that miR-375 is significantly downregulated in cancer cells. Our recent data on maxillary sinus, hypopharyngeal and esophageal squamous cell carcinomas have suggested that miR-375 is frequently downregulated and functions as a tumor suppressor that targets several oncogenic genes in cancer cells. In this review, we focus on several types of human squamous cell carcinoma and describe the aberrant expression of miRNAs and the cancer pathways they regulate in these diseases.
Similar content being viewed by others
Introduction
Head and neck squamous cell carcinoma (HNSCC), the sixth most common malignancy worldwide, arises in the oral cavity, oropharynx, larynx and hypopharynx.1 In spite of considerable advances in multimodal therapy, including surgery, radiotherapy and chemotherapy, the overall 5-year survival rate for patients with HNSCC is only 40–50%.2 Furthermore, the survival rate has not markedly improved in recent decades because of locoregional recurrences, distant metastases and second primary tumors.2 Esophageal cancer is similarly common, being the eighth most common cancer worldwide and ranks sixth among the leading causes of cancer mortality.3 Esophageal cancer can be divided into two main forms with distinct pathological characteristics: squamous cell carcinoma and adenocarcinoma. Esophageal squamous cell carcinoma (ESCC) is the most common type in eastern Asia, including Japan.4 Patients with early stage disease who undergo curative surgery have a 5-year overall survival rate of 30–35%.5 Patients in the advanced stages seldom survive more than 5 years, despite aggressive chemotherapy or chemoradiotherapy.5
To develop evidence-based novel therapies for HNSCC and ESCC, understanding at the molecular level is indispensable. However, most studies on human cancers have focused mainly on protein-coding genes, and our understanding of alterations in non-protein-coding sequences in cancer is largely unclear. In the post-genome sequencing era, it is crucial to find novel molecular mechanisms based on recent genome-wide studies, including non-coding RNA in human oncogenesis, including HNSCC and ESCC.
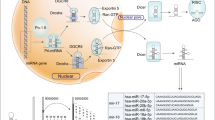
RNA can be divided into two categories: protein-coding RNA and non-coding RNA.6 MicroRNAs (miRNAs) are a class of small, non-coding RNA molecules, consisting of 19–22 nucleotides, that are involved in a variety of biological processes, including development, differentiation, apoptosis and cell proliferation.7 They regulate gene expression through translational repression and messenger RNA cleavage.8 Bioinformatic predictions suggest that miRNAs regulate more than 30% of protein-coding genes.7 So far, 1527 human miRNAs have been registered in miRBase release 18.0 (http://microrna.sanger.ac.uk/).
In the cancer research field, a growing body of evidence suggests that miRNAs are aberrantly expressed in many types of human cancers, including HNSCC and ESCC, and that they have significant roles in initiation, development and metastasis of these cancers.9, 10 Some highly expressed miRNAs could function as oncogenes by repressing tumor suppressors, whereas low-level miRNAs could function as tumor suppressors by negatively regulating oncogenes.10 Aberrant expression of miRNAs in human cancer cells causes destruction of miRNA-regulated messenger RNA networks. Therefore, identification of the miRNA-regulated pathways is important for the next generation of human cancer research.
Genome-wide miRNA expression signatures can rapidly and precisely reveal aberrant expression of miRNA in cancers. Thus, we have conducted miRNA expression signature analyses and searched for tumor-suppressive miRNAs in various types of cancers.11, 12, 13, 14, 15 Our previous studies of HNSCC and ESCC signatures found that miR-375 was significantly reduced in cancer tissues13, 15 and functioned as a tumor suppressor.16, 17 Other research groups also showed downregulation of miR-375 in HNSCC and ESCC and other types of human cancers.18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31
In this review, we focus on HNSCC and ESCC and describe the aberrant expression of miRNAs in these cancers and the cancer pathways they may regulate in these diseases.
Cloning of miR-375 and its function in normal cells
miR-375 was originally cloned and identified as an evolutionarily conserved pancreatic islet-specific miRNA.32 Northern blot analysis showed that miR-375 was highly expressed in mouse pancreatic islet cells, and not found in other tissues, including the liver, lung, fat, intestine, brain, kidney, spleen, heart and testis.32 In zebrafish embryo, miR-375 is expressed in pancreatic islet and pituitary gland.33 So, the functions of miR-375 in normal cells have been investigated mainly in pancreatic islet cells. In murine pancreatic beta cells, overexpression of miR-375 suppressed glucose-induced insulin secretion and, conversely, that inhibition of endogenous miR-375 function enhanced insulin secretion. miR-375 negatively regulated glucose-stimulated insulin secretion by directly regulating the expression of myotrophin, a gene involved in actin depolymerization and, potentially, vesicular fusion.32 miR-375 also negatively regulates the expression of 3′-phosphoinositide-dependent protein kinase 1 (PDK1) in pancreatic beta cells and, subsequently, decreases insulin transcription.34 miR-375−/− mice were found to develop hyperglycemia and hyperglucagonemia, with increased pancreatic alpha cells and decreased pancreatic beta cells.35 These studies show that miR-375 has an important role in insulin secretion and glucose metabolism, and may thereby become a novel therapeutic target in diabetes. miR-375 is also reported to contribute to adipocyte differentiation. Overexpression of miR-375 enhanced 3T3-L1 adipocyte differentiation through modulation of the ERK-PPARγ2-aP2 pathway.36 Functional analysis of miR-375 in normal cells is important, and elucidation of the molecular networks regulated by miR-375 in several types of human cells is necessary.
Aberrant expression and functional significance of miR-375 in HNSCC and ESCC
Screening for differentially expressed miRNAs in cancer cells is important as a first step in understanding the role of miRNAs in cancer. Genome-wide miRNA expression signatures can rapidly and precisely reveal aberrant expression of miRNA in cancer cells. As cases in point, several groups have conducted searches for miRNA expression signatures in HNSCC using PCR-based, microarray-based and sequencing-based methods. In this review, we have listed nine miRNA expression signatures in HNSCC and ESCC.12, 13, 15, 18, 19, 20, 21, 22, 23 Differentially expressed miRNAs are summarized based on the nine signatures in Table 1. Recently, we conducted miRNA expression signature analyses in hypopharyngeal-SCC, maxillary sinus SCC and ESCC using clinical specimens.12, 13, 15 Our expression signatures revealed that miR-1, miR-30a-3p, miR-139-5p, miR-143, miR-204 and miR-375 were commonly downregulated in cancer tissues when compared with adjacent, non-cancerous tissues. These commonly downregulated miRNAs have been selected by other research groups, as well, suggesting that it is important to investigate their functional significance and to identify the molecular targets of these miRNAs to provide novel information on human SCC oncogenesis.
Among the miRNAs identified in our expression signature analysis, miR-375 was found to be the most commonly downregulated, being downregulated in all signatures in HNSCC and ESCC. In spite of the difference in methods of expression analysis, miR-375 was commonly downregulated in those signatures that suggests downregulation of miR-375 is important for SCC oncogenesis. Based on this conclusion, the functional significance of miR-375 in HNSCC was recently investigated by our group. The restoration of miR-375 caused significant inhibition of cell proliferation and induction of cell apoptosis in SAS and FaDu cell lines derived from HNSCC.16 In maxillary sinus SCC, restoration of miR-375 also inhibited cell proliferation and invasion in IMC-3 cells.17 Similarly, another group demonstrated that transfection of miR-375 into FaDu, UTSCC-8 and UTSCC-42a cell lines reduced proliferation and clonogenicity.19
miRNAs are unique in their ability to regulate many protein-coding genes. Bioinformatic predictions suggest that miRNAs regulate more than 30% of protein-coding genes.7 The elucidation of new molecular pathways regulated by tumor-suppressive miR-375 is important for our understanding of human SCC oncogenesis. Based on this view, we also performed molecular target searches for miR-375 in cancer cells by using genome-wide gene expression analysis and luciferase reporter assays in miR-375 transfectants. Our data revealed that the metadherin (MTDH) and lactate dehydrogenase B were direct targets of miR-375 in HNSCC and maxillary sinus SCC.16, 17 Silencing of the MTDH gene caused significant inhibition of cancer cell proliferation, suggesting MTDH functions as an oncogene in HNSCC.16 Very interestingly, another group also reported that MTDH was regulated by miR-375 and that both miR-375 overexpression and MTDH knockdown in HNSCC cell lines resulted in significant decreases in tumor formation in SCID mice.24
In ESCC, expression signatures revealed that miR-375 was significantly reduced in cancer cells.12, 22, 23 The downregulation of miR-375 was caused by hypermethylation of the promoter region, and ectopic expression of miR-375 inhibited tumor cell growth and metastasis in vitro and in vivo by targeting insulin-like growth factor 1 receptor.23 Another study also showed a tumor-suppressive function for miR-375 in esophageal cancer cell lines and identified PDK1 as a target of miR-375.25
The prognosis of both HNSCC and ESCC patients at advanced stages is very poor, despite advances in medical treatments. For this reason, the search for new therapeutic targets is important. Tumor-suppressive miR-375-mediated pathways provide new potential therapeutic targets in HNSCC and ESCC.
Next, we describe the clinical applications of miR-375 in HNSCC and ESCC. Several groups have reported that reduced expression of miR-375 could be a biomarker for HNSCC.18, 21, 37 The expression levels of miR-375 were found to be lower in oral SCC patient oral rinse and saliva compared with those in healthy controls.21 The expression ratio miR-221:miR-375 could be useful for distinguishing HNSCC from normal epithelium.18 Another report showed that low miR-375 expression levels significantly correlated with cancer survival and distant metastasis.37 In ESCC, plasma concentrations of miR-375 were shown to be significantly lower in ESCC patients than in controls.38 Furthermore, low expression levels of miR-375 were significantly correlated with advanced stage, distant metastasis, poor overall survival and disease-free survival in ESCC patients.23
Recent topics of miRNA study are the existence of circulating miRNAs in the blood and body fluids of humans, especially cancer patients. Thus, a number of reports in the past several years have shown that circulating miRNAs in plasma or urine might be used as disease biomarkers and prognostic predictors. These studies will continue to provide evidence for which of the circulating miRNAs may be useful biomarkers in cancer.
Aberrant expression of miR-375 and its target genes in other types of human cancers
Aberrant expression of miR-375 and indications of its involvement in other types of cancer have also been reported (Table 2).16, 17, 23, 24, 25, 26, 27, 28, 29, 30, 31, 39, 40, 41, 42 Downregulation of miR-375 has been reported for gastric cancer, cervical cancer, pancreatic ductal adenocarcinoma and hepatocellular carcinoma.26, 27, 29, 30, 43, 44, 45 In gastric cancer, ectopic expression of miR-375 in cancer cells reduced cell viability and induced apoptosis by targeting PDK1 and tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein ζ (YWHAZ).26 Overexpression of miR-375 suppressed gastric cancer cell proliferation in vitro and in vivo by targeting Janus kinase 2.27 In hepatocellular carcinoma, it was reported that restoration of miR-375 in cancer cells decreased cell proliferation, clonogenicity, migration and invasion, and induced G1 arrest and apoptosis.30 Interestingly, this report demonstrated that MTDH was directly regulated by miR-375, as it is in HNSCC.16, 24, 30 Another report indicated that miR-375 inhibits proliferation and invasion of hepatocellular carcinoma cells by targeting yes-associated protein (YAP).29
We have described miR-375 function as a tumor suppressor in human cancers. However, in some types of cancer, miR-375 functions as an oncogene. In a breast cancer study, miR-375 was upregulated in estrogen receptor alpha-positive cell lines, and miR-375 overexpression, as well as facilitating cell proliferation, induced estrogen receptor alpha upregulation through regulation of RASD1, a negative regulator of estrogen receptor alpha.41 miRNA expression profiling in invasive lobular carcinoma of the breast showed that miR-375 was upregulated when compared with normal breast epithelium.40 Ectopic expression of miR-375 in non-tumorigenic breast epithelial MCF-10 A cells induced loss of cellular organization and a hyperplastic phenotype.40 In prostate cancer (PC), both deep sequencing and microarray analysis of miRNA expression revealed that miR-375 was significantly upregulated in tumor tissues.46 Serum levels of miR-375 in PC patients are upregulated compared with those of healthy controls.47 In addition, miR-375 was significantly upregulated in serum exosome fractions of metastatic PC patients relative to those of patients with non-recurrent PC.48 In lung cancer with neuroendocrine features, the transcription factor ASH1/ASCL1 is a key regulator in this disease.39 This study demonstrated that miR-375 was induced by ASH1/ASCL1 in lung cancer cells. Furthermore, it was indicated that miR-375 directly regulated YAP1, which had growth inhibitory activities in neuroendocrine-lung cancer cells.39 These findings indicate miR-375 can function as an oncogene in some types of cancer.
Is the function of miR-375 an oncogene or a tumor suppressor? This phenomenon of seemingly opposing activities of miR-375 has been described in several articles, yet it is very difficult to explain this phenomenon at present. Individual miRNAs may have different roles and regulate different target genes in different types of cancer, so it is necessary to investigate the expression patterns of both miRNAs and messenger RNAs in specific cancers to understand how miRNAs regulate cancer networks in the future.
In silico analysis of miR-375-regulated pathways
miRNAs are unique in their ability to regulate many protein-coding genes. One miRNA can have hundreds or even thousands of potential gene targets, and bioinformatic predictions suggest that miRNAs regulate more than 30% of protein-coding genes.7 Our interest is in the elucidation of the miRNA-regulated molecular network in several types of cells.
In this review, we have applied in silico analysis to identify the biological processes or pathways potentially regulated by miR-375. Using the TargetScan database, a total of 2267 putative gene targets of miR-375 were identified and are listed (Supplementary Table 1). Among 2267 genes, we investigated differentially expressed genes that were regulated by miR-375 in HNSCC cells. miR-375-regulated genes were identified based on our expression signatures of miR-375 transfectants in HNSCC cancer cells (Supplementary Table 1). These data were registered on the GEO database (accession number: GSE26032). Next, these candidate target genes were assigned to pathways using GENECODIS software analysis,49, 50 and statistically enriched pathways were identified. The GENECODIS software assigned a great many of the putative miR-375 targets to known pathways in KEGG (Kyoto Encyclopedia of Genes and Genomes),51 and these data facilitate the understanding of miRNA-regulated molecular networks in human cells. According to the GENECODIS software analysis, several pathways were identified, such as ‘pathways in cancer’, ‘MAPK signaling pathway’, ‘focal adhesion’ and ‘cytokine-cytokine receptor interaction’ (Figure 1).
Workflow for the in silico analysis of gene targets of miR-375. A total of 2267 genes were identified by the TargetScan program as predicted targets of miR-375. The genes were then analyzed and categorized with KEGG and pathways by the GENECODIS program (left). Twenty significantly enriched signaling pathways are shown in descending order of the number of genes contained in each pathway (right).
We focused on the ‘pathways in cancer’, as it was identified as the most enriched pathway among the miR-375 target genes. Among 2267 genes (these genes have putative target site(s) for miR-375 in their 3′ untranslated regions), 48 were assigned to this pathway (Table 3) and are highlighted in the KEGG MAP (Figure 2). When we analyzed the ‘pathways in cancer’, several genes were found to have been implicated in HNSCC. For example, overexpression of epidermal growth factor receptor has been shown in HNSCC,52, 53 and overexpression of epidermal growth factor receptor is a negative prognostic factor in HNSCC patients.53, 54 The mammalian target of rapamycin (mTOR) pathway was shown to be overactivated in HNSCC by investigating the phosphorylation status of the ribosomal protein S6, a well-known downstream target of mTOR.55 These genes have been used to develop molecularly targeted therapeutic agents for human cancer, such as Gefinitib (Iressa, also known as ZD1839), Erlotinib (marketed as Tarceva) and Cetuximab (marketed as Erbitux), which target epidermal growth factor receptor56, and Everolimus (marketed as Afinitor) and Temsirolimus (marketed as Torisel), which target mTOR.57 Cetuximab received Food and Drug Administration approval for treatment of locally advanced HNSCC in combination with radiotherapy. Overexpression of the genes targeted by these antagonists may result in the reduction of miR-375. In this way, the analysis of molecular pathways that started with identification of cancer-related miRNAs has provided new insights into human oncogenesis.
Conclusions
In this review, we have described aberrant expression of miR-375 in human cancers. In HNSCC and ESCC, miR-375 is frequently downregulated in cancer cells and functions as a tumor suppressor. Identification of molecular targets regulated by miRNAs is a big challenge for current cancer research. The oncogenic and/or tumor-suppressive miRNA-regulated novel cancer pathways could provide new information into the molecular mechanisms of human oncogenesis, and cancer progression and metastasis.
Accession codes
References
Jemal, A., Siegel, R., Xu, J., Ward, E. Cancer statistics. CA Cancer J. Clin. 60, 277–300 (2010).
Leemans, C. R., Braakhuis, B. J., Brakenhoff, R. H. The molecular biology of head and neck cancer. Nat. Rev. Cancer 11, 9–22 (2011).
Enzinger, P. C., Mayer, R. J. Esophageal cancer. N. Engl. J. Med. 349, 2241–2252 (2003).
Akutsu, Y., Matsubara, H. The significance of lymph node status as a prognostic factor for esophageal cancer. Surg. Today 41, 1190–1195 (2011).
Lagergren, J., Lagergren, P. Oesophageal cancer. BMJ 341, c6280 (2010).
Mattick, J. S. RNA regulation: a new genetics? Nat. Rev. Genet. 5, 316–323 (2004).
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Filipowicz, W., Bhattacharyya, S. N., Sonenberg, N. Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat. Rev. Genet. 9, 102–114 (2008).
Tran, N., O'Brien, C. J., Clark, J., Rose, B. Potential role of micro-RNAs in head and neck tumorigenesis. Head Neck 32, 1099–1111 (2010).
Esquela-Kerscher, A., Slack, F. J. Oncomirs - microRNAs with a role in cancer. Nat. Rev. Cancer. 6, 259–269 (2006).
Ichimi, T., Enokida, H., Okuno, Y., Kunimoto, R., Chiyomaru, T., Kawamoto, K. et al. Identification of novel microRNA targets based on microRNA signatures in bladder cancer. Int. J. Cancer 125, 345–352 (2009).
Kano, M., Seki, N., Kikkawa, N., Fujimura, L., Hoshino, I., Akutsu, Y. et al. miR-145, miR-133a and miR-133b: tumor-suppressive miRNAs target FSCN1 in esophageal squamous cell carcinoma. Int. J. Cancer 127, 2804–2814 (2010).
Kikkawa, N., Hanazawa, T., Fujimura, L., Nohata, N., Suzuki, H., Chazono, H. et al. miR-489 is a tumour-suppressive miRNA target PTPN11 in hypopharyngeal squamous cell carcinoma (HSCC). Br. J. Cancer 103, 877–884 (2010).
Moriya, Y., Nohata, N., Kinoshita, T., Mutallip, M., Okamoto, T., Yoshida, S. et al. Tumor suppressive microRNA-133a regulates novel molecular networks in lung squamous cell carcinoma. J. Hum. Genet. 57, 38–45 (2011).
Nohata, N., Hanazawa, T., Kikkawa, N., Sakurai, D., Fujimura, L., Chiyomaru, T. et al. Tumour suppressive microRNA-874 regulates novel cancer networks in maxillary sinus squamous cell carcinoma. Br. J. Cancer 105, 833–841 (2011).
Nohata, N., Hanazawa, T., Kikkawa, N., Mutallip, M., Sakurai, D., Fujimura, L. et al. Tumor suppressive microRNA-375 regulates oncogene AEG-1/MTDH in head and neck squamous cell carcinoma (HNSCC). J. Hum. Genet. 56, 595–601 (2011).
Kinoshita, T., Nohata, N., Yoshino, H., Hanazawa, T., Kikkawa, N., Fujimura, L. et al. Tumor suppressive microRNA-375 regulates lactate dehydrogenase B in maxillary sinus squamous cell carcinoma. Int. J. Oncol. 40, 185–193 (2012).
Avissar, M., Christensen, B. C., Kelsey, K. T., Marsit, C. J. MicroRNA expression ratio is predictive of head and neck squamous cell carcinoma. Clin. Cancer Res. 15, 2850–2855 (2009).
Hui, A. B., Lenarduzzi, M., Krushel, T., Waldron, L., Pintilie, M., Shi, W. et al. Comprehensive microRNA profiling for head and neck squamous cell carcinomas. Clin. Cancer Res. 16, 1129–1139 (2010).
Lajer, C. B., Nielsen, F. C., Friis-Hansen, L., Norrild, B., Borup, R., Garnaes, E. et al. Different miRNA signatures of oral and pharyngeal squamous cell carcinomas: a prospective translational study. Br. J. Cancer 104, 830–840 (2011).
Wiklund, E. D., Gao, S., Hulf, T., Sibbritt, T., Nair, S., Costea, D. E. et al. MicroRNA alterations and associated aberrant DNA methylation patterns across multiple sample types in oral squamous cell carcinoma. PLoS One 6, e27840 (2011).
Mathe, E. A., Nguyen, G. H., Bowman, E. D., Zhao, Y., Budhu, A., Schetter, A. J. et al. MicroRNA expression in squamous cell carcinoma and adenocarcinoma of the esophagus: associations with survival. Clin. Cancer Res. 15, 6192–6200 (2009).
Kong, K. L., Kwong, D. L., Chan, T. H., Law, S. Y., Chen, L., Li, Y. et al. MicroRNA-375 inhibits tumour growth and metastasis in oesophageal squamous cell carcinoma through repressing insulin-like growth factor 1 receptor. Gut 61, 33–42 (2012).
Hui, A. B., Bruce, J. P., Alajez, N. M., Shi, W., Yue, S., Perez-Ordonez, B. et al. Significance of dysregulated metadherin and microRNA-375 in head and neck cancer. Clin. Cancer Res. 17, 7539–7550 (2011).
Li, X., Lin, R., Li, J. Epigenetic silencing of microRNA-375 regulates PDK1 expression in esophageal cancer. Dig. Dis. Sci. 56, 2849–2856 (2011).
Tsukamoto, Y., Nakada, C., Noguchi, T., Tanigawa, M., Nguyen, L. T., Uchida, T. et al. MicroRNA-375 is downregulated in gastric carcinomas and regulates cell survival by targeting PDK1 and 14-3-3zeta. Cancer Res. 70, 2339–2349 (2010).
Ding, L., Xu, Y., Zhang, W., Deng, Y., Si, M., Du, Y. et al. MiR-375 frequently downregulated in gastric cancer inhibits cell proliferation by targeting JAK2. Cell Res. 20, 784–793 (2010).
Basu, A., Alder, H., Khiyami, A., Leahy, P., Croce, C. M., Haldar, S. MicroRNA-375 and microRNA-221: potential noncoding RNAs associated with antiproliferative activity of benzyl isothiocyanate in pancreatic cancer. Genes Cancer 2, 108–119 (2011).
Liu, A. M., Poon, R. T., Luk, J. M. MicroRNA-375 targets hippo-signaling effector YAP in liver cancer and inhibits tumor properties. Biochem. Biophys. Res. Commun. 394, 623–627 (2010).
He, X.-X., Chang, Y., Meng, F.-Y., Wang, M.-Y., Xie, Q.-H., Tang, F. et al. MicroRNA-375 targets AEG-1 in hepatocellular carcinoma and suppresses liver cancer cell growth in vitro and in vivo. Oncogene 31, 3357–3369 (2012).
Mazar, J., DeBlasio, D., Govindarajan, S. S., Zhang, S., Perera, R. J. Epigenetic regulation of microRNA-375 and its role in melanoma development in humans. FEBS Lett 585, 2467–2476 (2011).
Poy, M. N., Eliasson, L., Krutzfeldt, J., Kuwajima, S., Ma, X., Macdonald, P. E. et al. A pancreatic islet-specific microRNA regulates insulin secretion. Nature 432, 226–230 (2004).
Wienholds, E., Kloosterman, W. P., Miska, E., Alvarez-Saavedra, E., Berezikov, E., de Bruijn, E. et al. MicroRNA expression in zebrafish embryonic development. Science 309, 310–311 (2005).
El Ouaamari, A., Baroukh, N., Martens, G. A., Lebrun, P., Pipeleers, D., van Obberghen, E. miR-375 targets 3'-phosphoinositide-dependent protein kinase-1 and regulates glucose-induced biological responses in pancreatic beta-cells. Diabetes 57, 2708–2717 (2008).
Poy, M. N., Hausser, J., Trajkovski, M., Braun, M., Collins, S., Rorsman, P. et al. miR-375 maintains normal pancreatic alpha- and beta-cell mass. Proc. Natl Acad. Sci. USA 106, 5813–5818 (2009).
Ling, H. Y., Wen, G. B., Feng, S. D., Tuo, Q. H., Ou, H. S., Yao, C. H. et al. MicroRNA-375 promotes 3T3-L1 adipocyte differentiation through modulation of extracellular signal-regulated kinase signalling. Clin. Exp. Pharmacol. Physiol. 38, 239–246 (2011).
Harris, T., Jimenez, L., Kawachi, N., Fan, J. B., Chen, J., Belbin, T. et al. Low-level expression of miR-375 correlates with poor outcome and metastasis while altering the invasive properties of head and neck squamous cell carcinomas. Am. J. Pathol. 180, 917–928 (2012).
Komatsu, S., Ichikawa, D., Takeshita, H., Tsujiura, M., Morimura, R., Nagata, H. et al. Circulating microRNAs in plasma of patients with oesophageal squamous cell carcinoma. Br. J. Cancer 105, 104–111 (2011).
Nishikawa, E., Osada, H., Okazaki, Y., Arima, C., Tomida, S., Tatematsu, Y. et al. miR-375 is activated by ASH1 and inhibits YAP1 in a lineage-dependent manner in lung cancer. Cancer Res. 71, 6165–6173 (2011).
Giricz, O., Reynolds, P. A., Ramnauth, A., Liu, C., Wang, T., Stead, L. et al. Hsa-miR-375 is differentially expressed during breast lobular neoplasia and promotes loss of mammary acinar polarity. J. Pathol. 226, 108–119 (2012).
de Souza Rocha Simonini, P., Breiling, A., Gupta, N., Malekpour, M., Youns, M., Omranipour, R. et al. Epigenetically deregulated microRNA-375 is involved in a positive feedback loop with estrogen receptor alpha in breast cancer cells. Cancer Res. 70, 9175–9184 (2010).
Szczyrba, J., Nolte, E., Wach, S., Kremmer, E., Stohr, R., Hartmann, A. et al. Downregulation of Sec23A protein by miRNA-375 in prostate carcinoma. Mol. Cancer Res. 9, 791–800 (2011).
Wang, F., Li, Y., Zhou, J., Xu, J., Peng, C., Ye, F. et al. miR-375 is down-regulated in squamous cervical cancer and inhibits cell migration and invasion via targeting transcription factor SP1. Am. J. Pathol. 179, 2580–2588 (2011).
Bhatti, I., Lee, A., James, V., Hall, R.I., Lund, J.N., Tufarelli, C. et al. Knockdown of microRNA-21 inhibits proliferation and increases cell death by targeting programmed cell death 4 (PDCD4) in pancreatic ductal adenocarcinoma. J. Gastrointest. Surg. 15, 199–208 (2011).
Ladeiro, Y., Couchy, G., Balabaud, C., Bioulac-Sage, P., Pelletier, L., Rebouissou, S. et al. MicroRNA profiling in hepatocellular tumors is associated with clinical features and oncogene/tumor suppressor gene mutations. Hepatology 47, 1955–1963 (2008).
Wach, S., Nolte, E., Szczyrba, J., Stohr, R., Hartmann, A., Orntoft, T. et al. MicroRNA profiles of prostate carcinoma detected by multiplatform microRNA screening. Int. J. Cancer 130, 611–621 (2012).
Selth, L. A., Townley, S., Gillis, J. L., Ochnik, A. M., Murti, K., Macfarlane, R. J. et al. Discovery of circulating microRNAs associated with human prostate cancer using a mouse model of disease. Int. J. Cancer 131, 652–661 (2012).
Bryant, R. J., Pawlowski, T., Catto, J. W., Marsden, G., Vessella, R. L., Rhees, B. et al. Changes in circulating microRNA levels associated with prostate cancer. Br. J. Cancer 106, 768–774 (2012).
Carmona-Saez, P., Chagoyen, M., Tirado, F., Carazo, J. M., Pascual-Montano, A. GENECODIS: a web-based tool for finding significant concurrent annotations in gene lists. Genome Biol. 8, R3 (2007).
Nogales-Cadenas, R., Carmona-Saez, P., Vazquez, M., Vicente, C., Yang, X., Tirado, F. et al. GeneCodis: interpreting gene lists through enrichment analysis and integration of diverse biological information. Nucleic Acids Res. 37, W317–W322 (2009).
Kanehisa, M., Goto, S., Sato, Y., Furumichi, M., Tanabe, M. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res. 40, D109–D114 (2012).
Grandis, J. R., Tweardy, D. J. Elevated levels of transforming growth factor alpha and epidermal growth factor receptor messenger RNA are early markers of carcinogenesis in head and neck cancer. Cancer Res 53, 3579–3584 (1993).
Dassonville, O., Formento, J. L., Francoual, M., Ramaioli, A., Santini, J., Schneider, M. et al. Expression of epidermal growth factor receptor and survival in upper aerodigestive tract cancer. J. Clin. Oncol. 11, 1873–1878 (1993).
Maurizi, M., Almadori, G., Ferrandina, G., Distefano, M., Romanini, M. E., Cadoni, G. et al. Prognostic significance of epidermal growth factor receptor in laryngeal squamous cell carcinoma. Br. J. Cancer 74, 1253–1257 (1996).
Amornphimoltham, P., Patel, V., Sodhi, A., Nikitakis, N. G., Sauk, J. J., Sausville, E. A. et al. Mammalian target of rapamycin, a molecular target in squamous cell carcinomas of the head and neck. Cancer Res. 65, 9953–9961 (2005).
Seshacharyulu, P., Ponnusamy, M. P., Haridas, D., Jain, M., Ganti, A. K., Batra, S. K. Targeting the EGFR signaling pathway in cancer therapy. Expert Opin. Ther. Targets 16, 15–31 (2012).
Freudlsperger, C., Burnett, J. R., Friedman, J. A., Kannabiran, V. R., Chen, Z., Van Waes, C. EGFR-PI3K-AKT-mTOR signaling in head and neck squamous cell carcinomas: attractive targets for molecular-oriented therapy. Expert Opin. Ther. Targets 15, 63–74 (2011).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Kinoshita, T., Hanazawa, T., Nohata, N. et al. The functional significance of microRNA-375 in human squamous cell carcinoma: aberrant expression and effects on cancer pathways. J Hum Genet 57, 556–563 (2012). https://doi.org/10.1038/jhg.2012.75
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2012.75
Keywords
This article is cited by
-
Circulating microRNAs modulating glycolysis as non-invasive prognostic biomarkers of HNSCC
European Archives of Oto-Rhino-Laryngology (2021)
-
The microRNA-375 as a potentially promising biomarker to predict the prognosis of patients with head and neck or esophageal squamous cell carcinoma: a meta-analysis
European Archives of Oto-Rhino-Laryngology (2019)
-
MiR-375 inhibits the hepatocyte growth factor-elicited migration of mesenchymal stem cells by downregulating Akt signaling
Cell and Tissue Research (2018)
-
The microRNA signatures: aberrantly expressed microRNAs in head and neck squamous cell carcinoma
Journal of Human Genetics (2017)
-
MicroRNA expression signature of oral squamous cell carcinoma: functional role of microRNA-26a/b in the modulation of novel cancer pathways
British Journal of Cancer (2015)