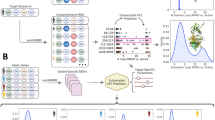
Abstract
Cytochrome c oxidase (COX) of the electron transport system is thought to be the rate-limiting step in cellular respiration and is found mutated in numerous human pathologies. Here, we employ quaternary three-dimensional (3-D) modeling to construct a model for human COX. The model was used to predict the functional consequences of amino-acid mutations based on phylogenetic conservation of amino acids together with volume and/or steric perturbations, participation in subunit–subunit interfaces and non-covalent energy loss or incompatibilities. These metrics were combined and interpreted for potential functional impact. A notable strength of the 3-D model is that it can interpret and predict the structural consequences of amino-acid variation in all 13 protein subunits. Importantly, the influence of compensatory changes can also be modeled. We examine mutations listed in the human mutation database Mitomap, and in 100 older men, and compare the results from the 3-D model against the automated MutPred web application tool. In combination, these comparisons suggest that the 3-D model predicts more functionally significant mutations than does MutPred. We conclude that the model has useful functional prediction capability but may need modification as functional data on specific mutations becomes known.
Similar content being viewed by others
Introduction
Cytochrome c oxidase (COX, complex IV) of the mitochondrial electron transport system catalyzes the reduction of molecular oxygen1 and is hypothesized to be the rate-limiting step in oxidative phosphorylation.2 COX consists of two copper binding sites (CuA and CuB), two hemes (a and a3), as well as a zinc and a magnesium ion. With the transfer of each electron, two protons are pumped through COX from the inner mitochondrial membrane to the intermembrane space. In mammals, COX is composed of 13 subunits consisting of 3 mitochondrial DNA (mtDNA) and 10 nuclear-encoded genes. The three genes expressed from the mtDNA (MT-CO1, MT-CO2 and MT-CO3) are the largest of the subunits and form the catalytic core of the enzyme.3 The 10 nuclear-encoded COX genes form the remainder of the enzyme and are directly regulated by nuclear respiratory factors 1 and 2.4, 5 In mammals, Osada and Akashi6 present evidence for ‘compensatory’ adaptive substitutions in nuclear DNA-encoded mitochondrial proteins to prevent fitness decline in primate COX.6 However, the frequency of such compensatory mutations within humans is currently unknown at least partly because a robust model of human COX has not been developed. A goal of this study is to develop a robust three-dimensional (3-D) model for human mitochondrial COX. We then map known mutations onto the model and determine the frequency of mutations in an aged population that are predicted to be functionally significant.
Mutations in the mtDNA-encoded COX genes have been experimentally associated with COX deficiency in various clinical phenotypes, including motor neuron disease,7 mitochondrial encephalomyopathy,8 mitochondrial myopathy,9, 10 myoglobinuria,11 mitochondrial myopathy, encephalopathy, lactic acidosis and stroke-like (MELAS) episodes,12 and have been associated with aging13 and prostate cancer.14 Mutations in mtDNA-encoded COX genes can affect single tissues or multiple systems with the severity of COX deficiency varying in different tissues.15 Further, the severity of COX deficiency may be associated with heteroplasmic load (the presence of more than one mtDNA type) with mitochondrial dysfunction occurring when one mtDNA type harboring a deleterious mutation reaches a threshold level.16, 17, 18
In concert with experimental studies, bioinformatic tools have been developed to predict the significance of amino-acid variation on protein structure and function.19, 20, 21 Many of these algorithms use phylogenetic information to base their predictions on the effect of amino-acid variation within the context of multiple sequence alignment. 3-D modeling is a further development that can be used to determine the specific interaction of a mutant residue with neighboring residues within individual proteins and protein complexes.20, 22 Thus, bioinformatic protein modeling is a powerful strategy that can be utilized to make informed predictions into the functional importance of protein amino-acid variation. In mammals, the X-ray crystallography-solved structure for COX has been derived from bovine heart tissue23 and can be used to form the foundation for generating a model of the complete 13 subunit human COX structure.
In a previous study, we used the homology modeling approach to develop a 3-D model of COX for the fly Drosophila simulans.22 This approach enabled us to investigate the structural and functional consequences of a naturally occurring deletion in a nuclear-encoded subunit. Here we use a similar approach to develop an improved quaternary 3-D model for human mitochondrial COX using the Modeller 9.10 program and protein template PDB: 3ASO.24, 25 We then compared the 3-D model against the MutPred web application tool.26 MutPred was developed to classify an amino-acid substitution as disease-associated or neutral in humans. First, we modeled COX variants listed in the human mutation database, Mitomap.27 This focused approach allowed us to model and make predictions on known functional COX mutations and to quantify the likelihood of variants altering structural conformation and potentially being functionally important. Next, we sequenced the three mtDNA COX subunits from 100 individuals randomly selected from an aged cohort. In combination, we suggest that the model has the potential to be useful for predicting the functional consequences of COX mutations; however, the model may need modification as functional data on specific mutations becomes known. This study did not include an analysis of any mutations found in the COX assembly genes.
Materials and methods
Quaternary 3-D model
The 3-D model of human COX was generated by homology modeling to bovine COX (PDB: 3ASO, 2.30 Å resolution) using Modeller 9.10.24, 25 The human and bovine (crystallographically resolved) sequence alignment was extracted and the unmatched termini of the human sequence trimmed to match the bovine termini. For example, the first 25 amino acids from COX4I1, the first 45 amino acids from COX5A and the first 31 amino acids from COX6A1 were all trimmed before analysis. This alignment was used to generate three monomeric multisubunit human COX models, including hemes a and a3, copper centers and additional metals resolved in the bovine structure. The high-resolution lipid structures present in bovine COX were not modeled.
The only structural metric that was not optimized was a small fraction of non-standard side-chain rotomers, an outcome of minimization not deemed critical for the qualitative mutant evaluations here.
Structural parameters measured for functional prediction
The qualitative functional impact of each identified mutation was predicted from four unique parameters. The structure-based parameters included sequence conservation derived from the multiple sequence alignment of 16 mammal and four additional vertebrate sequences. This was combined with the three parameters of local volume or steric perturbation, subunit–subunit interaction interface perturbation and non-covalent energy of interaction loss or incompatibility.
Sequence conservation was determined from the multiple sequence alignment of all COX sequences from 20 species (Supplementary Table 1). More distantly related sequences were avoided to reduce the potential for misinterpretation of conservation arising from second site revertants. Sequences were aligned using PROMALS for each of the 13 COX subunits with the inclusion of bovine complete and crystallographically resolved sequences.28 For each mutant position analyzed, the position-specific percent identity was determined across the 20 sequences. Because of the limited number of sequences in the alignments and the high conservation of COX, a stringent criteria for mutant impact was used. A mutant was interpreted as having high potential impact if the respective sequence position was 95–100% conserved (exact residue), while a mild impact was assigned if the conservation was between 75 and 94%. Residues providing <74% conservation were considered to be of little impact.
The internal structural parameters were based on a 5 Å interaction sphere around the site of mutation. This volume represents a conservative distance for all non-covalent and nearby covalent residue contacts. The distance was based on an empirically determined van der Waals contact distance for methane of 4.01 Å.29 Local volume or steric effects were evaluated based on the apparent density of side-chain and/or backbone packing within this sphere, indicated by the number of residues within the interaction distance. The potential impact of additional steric restrains imposed by the backbone trajectory was also interpreted (for example, proline angle restriction). A significant impact was assigned when a side-chain methyl group was removed from a densely packed environment. This type of change would lead to cavity formation and side-chain repacking, which can be energetically costly.30, 31
Subunit–subunit contacts were also determined within the 5 Å radius about each mutation. A mild impact in this case was assigned if only one or a few atoms from the subunit interface were in contact. Typically, contact across subunit interfaces constituted significantly greater numbers of atoms and residues and the impact of a change was considered greater.
Non-covalent energy perturbations were evaluated based on the mutant side-chain environment and its characteristics (for example, hydrophobic, polar, electrostatic). Included in this category were mutant residues having an immediate impact/perturbation on the COX hemes or other metal centers. A significant impact was assigned when an unpaired charge was generated in an otherwise hydrophobic core.
Ranking the functional significance of the mutations
For each parameter, the relative impact of mutation was evaluated and scored as having no impact (0), mild impact (0.5) or substantial impact (1) across the four parameters. The cumulative score was interpreted as the likelihood of any given mutation to affect the function of COX; that is, ‘0+0+0.5+1’ (total score of 1.5) is very unlikely to affect function, while ‘1+1+1+1’ (a maximum score of 4) is very likely to lead to a functional defect. A predictive ratio score between 0 and 1 was then derived from the total cumulative score and the maximum score to determine functionality. In this study, mutations generating normalized ratio scores of 0.6 and above were considered functionally important. However, this score may need to be modified as additional functional data on the significance of specific mutations becomes known.
COX variants listed in the human mutation database
First, we modeled COX variants listed in the human mutation database Mitomap.27 It is important to note that the variants listed in Mitomap do not represent a random sample of the human population. Further, a substantial number of variants are not reported in the literature and represent direct submissions without supporting correlative data.
We then tested whether mutations predicted by the computer program MutPred have any significant effects on function.26 Mutations generating MutPred scores above 0.5 and assigned with AH (actionable hypothesis, may have influence on disease), CH (confident hypothesis, confidence that variant is disease causing) or VCH (very confident hypothesis, high confidence level that variant is disease causing) were considered to be functional.26
Assaying COX variation from an aged cohort
To determine the frequency of mutations predicted to be functional by the 3-D model and by MutPred, we isolated DNA and sequenced the three core COX genes from 100 aged males. Males were randomly selected from the Concord Health and Aging in Men Project (CHAMP) based in Sydney, Australia.32 Aged individuals were chosen because any identifiable COX variant in this study population is unlikely to be highly deleterious given that these individuals are long-lived. Overall, the cohort that we studied was age 77±5.0 years, weight 79.8±12.3 kg and body mass index 28.1±4.1. DNA samples for mutational screening were obtained from peripheral blood leukocytes because somatic mutations are not expected to accumulate with aging in blood.33 The study was approved by the Concord Hospital Human Research Ethics Committee, and all participants gave informed consent.
Total DNA was extracted using standard molecular procedures. A single 4.3-kb DNA fragment encompassing the three mitochondrial genes MT-CO1, MT-CO2 and MT-CO3 was polymerase chain reaction amplified from 100 CHAMP individuals using oligonucleotide primers MT-CO1F (5′-AGGTTTGAAGCTGCTTCTTCG-3′; positions 5771–5791) and MT-CO3R (5′-ATTAGTAGTAAGGCTAGGAGG-3′; positions 10 111–10 091). Primer positions are based on the revised Cambridge Reference Sequence (GenBank accession number NC_012920) and each primer sequence was analyzed by Primer-BLAST (http://www.ncbi.nlm.nih.gov/tools/primer-blast/) to ensure product specificity. A 100 ng of DNA using the Crimson LongAmp Taq DNA Polymerase (New England Biolabs, Ipswich, MA, USA) was amplified using a manual hot start of 98 °C for 2 min, followed by 95 °C for 3 min, 40 cycles of 95 °C for 15 s, 56 °C for 1 min and 65 °C for 3.5 min. For the last 30 cycles, the elongation step at 65 °C was increased by 5 s per cycle. This was followed by a further incubation at 68 °C for 10 min. Polymerase chain reaction fragments were purified using ExoSap (USB Amersham, Buckinghamshire, UK) before sequencing.
To identify mtDNA sequence variation, each 4.3-kb polymerase chain reaction fragment was directly sequenced using nested primers specific for MT-CO1 (Seq1, 5′-TTCAATATGAAAATCACCTC-3′, positions 5801–5820; Seq2, 5′-ACCTAACCATCTTCTCCTT-3′, positions 6334–6352; Seq3, 5′-GATTCATCTTTCTTTTCACC-3′, positions 6931–6950), MT-CO2 (Seq4, 5′-AACCATTTCATAACTTTGTC-3′, positions 7531–7550; Seq5, 5′-AATAATTACATCACAAGACG-3′, positions 8041–8060) and MT-CO3 (Seq6, 5′-GAAATCGCTGTCGCCTTA-3′, positions 9133–9150) to cover each gene region. Polymerase chain reaction templates were denatured at 96 °C for 1 min followed by primer extension in the presence of Big Dye Terminators v.3.1 (Applied Biosystems, Mulgrave, VIC, Australia) for 30 cycles of 96 °C for 15 s, 50 °C for 15 s and 60 °C for 4 min. Trace sequences were analyzed using the Sequencher 5.0 gene analysis software (Gene Codes, Ann Arbor, MI, USA).
Results
Quaternary 3-D model
The top scoring human COX structure was structurally aligned with the original bovine COX (PBD: 3ASO) and sequence identities were extracted from the resulting subunit pairings using UCSF Chimera 1.6 (Supplementary Table 2).34 The initial model also included the hemes and other metal centers. Evaluation of the initial model showed a significant number of structural clashes as determined by van der Waals overlap >0.6 Å. The initial model had 363 structural clashes compared with 27 in the original bovine COX.
To refine the initial structure, we used 200 steps of steepest decent and 100 steps of conjugate gradient minimization, resulting in 26 and 4 clashes, respectively. Structure refinement also improved protein model validity scores. One metric for reliability was the Molprobity score, which was 2.07 for the original bovine structure, 3.16 for the initial model, 2.33 for the steepest decent outcome and 2.05 for the conjugate gradient final model.35 In addition, each of the human COX structures was evaluated with the protein validation tools Molprobity, ERRAT2, PROCHECK and WHATCHECK, and compared with the bovine COX crystal structure.35, 36, 37, 38
Structural parameters measured for functional prediction
The impact of mutation on COX function was inferred from a stringent sequence conservation evaluation and from three unique structural parameters. Only mutations containing several of the structurally measured perturbations were inferred to have a functional impact on human COX.
The sequence identity between 20 species across all 13 subunits ranged from 72% for subunit MT-CO2 to 91% for subunit MT-CO1 (Supplementary Table 2). Although the MT-CO2 subunit sequence identity was only 72%, the sequence similarity based on Blosum62 similarity groups was 87%, indicating that an unambiguous alignment could be achieved for reliable structure generation. In addition, no gaps were present in the alignment of any subunit pairing; thus, no underdetermined unstructured regions were introduced into the human COX homology model. In a large multidomain protein like COX, a conservative change of isoleucine to valine (that is, the loss of a single methyl group) would not be expected to impact negatively on protein function. This statement remains true unless several other factors are involved, such as direct perturbation of a prosthetic group or a local rearrangement leading to extensive repacking and loss of favorable distant interactions.
The structure-based parameters included local volume or steric perturbation, subunit–subunit interaction interface perturbation and non-covalent energy of interaction loss or incompatibility. A qualitative assessment of side-chain atom addition or subtraction was interpreted from local packing density, potential for significant propagation of steric rearrangement (necessary rearrangement of many other amino acids) and the potential impact on hemes and other metal centers. For example, rather subtle and conservative mutations like L196I (in MT-CO1) involved in tight helix–helix packing, V38I (in MT-CO2) making direct contact with a heme prosthetic group (Table 1) and I175V (in MT-CO2) indirectly influencing the CuB ligands (Supplementary Table 3) are all predicted to have a significant impact on human COX function. However, surface-exposed mutations, either facing the aqueous environment or the lipid bilayer, were interpreted to have little or no impact on COX function. The lipid-exposed mutations included V91A and F251L (in MT-CO3) (Table 1) and V193I and I419V (in MT-CO1) (Supplementary Table 3), with each mutation being inferred to be inconsequential to COX function. The unique volume and steric properties of glycine and proline were also considered. For example, glycines in very tight turns when mutated could introduce steric limitations, like G125D (in MT-CO1), or interfere with very close helix–helix packing, like G170S (in MT-CO3) (Table 1). Likewise, steric limitations introduced with proline mutation can influence turn and secondary structure conformations. Proline mutants L135P (in MT-CO2) and S195P (in MT-CO3) are examples and were deemed significant to COX function (Table 1).
Mutations causing a reduction of subunit–subunit interaction energy could presumably reduce COX levels in the mitochondrial membrane. Mutations located in transmembrane helices are predicted to cause the greatest structural perturbations because of changes in packing density and regularity of the backbone structure. Any time a mutation resulted in a significant cross-subunit (subunit–subunit) interaction in the 3-D model, we assumed there would be a significant impact on protein assembly, stability or metal center assembly and environment. For example, the MT-CO1 mutation, F94C (Supplementary Table 3), makes significant hydrophobic contacts with MT-CO3 within the membrane spanning part of this interface. The loss of volume and hydrophobic surface with the mutation could reduce the interaction energy between the two subunits. The MT-CO2 mutation, K98Q (Table 1), makes extensive contact with COX6B1 on the matrix side of the membrane as well as an internal salt bridge that would be broken because of the mutation. Both the loss of a counter charge to a nearby glutamic acid and the change in side-chain volume would be expected to weaken the interaction between MT-CO2 and COX6B1. The MT-CO3 mutation, F35S (Supplementary Table 3), makes extensive hydrophobic contacts with both MT-CO1 and COX6A1 near the membrane/solvent interface. The dramatic change to serine would change the volume greatly as well as the local polarity, both of which would be expected to reduce the stability of the subunit interactions.
Non-covalent interaction perturbations were inferred when polar residues or those having charged side chains occurred in an otherwise hydrophobic environment (for example, I280T in MT-CO1 and M1T or M29K in MT-CO2) or when mutations caused disruption of an interacting polar or charged side chains (for example, S142F in MT-CO1, K98Q in MT-CO2 and S14N or S195P in MT-CO3) (Table 1). A perturbation was also assigned when rearrangement of the local side-chain was inferred to disrupt local hydrogen bonding or electrostatic interactions. However, these were assumed to have only a small impact on COX function (for example, F219L in MT-CO3) (Table 1).
COX variants listed in the human mutation database
Currently, 162 COX amino-acid mutations are listed in Mitomap.27 Of these, 40 have known biochemical functionality (36 are reported with a biochemical defect, while four variants have normal function) and 122 are not biochemically characterized (Figure 1).
Comparisons between Modeller and MutPred for functional mutation prediction. a and b represent the total number of COX mutations with reported biochemical defects analyzed for functional prediction. c and d represent the total number of COX mutations with unknown functional activity analyzed for functional prediction. Light gray bars, the number of mutations predicted to be non-functional; dark gray bars, the number of mutations assigned a confidence level of being functional; and black bars, the number of mutations assigned a very confident level of being functional.
Of the 36 variants with reduced function, 34 are in mtDNA core COX subunits (MT-CO1=15; MT-CO2=8; MT-CO3=11) and two are in nuclear-encoded subunits (COX4I2=1; COX6B1=1). In total, 3-D model analysis of all 36 mutations predicted that 89% (32/36) were likely to be functionally important in a nuclear background that does not harbor compensatory mutations (Table 1). MutPred predictions estimated that 33% (12/36) of the disease-associated mutations with a reported biochemical defect would be functional.
3-D model analysis of the four variants with reported normal activity (A41T, K98Q in MT-CO2, and V91A, F251L in MT-CO3) supported the biochemical data except for the MT-CO2 K98Q variant, which produced a false-positive result (Table 1). MutPred analysis of these same four variants produced false-positive data for K98Q (in MT-CO2) and V91A (in MT-CO3) (Table 1).
A total of 122 non-functionally characterized core COX amino-acid mutations in Mitomap27 were mapped onto the 3-D model (MT-CO1=52; MT-CO2=31; MT-CO3=39). The model predicts that 41% (50/122) core COX variants may be functionally important in a nuclear background without compensatory mutations (Supplementary Table 3). MutPred analysis of the non-functionally characterized core COX mutations estimated that 17% (21/122) would be functional (Supplementary Table 3).
Assaying COX variation from an aged cohort
We sequenced the three core COX genes from 100 aged men to determine the prevalence of mtDNA mutations and estimate the number of functional changes determined by the 3-D model and by MutPred.32 Sequence analysis revealed a total of 69 mtDNA mutations (MT-CO1=38; MT-CO2=17; MT-CO3=14). Of these, 57 (82.6%) were synonymous and 12 (17.4%) were amino-acid variants. From the 12 mutations that caused an amino-acid change, eight are reported in Mitomap.27 Of these eight, three are associated with prostate cancer, while five come from direct submissions and have unknown function (Table 2). The remaining four amino-acid changes are potentially novel.
The 3-D model predictions suggested that four of the 12 mutations (found in four of 100 individuals) were likely to alter the structural integrity of the COX complex and were functionally important (Table 2), assuming that compensatory nuclear mutations have also not occurred. Of the four variants predicted to be functional, two of these, A120T (in MT-CO1) and V136L (in MT-CO3), were predicted to interact with the nuclear-encoded subunits of COX7C and COX6A1, respectively, while the S330G variant (in MT-CO1) was predicted to interact with the mitochondrial-encoded MT-CO2 subunit. The remaining V143M variant (in MT-CO1) was not predicted to interact with any other subunit. MutPred predicted that one (MT-CO1, V193I) of these 12 amino-acid variants would be functional.
Discussion
Here we develop a quaternary 3-D model for human mitochondrial COX. A strength of the model is that it can interpret and predict the structural consequences of amino-acid variation in all 13 protein subunits. Importantly, the influence of compensatory changes can also be modeled. In this study, we compare the results from the 3-D model against the MutPred web application tool.26 However, direct comparisons between the 3-D model and MutPred need to be made with caution as the 3-D model requires an in-depth understanding of bioinformatics modeling. In contrast, MutPred is a fully automated web-based tool that can quickly analyze both structural and functional properties of each protein independently.
First, we investigated mutations reported in Mitomap. The 3-D model predicted that 89% (32/36) of variants previously reported with a measurable biochemical defect would be functionally important. In contrast, MutPred predicted that 33% (12/36) of these same variants would be functionally important. For non-biochemically characterized variants, the 3-D model predicted that 41% (50/122) would influence COX function, while MutPred predicted 17% (21/122) would be functional (Supplementary Table 3). The proportion of mutations that is predicted to be deleterious by both the 3-D model and by MutPred suggests that the data present in Mitomap cannot be used to infer the frequency of deleterious mutations in a normal population. Elliott et al.68 looked for 10 well-known rare inherited mitochondrial diseases, and found that 1/200 harbored them at very low levels of heteroplasmy, while Schaefer et al.69 found that the frequency of mtDNA mutations in disease occurs in approximately 1/4000 individuals.
To test for the frequency of mutations predicted to be functional from within a known population, we isolated DNA and sequenced the three core COX genes from 100 subjects randomly selected from a cohort of 1705 aged individuals enrolled on the Sydney CHAMP study.32 Sequencing of DNA from older men (>70 years) indicates that about 20% (21/100) of the male aged population contain a COX amino-acid mutation. In total, 12 amino-acid variants were found in the aged cohort, with 4 of these predicted to be functional by the 3-D model (Table 2). In contrast, MutPred predicted that one of these 12 amino-acid variants would be functional. In combination with the results obtained from examining the mutations listed in Mitomap, these data show that the 3-D model predicts more functionally significant mutations than does MutPred. At this time, it seems likely that either the 3-D model is overpredicting functional mutations and/or MutPred is underpredicting them. In this study, mutations generating normalized ratio scores of 0.6 and above were considered functionally important. This score may need to be modified as additional functional data on the significance of specific mutations becomes known.
There are several limitations to the current 3-D model. The model is incapable of analyzing some functional properties of protein residues (i.e., DNA-binding residues, phosphorylation and methylation sites). Furthermore, structural analysis on the locations of putative proton/water channels, specific binding surfaces (i.e., the cytochrome c binding surface) or specific lipid binding sites (lipids that could be involved in protein activity) were not measured. In this initial work, we decided not to specify amino acids directly involved in proton translocation through the COX D and K proton channels. These channels have been best described in bacterial COX mutagenesis and crystallographic studies.70 The current model and, in fact, the bovine COX template do not contain the crystallographically resolved water molecules comprising these proton channels. Some of the impact of mutating proton channel residues would be accounted for in the current analysis as energetic perturbations in highly conserved positions. With respect to the cytochrome c binding surface, the exact location(s) of interaction has not yet been reported nor has the impact of individual amino-acid changes. Consideration could be given to clusters of asparate and/or glutamate in future models as cytochrome c itself contains positive amino acids on its surface surrounding its heme. Perhaps, the most difficult functional interaction to characterize is with structurally/functionally important lipids. For example, COX function is known to depend on cardiolipin, but the location of these and other lipids vary or do not exist in current crystallographic structures. This is a likely source of error in predicting mutational impact on the unusually poorly packed MT-CO3, which has significant lipid exposure. In the amino-acid conservation analysis, a fairly stringent metric was used and no consideration was given to similar substitutions (i.e., glutamic acid to aspartic acid). Finally, there is also some potential for double counting importance between volume and energy as they are not completely independent.
This study strongly suggests that the 3-D model will be a useful tool for the functional determination of specific COX mutations. An advantage of the 3-D model is that the loss of function criteria can be adjusted and that insight can be gained about the presence of compensatory mechanisms. Additional sampling of mitochondrial and nuclear-encoded genes in young and older individuals along with the functional testing of the bioenergetic consequences of specific changes is however required to fully assess the robustness of the model and the importance of compensatory mechanisms in humans. Furthermore, this study may form the basis of future studies to generate 3-D models for each of the mitochondrial electron transport system complexes.
Accession codes
References
Shoubridge, E. A. Cytochrome c oxidase deficiency. Am. J. Med. Genet. 106, 46–52 (2001).
Villani, G., Greco, M., Papa, S. & Attardi, G. Low reserve of cytochrome c oxidase capacity in vivo in the respiratory chain of a variety of human cell types. J. Biol. Chem. 273, 31829–31836 (1998).
Barrientos, A., Barros, M. H., Valnot, I., Rotig, A., Rustin, P. & Tzagoloff, A. Cytochrome oxidase in health and disease. Gene 286, 53–63 (2002).
Ongwijitwat, S. & Wong-Riley, M. T. Is nuclear respiratory factor 2 a master transcriptional coordinator for all ten nuclear-encoded cytochrome c oxidase subunits in neurons? Gene 360, 65–77 (2005).
Dhar, S. S., Ongwijitwat, S. & Wong-Riley, M. T. Nuclear respiratory factor 1 regulates all ten nuclear-encoded subunits of cytochrome c oxidase in neurons. J. Biol. Chem. 283, 3120–3129 (2008).
Osada, N. & Akashi, H. Mitochondrial–nuclear interactions and accelerated compensatory evolution: evidence from the primate cytochrome C oxidase complex. Mol. Biol. Evol. 29, 337–346 (2012).
Comi, G. P., Bordoni, A., Salani, S., Franceschina, L., Sciacco, M., Prelle, A. et al. Cytochrome c oxidase subunit I microdeletion in a patient with motor neuron disease. Ann. Neurol. 43, 110–116 (1998).
Clark, K. M., Taylor, R. W., Johnson, M. A., Chinnery, P. F., Chrzanowska-Lightowlers, Z. M., Andrews, R. M. et al. An mtDNA mutation in the initiation codon of the cytochrome C oxidase subunit II gene results in lower levels of the protein and a mitochondrial encephalomyopathy. Am. J. Hum. Genet. 64, 1330–1339 (1999).
Rahman, S., Taanman, J. W., Cooper, J. M., Nelson, I., Hargreaves, I., Meunier, B. et al. A missense mutation of cytochrome oxidase subunit II causes defective assembly and myopathy. Am. J. Hum. Genet. 65, 1030–1039 (1999).
Horvath, R., Schoser, B. G., Muller-Hocker, J., Volpel, M., Jaksch, M. & Lochmuller, H. Mutations in mtDNA-encoded cytochrome c oxidase subunit genes causing isolated myopathy or severe encephalomyopathy. Neuromuscul. Disord. 15, 851–857 (2005).
Hoffbuhr, K. C., Davidson, E., Filiano, B. A., Davidson, M., Kennaway, N. G. & King, M. P. A pathogenic 15-base pair deletion in mitochondrial DNA-encoded cytochrome c oxidase subunit III results in the absence of functional cytochrome c oxidase. J. Biol. Chem. 275, 13994–14003 (2000).
Tam, E. W., Feigenbaum, A., Addis, J. B., Blaser, S., Mackay, N., Al-Dosary, M. et al. A novel mitochondrial DNA mutation in COX1 leads to strokes, seizures, and lactic acidosis. Neuropediatrics 39, 328–334 (2008).
Bratic, I. & Trifunovic, A. Mitochondrial energy metabolism and ageing. Biochim. Biophys. Acta 1797, 961–967 (2010).
Petros, J. A., Baumann, A. K., Ruiz-Pesini, E., Amin, M. B., Sun, C. Q., Hall, J. et al. mtDNA mutations increase tumorigenicity in prostate cancer. Proc. Natl Acad. Sci. USA 102, 719–724 (2005).
Diaz, F. Cytochrome c oxidase deficiency: patients and animal models. Biochim. Biophys. Acta 1802, 100–110 (2010).
Raha, S., Merante, F., Shoubridge, E., Myint, A. T., Tein, I., Benson, L. et al. Repopulation of rho0 cells with mitochondria from a patient with a mitochondrial DNA point mutation in tRNA(Gly) results in respiratory chain dysfunction. Hum. Mutat. 13, 245–254 (1999).
Baranowska, I., Jaderlund, K. H., Nennesmo, I., Holmqvist, E., Heidrich, N., Larsson, N. G. et al. Sensory ataxic neuropathy in golden retriever dogs is caused by a deletion in the mitochondrial tRNATyr gene. PLoS Genet. 5, e1000499 (2009).
Hiona, A., Sanz, A., Kujoth, G. C., Pamplona, R., Seo, A. Y., Hofer, T. et al. Mitochondrial DNA mutations induce mitochondrial dysfunction, apoptosis and sarcopenia in skeletal muscle of mitochondrial DNA mutator mice. PLoS One 5, e11468 (2010).
Dantzer, J., Moad, C., Heiland, R. & Mooney, S. MutDB services: interactive structural analysis of mutation data. Nucleic Acids Res. 33, W311–W314 (2005).
Pinto, J. R., Parvatiyar, M. S., Jones, M. A., Liang, J., Ackerman, M. J. & Potter, J. D. A functional and structural study of troponin C mutations related to hypertrophic cardiomyopathy. J. Biol. Chem. 284, 19090–19100 (2009).
Hicks, S., Wheeler, D. A., Plon, S. E. & Kimmel, M. Prediction of missense mutation functionality depends on both the algorithm and sequence alignment employed. Hum. Mutat. 32, 661–668 (2011).
Melvin, R. G., Katewa, S. D. & Ballard, J. W. O. A candidate complex approach to study functional mitochondrial DNA changes: sequence variation and quaternary structure modeling of Drosophila simulans cytochrome c oxidase. J. Mol. Evol. 66, 232–242 (2008).
Tsukihara, T., Aoyama, H., Yamashita, E., Tomizaki, T., Yamaguchi, H., Shinzawa-Itoh, K. et al. Structures of metal sites of oxidized bovine heart cytochrome c oxidase at 2.8 A. Science 269, 1069–1074 (1995).
Suga, M., Yano, N., Muramoto, K., Shinzawa-Itoh, K., Maeda, T., Yamashita, E. et al. Distinguishing between Cl− and O2(2−) as the bridging element between Fe3+ and Cu2+ in resting-oxidized cytochrome c oxidase. Acta Crystallogr. D 67, 742–744 (2011).
Sali, A. & Blundell, T. L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Li, B., Krishnan, V. G., Mort, M. E., Xin, F., Kamati, K. K., Cooper, D. N. et al. Automated inference of molecular mechanisms of disease from amino acid substitutions. Bioinformatics 25, 2744–2750 (2009).
Ruiz-Pesini, E., Lott, M. T., Procaccio, V., Poole, J. C., Brandon, M. C., Mishmar, D. et al. An enhanced MITOMAP with a global mtDNA mutational phylogeny. Nucleic Acids Res. 35, D823–D828 (2007).
Pei, J. & Grishin, N. V. PROMALS: towards accurate multiple sequence alignments of distantly related proteins. Bioinformatics 23, 802–808 (2007).
Reid, B. P. O. L. M. & Sparks, R. K. Methane–methane isotropic interaction potential from total differential cross sections. J. Chem. Phys. 11, 5656 (1985).
Ratnaparkhi, G. S. & Varadarajan, R. Thermodynamic and structural studies of cavity formation in proteins suggest that loss of packing interactions rather than the hydrophobic effect dominates the observed energetics. Biochemistry 39, 12365–12374 (2000).
Holder, J. B., Bennett, A. F., Chen, J., Spencer, D. S., Byrne, M. P. & Stites, W. E. Energetics of side chain packing in staphylococcal nuclease assessed by exchange of valines, isoleucines, and leucines. Biochemistry 40, 13998–14003 (2001).
Cumming, R. G., Handelsman, D., Seibel, M. J., Creasey, H., Sambrook, P., Waite, L. et al. Cohort Profile: the Concord Health and Ageing in Men Project (CHAMP). Int. J. Epidemiol. 38, 374–378 (2009).
t Hart, L. M., Jansen, J. J., Lemkes, H. H., de Knijff, P. & Maassen, J. A. Heteroplasmy levels of a mitochondrial gene mutation associated with diabetes mellitus decrease in leucocyte DNA upon aging. Hum. Mutat. 7, 193–197 (1996).
Pettersen, E. F., Goddard, T. D., Huang, C. C., Couch, G. S., Greenblatt, D. M., Meng, E. C. et al. UCSF chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Davis, I. W., Leaver-Fay, A., Chen, V. B., Block, J. N., Kapral, G. J., Wang, X. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Colovos, C. & Yeates, T. O. Verification of protein structures: patterns of nonbonded atomic interactions. Protein Sci. 2, 1511–1519 (1993).
Laskowski, R. A., Rullmannn, J. A., MacArthur, M. W., Kaptein, R. & Thornton, J. M. AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J. Biomol. NMR 8, 477–486 (1996).
Hooft, R. W., Vriend, G., Sander, C. & Abola, E. E. Errors in protein structures. Nature 381, 272 (1996).
Uusimaa, J., Finnila, S., Vainionpaa, L., Karppa, M., Herva, R., Rantala, H. et al. A mutation in mitochondrial DNA-encoded cytochrome c oxidase II gene in a child with Alpers–Huttenlocher-like disease. Pediatrics 111, e262–e268 (2003).
Mkaouar-Rebai, E., Ellouze, E., Chamkha, I., Kammoun, F., Triki, C. & Fakhfakh, F. Molecular–clinical correlation in a family with a novel heteroplasmic Leigh syndrome missense mutation in the mitochondrial cytochrome c oxidase III gene. J. Child Neurol. 26, 12–20 (2011).
Tiranti, V., Corona, P., Greco, M., Taanman, J. W., Carrara, F., Lamantea, E. et al. A novel frameshift mutation of the mtDNA COIII gene leads to impaired assembly of cytochrome c oxidase in a patient affected by Leigh-like syndrome. Hum. Mol. Genet. 9, 2733–2742 (2000).
Brown, M. D., Yang, C. C., Trounce, I., Torroni, A., Lott, M. T. & Wallace, D. C. A mitochondrial DNA variant, identified in Leber hereditary optic neuropathy patients, which extends the amino acid sequence of cytochrome c oxidase subunit I. Am. J. Hum. Genet. 51, 378–385 (1992).
Abu-Amero, K. K. & Bosley, T. M. Mitochondrial abnormalities in patients with LHON-like optic neuropathies. Invest. Ophthalmol. Vis. Sci. 47, 4211–4220 (2006).
Manfredi, G., Schon, E. A., Moraes, C. T., Bonilla, E., Berry, G. T., Sladky, J. T. et al. A new mutation associated with MELAS is located in a mitochondrial DNA polypeptide-coding gene. Neuromuscul. Disord. 5, 391–398 (1995).
Varlamov, D. A., Kudin, A. P., Vielhaber, S., Schroder, R., Sassen, R., Becker, A. et al. Metabolic consequences of a novel missense mutation of the mtDNA CO I gene. Hum. Mol. Genet. 11, 1797–1805 (2002).
Hamblet, N. S., Ragland, B., Ali, M., Conyers, B. & Castora, F. J. Mutations in mitochondrial-encoded cytochrome c oxidase subunits I, II, and III genes detected in Alzheimer’s disease using single-strand conformation polymorphism. Electrophoresis 27, 398–408 (2006).
Broker, S., Meunier, B., Rich, P., Gattermann, N. & Hofhaus, G. MtDNA mutations associated with sideroblastic anaemia cause a defect of mitochondrial cytochrome c oxidase. Eur. J. Biochem. 258, 132–138 (1998).
Greaves, L. C., Barron, M. J., Plusa, S., Kirkwood, T. B., Mathers, J. C., Taylor, R. W. et al. Defects in multiple complexes of the respiratory chain are present in ageing human colonic crypts. Exp. Gerontol. 45, 573–579 (2010).
Namslauer, I. & Brzezinski, P. A mitochondrial DNA mutation linked to colon cancer results in proton leaks in cytochrome c oxidase. Proc. Natl Acad. Sci. USA 106, 3402–3407 (2009).
Lucioli, S., Hoffmeier, K., Carrozzo, R., Tessa, A., Ludwig, B. & Santorelli, F. M. Introducing a novel human mtDNA mutation into the Paracoccus denitrificans COX I gene explains functional deficits in a patient. Neurogenetics 7, 51–57 (2006).
Bortot, B., Barbi, E., Biffi, S., Angelini, C., Faleschini, E., Severini, G. M. et al. Two novel cosegregating mutations in tRNAMet and COX III, in a patient with exercise intolerance and autoimmune polyendocrinopathy. Mitochondrion 9, 123–129 (2009).
Herrero-Martin, M. D., Pineda, M., Briones, P., Lopez-Gallardo, E., Carreras, M., Benac, M. et al. A new pathologic mitochondrial DNA mutation in the cytochrome oxidase subunit I (MT-CO1). Hum. Mutat. 29, E112–E122 (2008).
Wei, Y. L., Yu, C. A., Yang, P., Li, A. L., Wen, J. Y., Zhao, S. M. et al. Novel mitochondrial DNA mutations associated with Chinese familial hypertrophic cardiomyopathy. Clin. Exp. Pharmacol. Physiol. 36, 933–939 (2009).
Massa, V., Fernandez-Vizarra, E., Alshahwan, S., Bakhsh, E., Goffrini, P., Ferrero, I. et al. Severe infantile encephalomyopathy caused by a mutation in COX6B1, a nucleus-encoded subunit of cytochrome c oxidase. Am. J. Hum. Genet. 82, 1281–1289 (2008).
Wong, L. J., Dai, P., Tan, D., Lipson, M., Grix, A., Sifry-Platt, M. et al. Severe lactic acidosis caused by a novel frame-shift mutation in mitochondrial-encoded cytochrome c oxidase subunit II. Am. J. Med. Genet. 102, 95–99 (2001).
Valente, L., Piga, D., Lamantea, E., Carrara, F., Uziel, G., Cudia, P. et al. Identification of novel mutations in five patients with mitochondrial encephalomyopathy. Biochim. Biophys. Acta 1787, 491–501 (2009).
Bruno, C., Martinuzzi, A., Tang, Y., Andreu, A. L., Pallotti, F., Bonilla, E. et al. A stop-codon mutation in the human mtDNA cytochrome c oxidase I gene disrupts the functional structure of complex IV. Am. J. Hum. Genet. 65, 611–620 (1999).
Campos, Y., Garcia-Redondo, A., Fernandez-Moreno, M. A., Martinez-Pardo, M., Goda, G., Rubio, J. C. et al. Early-onset multisystem mitochondrial disorder caused by a nonsense mutation in the mitochondrial DNA cytochrome C oxidase II gene. Ann. Neurol. 50, 409–413 (2001).
Hanna, M. G., Nelson, I. P., Rahman, S., Lane, R. J., Land, J., Heales, S. et al. Cytochrome c oxidase deficiency associated with the first stop-codon point mutation in human mtDNA. Am. J. Hum. Genet. 63, 29–36 (1998).
Karadimas, C. L., Greenstein, P., Sue, C. M., Joseph, J. T., Tanji, K., Haller, R. G. et al. Recurrent myoglobinuria due to a nonsense mutation in the COX I gene of mitochondrial DNA. Neurology 55, 644–649 (2000).
Keightley, J. A., Hoffbuhr, K. C., Burton, M. D., Salas, V. M., Johnston, W. S., Penn, A. M. et al. A microdeletion in cytochrome c oxidase (COX) subunit III associated with COX deficiency and recurrent myoglobinuria. Nat. Genet. 12, 410–416 (1996).
Horvath, R., Scharfe, C., Hoeltzenbein, M., Do, B. H., Schroder, C., Warzok, R. et al. Childhood onset mitochondrial myopathy and lactic acidosis caused by a stop mutation in the mitochondrial cytochrome c oxidase III gene. J. Med. Genet. 39, 812–816 (2002).
Kollberg, G., Moslemi, A. R., Lindberg, C., Holme, E. & Oldfors, A. Mitochondrial myopathy and rhabdomyolysis associated with a novel nonsense mutation in the gene encoding cytochrome c oxidase subunit I. J. Neuropathol. Exp. Neurol. 64, 123–128 (2005).
Abu-Amero, K. K., Bosley, T. M. & Morales, J. Analysis of nuclear and mitochondrial genes in patients with pseudoexfoliation glaucoma. Mol. Vis. 14, 29–36 (2008).
Shteyer, E., Saada, A., Shaag, A., Al-Hijawi, F. A., Kidess, R., Revel-Vilk, S. et al. Exocrine pancreatic insufficiency, dyserythropoeitic anemia, and calvarial hyperostosis are caused by a mutation in the COX4I2 gene. Am. J. Hum. Genet. 84, 412–417 (2009).
McFarland, R., Taylor, R. W., Chinnery, P. F., Howell, N. & Turnbull, D. M. A novel sporadic mutation in cytochrome c oxidase subunit II as a cause of rhabdomyolysis. Neuromuscul. Disord. 14, 162–166 (2004).
Marotta, R., Chin, J., Kirby, D. M., Chiotis, M., Cook, M. & Collins, S. J. Novel single base pair COX III subunit deletion of mitochondrial DNA associated with rhabdomyolysis. J. Clin. Neurosci. 18, 290–292 (2011).
Elliott, H. R., Samuels, D. C., Eden, J. A., Relton, C. L. & Chinnery, P. F. Pathogenic mitochondrial DNA mutations are common in the general population. Am. J. Hum. Genet. 83, 254–260 (2008).
Schaefer, A. M., McFarland, R., Blakely, E. L., He, L., Whittaker, R. G., Taylor, R. W. et al. Prevalence of mitochondrial DNA disease in adults. Ann. Neurol. 63, 35–39 (2008).
Yoshikawa, S., Muramoto, K. & Shinzawa-Itoh, K. Proton-pumping mechanism of cytochrome C oxidase. Annu. Rev. Biophys. 40, 205–223 (2011).
Acknowledgements
We thank the investigators involved with the CHAMP study, including Professor Robert Cumming, Associate Professor Fiona Blyth, Associate Professor Vasikaran Naganathan, Professor David Handelsman and Melisa Litchfield for the kind provision of DNA and epidemiological data. We also thank members of the Ballard lab group for comments on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Horan, M., Rumbley, J., Melvin, R. et al. Quaternary protein modeling to predict the function of DNA variation found in human mitochondrial cytochrome c oxidase. J Hum Genet 58, 127–134 (2013). https://doi.org/10.1038/jhg.2012.144
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2012.144