Abstract
Vanishing white matter disease (VWM) is the first human hereditary disease known to be caused by defects in initiation of protein synthesis. Gene defects in each of the five subunits of eukaryotic translation initiation factor 2B (eIF2B α–ɛ) are responsible for the disease, although the mechanism of the pathogenesis is not well understood. In our previous study, four novel eIF2Bɛ mutations were found in Chinese patients: p.Asp62Val, p.Cys335Ser, p.Asn376Asp and p.Ser610-Asp613del. Functional analysis was performed on these mutations and the recently reported p.Arg269X. Our data showed that all resulted in a decrease in the guanine nucleotide exchange (GEF) activity of the eIF2B complex. p.Arg269X and p.Ser610-Asp613del mutants displayed the lowest activity, followed by p.Cys335Ser, p.Asn376Asp and p.Asp62Val. p.Arg269X and p.Ser610-Asp613del could not produce stable eIF2Bɛ, leading to almost complete loss-of-function. No evidence was obtained for the three missense mutations in changes in eIF2Bɛ protein level or eIF2BɛSer540 phosphorylation, and disruption of holocomplex assembly, or binding to eIF2. All patients in our study had the classical phenotype. p.Asp62Val and p.Asn376Asp mutations caused only mildly decreased GEF activity, were probably responsible for relatively mild phenotype in cases of classical VWM.
Similar content being viewed by others
Introduction
Vanishing white matter disease (VWM, OMIM 306896), also called childhood ataxia with central nervous system hypomyelination or eukaryotic translation initiation factor 2B (eIF2B)-related disorder, is one of the most prevalent inherited childhood leukoencephalopathies. The phenotype varies from a severe antenatal form to a mild adult-onset variant. The classical phenotype is characterized by early childhood onset of progressive neurological regression, usually with episodic deterioration provoked by febrile infection, minor head trauma or sudden fright.1, 2, 3 Death occurs after a period, which varies from several years to decades. The remarkable feature in brain magnetic resonance imaging is symmetric and diffuse fluid-like signal in cerebral white matter, which is consistent with the cystic degeneration found by pathological analysis.4
VWM is inherited in an autosomal recessive manner. Mutations in the five subunits of eIF2B (α, β, γ, δ and ɛ), encoded by EIF2B1–5, were identified in VWM in 2001–2002 (Leegwater et al.;5 van der Knaap et al.6). The eIF2B complex has a key role in mRNA translation initiation through its guanine nucleotide exchange (GEF) activity. The eIF2B complex promotes guanosine diphosphate (GDP)/guanosine triphosphate (GTP) exchange on eIF2, and is required to regenerate active GTP–eIF2 from its inactive GDP-bound form to allow continued translation initiation. In the GTP-bound state, eIF2 recruits the initiator Met-tRNA to the 40S subunit for translation initiation. Although the pathogenic pathway from the eIF2B defects to the degeneration of cerebral white matter is far from understood, VWM is considered the first human hereditary disease caused by defects in the initiation of protein synthesis.7, 8
To date, >120 VWM mutations have been identified in EIF2B1–5. More than half were found in EIF2B5 (57%), followed by EIF2B4 (16%), EIF2B2 (16%), EIF2B3 (7%) and EIF2B1 (4%).9, 10, 11 Most of the published studies were carried out in the Caucasian population. Since 2006, 12 Chinese children have been diagnosed with VWM that was confirmed by EIF2B1–5 gene sequencing.12 A total of 16 EIF2B mutations were found (Table 1) in the Chinese patients, with 11 in EIF2B5 (68.8%), 3 in EIF2B3 (18.8%) and 2 in EIF2B2 (12.5%). Eight mutations were not previously reported in other populations, and of these four are in EIF2B5.
Of the five eIF2B subunits, eIF2Bɛ is the largest and the most important, as it exhibits GEF activity when expressed alone. eIF2Bɛ is encoded by EIF2B5. EIF2B5 mutations impair the function of the eIF2B complex in diverse ways, including causing the formation of truncated proteins, disrupting holocomplex assembly and causing partial loss of GEF activity.13, 14, 15 The EIF2B5 mutations that have been functionally investigated are p.Val73Gly, p.Thr91Ala, p.Leu106Phe, p.Arg113His, p.Arg195His, p.Arg299His, p.Arg315His, p.Arg339Pro, p.Gly386Val, p.Val430Ala and p.Trp628Arg.13, 16, 17, 18, 19, 20 The EIF2B5 mutations we found in Chinese patients, which were not previously reported in other population, were p.Asp62Val (c.185A>T), p.Cys335Ser (c.1004G>C), p.Asn376Asp (c.1126A>G) and p.Ser610-Asp613del (c.1827-1838del), they locate from N- through C-terminus of eIF2Bɛ.12 Analysis of how these mutations give rise to the functional defects will help in define the critical sites in eIF2Bɛ involved in its function and regulation. Information about the functional defects that result from specific mutations will be helpful for delineating the genotype–phenotype correlations.
In this study, we analyzed how these EIF2B5 mutations, which recently described in Chinese patients, resulted in functional defects in eIF2Bɛ. Mutants were compared with wild type in terms of the level of eIF2Bɛ protein, interaction of the mutant subunit with the others to form the eIF2B complex, interaction with the eIF2 substrate, the phosphorylation level of eIF2Bɛ at the regulatory GSK3 (glycogen synthase kinase-3) site, and the GEF activity. In addition to the four recently described mutations, p.Arg269X, which was reported in 2008 (Maletkovic et al.11), was also studied. We summarized the phenotypes of four Chinese children who harbored these EIF2B5 mutations, and searched for genotype–phenotype correlations.
Materials and methods
Patients
The research was approved by Medical Ethics Committee of Peking University First Hospital. Clinical diagnosis of VWM was based on the criteria of van der Knaap et al.21 EIF2B1–5 mutation screening was performed for each patient and the parents. Of 12 confirmed VWM patients, EIF2B5 mutations not previously reported in other population were found in four. Data on these patients are in Table 2.
Expression vectors
Expression vectors for full-length wild-type human eIF2Bα–ɛ are as described previously.13 The eIF2Bα–δ complementary DNA in expression vectors was N-terminally tagged with Myc, and the vector with full-length eIF2Bɛ complementary DNA was tagged with both Myc and hexahistidine (His6) at the N-terminus. eIF2Bɛ mutations (p.Asp62Val, p.Arg269X, p.Cys335Ser, p.Asn376Asp and p.Ser610-Asp613del) were generated using the Quick-change site-directed mutagenesis kit (Stratagene, La Jolla, CA, USA) according to the manufacturer's instructions, and were confirmed by sequencing.
Cell culture, transfection, lysis and pull-down purification
Human embryonic kidney 293 (HEK 293) cells were grown in Dulbecco's modified Eagle's medium supplemented with 10% fetal bovine serum. Transfection with the mutants or wild type was conducted using Lipofectamine2000 (Invitrogen, Carlsbad, CA, USA) following the manufacturer's protocol. Cells were lysed 48 h after transfection. To study eIF2Bα–ɛ complex assembly, the interaction with eIF2, phosphorylation level or the GEF activity, recombinant complex containing His-Myc-tagged eIF2Bɛ was purified from HEK 293 lysates containing endogenous eIF2B. Purification was by pull-down with Ni-nitrilotriacetic acid-agarose (Qiagen, Shanghai, China). Pulled-down protein was analyzed by western blot, with anti-Myc to assess the eIF2Bα–ɛ complex formation, anti-eIF2 for eIF2B–eIF2 interaction, and anti-eIF2BɛSer540 (P) for phosphorylation level of eIF2BɛSer540.
Quantitative real-time reverse transcriptase-PCR
To determine whether transcript levels of the mutants resulted in reduced protein, quantitative real-time reverse transcriptase-PCR was performed. Total RNA was isolated from transfected HEK 293 cells with TRIzol reagent (Invitrogen). We used the TaqMan probe technology for analysis. PCR products are located in 5′-upstream of both two mutations, primers are 5′-CTTCTTCCCCATCTCCAAGGA-3′ (sense) and 5′-TACACCTGTGGCAGTCAGGAAT-3′(anti-sense). TaqMan probe for EIF2B5, with 5′-FAM and 3′-TAMRA, is 5′-CAGCCTCGGGTCCTCTTGCCC-3′. Glyceraldehyde 3-phosphate dehydrogenase is used as internal control, primers pairs are 5′-CAGTCAGCCGCATCTTCTTTT-3′(sense) and 5′-GTGACCAGGCGCCCAATAC-3′(anti-sense), TaqMan probe for glyceraldehyde 3-phosphate dehydrogenase is 5′-CGTCGCCAGCCGAGCCACA-3′.
GEF activity measurement
The activity of recombinant eIF2B complex, with His-Myc-tagged eIF2Bɛ, was tested by purification by pull-down as described above. GTP/GDP exchange activity was measured using the standard assay,13 with purified human eIF2.[3H]GDP as substrate. Amounts of purified recombinant eIF2B complex were determined by quantitative western blot using the LiCor Odyssey system, (LI-COR Biotechnology-UK Ltd, Cambridge, UK) and the amounts of protein in GEF activity assays adjusted accordingly.
Results
All five eIF2Bɛ mutations decrease eIF2B GEF activity
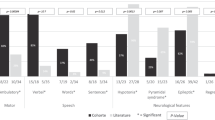
To determine whether the eIF2Bɛ mutations affected GEF activity, we measured this activity using purified recombinant eIF2B heteropentamers containing either mutant or wild-type eIF2Bɛ. All five mutant eIF2Bɛ consistently displayed decreased GEF activity compared with the wild type. The impairment varied from mild to severe. p.Arg269X and p.Ser610-Asp613del mutants displayed the lowest GEF activity, followed by p.Cys335Ser, p.Asn376Asp and p.Asp62Val (Figure 1).
Effects of five EIF2B5 mutations on the eukaryotic translation initiation factor 2B (eIF2B) guanine nucleotide exchange (GEF) activity. GEF activity was measured using purified recombinant eIF2B heteropentamers containing either mutant or wild-type eIF2Bɛ (n=3). All five mutant eIF2Bɛ consistently displayed decreased GEF activity compared with the wild type.
The p.Arg269X and p.Ser610-Asp613del mutations significantly reduce eIF2Bɛ protein level
Vectors containing the five eIF2Bɛ variants were transfected into HEK 293 cells for analysis by in vitro studies. The levels of eIF2Bɛ-Asp62Val, eIF2Bɛ-Cys335Ser and eIF2Bɛ-Asn376Asp were similar to the level of wild-type eIF2Bɛ, while eIF2Bɛ-Arg269X and eIF2Bɛ-Ser610-Asp613del were substantially reduced and were almost undetectable, which shown as very weak bands only when exposure was enhanced (Figure 2). To understand whether the reduction in the mutant protein resulted from a decrease in mRNA level, quantitative real-time reverse transcriptase-PCR was conducted. The level of the mRNA for p.Arg269X was comparable to the wild-type mRNA, while signal for p.Ser610-Asp613del mRNA was not detected suggesting that its stability may be impaired.
Protein expression of five mutant eukaryotic translation initiation factor 2B (eIF2B)ɛ. The five eIF2Bɛ variants were transfected into human embryonic kidney 293 (HEK 293) cells for analysis by in vitro studies (n=5). The levels of eIF2Bɛ-Asp62Val, Cys335Ser and Asn376Asp were similar to the level of wild-type eIF2Bɛ, while eIF2Bɛ-Arg269X and Ser610-Asp613del were substantially reduced and were almost undetectable.
Interaction of eIF2Bɛ with other eIF2B subunits is not affected by the p.Asp62Val, p.Cys335Ser or p.Asn376Asp mutations
eIF2Bɛ alone can mediate GEF, but GEF activity is greatly enhanced when eIF2Bɛ combines with the other four subunits to form a heteropentamer.22, 23 The binary subcomplex of eIF2Bɛ and eIF2Bγ is needed for holocomplex formation. Without the γ subunit, eIF2Bɛ was unable to bind a complex of eIF2Bαβδ. To determine whether the mutations affected the assembly of eIF2B complex, we compared eIF2Bɛγ subcomplex formation between the mutants and the wild type. HEK 293 cells were co-transfected with His-Myc-tagged eIF2Bɛ and Myc-tagged eIF2Bγ. Protein pulled down with Ni- nitrilotriacetic acid-agarose, containing recombinant eIF2Bɛγ, was analyzed by western blot, using anti-Myc. The amount of the pulled-down Myc-tagged eIF2Bγ was comparable between the three missense mutations and the wild type, while no Myc-tagged eIF2Bγ was pulled down from p.Arg269X or p.Ser610-Asp613del transfected cells (Figure 3). Therefore, p.Asp62Val, p.Cys335Ser and p.Asn376Asp did not affect the assembly of the eIF2Bɛγ subcomplex. The failure to detect pulled-down eIF2Bγ with p.Arg269X and p.Ser610-Asp613del probably reflects the greatly reduced amount of protein in the lysate coupled, perhaps, with inability of the truncated polypeptide to bind stably to eIF2Bγ. We further studied whether the mutations affected formation of the eIF2B heteropentamer by expressing the His-Myc-tagged eIF2Bɛ subunit together with all four other Myc-tagged subunits. Similar amounts of Myc-tagged α, β, γ, δ or ɛ subunits were pulled down from cells with the three missense mutations, and the wild-type eIF2Bɛ (Figure 3).
Effects of three EIF2B5 mutations on the Interaction of eukaryotic translation initiation factor 2B (eIF2B)ɛ with other eIF2B subunits. Human embryonic kidney 293 (HEK 293) cells were co-transfected with His-Myc-tagged eIF2Bɛ and Myc-tagged eIF2Bγ. Protein pulled down with Ni-nitrilotriacetic acid (NTA)-agarose, containing recombinant eIF2Bɛγ, was analyzed by western blot, using anti-Myc. p.Asp62Val, p.Cys335Ser and p.Asn376Asp did not affect the assembly of the eIF2Bɛγ subcomplex (Figure 3a). Then we expressed the His-Myc-tagged eIF2Bɛ subunit together with all four other Myc-tagged subunits. Similar amounts of Myc-tagged α, β, γ, δ or ɛ subunits were pulled down from cells with the three missense mutations and the wild type (Figures 3b and c). The failure to detect pulled-down other eIF2B subunits with p.Arg269X and p.Ser610-Asp613del is probably because of the significantly reduced amount of protein in the lysate (n=4).
The p.Asp62Val, p.Cys335Ser and p.Asn376Asp mutations do not affect the binding of eIF2B complex to eIF2
Binding of eIF2B complex to eIF2 (its substrate) is critical for its GEF activity. The eIF2 trimeric complex consists of α, β and γ subunits. Phosphorylation of eIF2α can inhibit the GEF activity of eIF2B, and is an important regulation mechanism for eIF2B.23, 24 We studied whether the binding of eIF2B complex to eIF2 or eIF2α (P) was affected by the three missense mutations. HEK 293 cells were co-transfected with His-Myc-tagged eIF2Bɛ and the other four Myc-tagged eIF2B subunits. Protein pulled down with Ni-nitrilotriacetic acid-agarose, containing recombinant eIF2B, was analyzed by western blot with anti-eIF2 or anti-eIF2α (P). The amount of eIF2 and eIF2α (P), was similar for the three missense mutations and the wild type (Figure 4). Thus, the reduced activity of the missense mutants is not due to impaired binding to substrate or enhanced binding to eIF2α(P).
Effects of three EIF2B5 mutations on the binding of eukaryotic translation initiation factor 2B (eIF2B) complex to eIF2. Human embryonic kidney 293 (HEK 293) cells were co-transfected with His-Myc-tagged eIF2Bɛ and the other four Myc-tagged eIF2B subunits. Protein pulled down with Ni-nitrilotriacetic acid (NTA)-agarose, containing recombinant eIF2B, was analyzed by western blot with anti-eIF2 or anti-eIF2α (P). The amount of eIF2 and eIF2α (P), was similar for the three missense mutations and the wild type (n=3).
The p.Asp62Val, p.Cys335Ser and p.Asn376Asp mutations do not affect the phosphorylation level of eIF2BɛSer540
Phosphorylation of eIF2Bɛ itself is one of the mechanisms that regulates its function, and eIF2BɛSer540, which is the target of GSK3, is a key regulatory phosphorylation site in eIF2Bɛ.23, 24, 25 To determine whether the mutations changed the phosphorylation level of the GSK-3 site, we analyzed purified His-Myc-tagged eIF2Bɛ by western blot, using anti-eIF2BɛSer540 (P) and anti-eIF2Bɛ, and calculated a semi-quantitative eIF2BɛSer540 (P)/eIF2Bɛ ratio for the mutants and the wild type. None of three missense mutations, p.Asp62Val, p.Cys335Ser and p.Asn376Asp, affected the phosphorylation level of eIF2Bɛ Ser540 (Figure 5).
Phosphorylation level of eukaryotic translation initiation factor 2B (eIF2B)ɛSer540 with three missense mutations. We analyzed purified His-Myc-tagged eIF2Bɛ by western blot, using anti-eIF2BɛSer540 (P) and anti-eIF2Bɛ, and calculated a semi-quantitative eIF2BɛSer540 (P)/eIF2Bɛ ratio for the mutants and the wild type. None of three missense mutations, p.Asp62Val, p.Cys335Ser and p.Asn376Asp, affected the phosphorylation level of eIF2Bɛ Ser540 (n=3).
Discussion
eIF2Bɛ is a multidomain protein (721 amino acids) that is widely expressed in eukaryotic cells. It is the largest subunit in the eIF2B heteropentamer complex, and contains the catalytic domain.26 The C-terminal residues (518–712) of eIF2Bɛ contain a minimal functional unit that interacts with eIF2, and exhibits GEF activity. Within the C-terminal region, residues 581–712 can interact with eIF2, and residues 518–580 are required for GEF activity, and are thus considered the ‘catalytic center’. The role of the N-terminal 500 residues of eIF2Bɛ is unclear, but is probably involved in interaction with the other eIF2B subunits.23, 24
Measurement of eIF2B GEF activity in patients’ transformed-lymphocytes was reported to be an important tool for the diagnosis of eIF2B-related disorders.27 We found that all these recently described eIF2Bɛ mutations in Chinese VWM children resulted in a decrease in eIF2B complex GEF activity. The p.Arg269X and p.Ser610-Asp613del mutants displayed the lowest GEF activity, followed by p.Cys335Ser, p.Asn376Asp and p.Asp62Val. Therefore, all the eIF2Bɛ mutations impaired the eIF2Bɛ catalytic function, and consequently affected normal translation initiation.
Our data showed that both p.Arg269X and p.Ser610-Asp613del resulted in significant produce of stable eIF2Bɛ, either because of a decrease in mRNA level or posttranslational degradation. Therefore, these two mutations lead to an almost loss-of-function and should be considered null mutations. The other three mutations did not change the protein level. As all of them are in the N-terminus, it is unlikely that they impair the catalytic function of eIF2Bɛ by interfering directly with GEF activity or substrate binding. On the basis of proposed pathways for regulating eIF2B, we next attempt to define the reason for the decreased GEF activity in the p.Asp62Val, p.Cys335Ser and p.Asn376Asp mutants.
Although eIF2Bɛ itself exhibits GEF activity, the formation of a five-subunit eIF2B complex stimulates GEF activity 10- to 40-fold in yeast or mammalian cells.22 The N-terminus of eIF2Bɛ is considered to be critical for complex formation. The subcomplex eIF2Bɛγ is reported to be required for holocomplex formation. We examined whether the N-terminal mutations interfered with assembly of eIF2Bɛγ or the heteropentamer. All the mutants examined here efficiently formed both types of complex indicating that p.Asp62, p.Cys335 and p.Asn376 in eIF2Bɛ are probably not critical residues for subunit interaction.
Direct phosphorylation of eIF2Bɛ has been described for mammalian eIF2B, which is another way of functional regulation.23, 24, 28 At least four protein kinases have been found to phosphorylate eIF2Bɛ, including CK1 (casein kinase1), CK2 (casein kinase2), DYRK (dual-specificity tyrosine phosphorylated and regulated kinase) and GSK3. Of these, GSK3 is thought to have greatest effect on regulation of eIF2B activity. Phosphorylation of human eIF2Bɛ by GSK3 at Ser540 inhibits eIF2B activity. In the presence of insulin, GSK3 activity is inhibited, thus eIF2Bɛ is dephosphorylated and therefore becomes more active. In our study, the p.Asp62Val, p.Cys335Ser and p.Asn376Asp mutations showed no effects on the phosphorylation of the GSK3 site. Their N-terminal location makes it unlikely that these mutations affect the other known phosphoryraltion sites directly, as most of them are in the C terminus. Indirect effects or changes in other potential phosphorylation sites are possible.
Another mechanism of regulation is phosphorylation of the eIF2α subunit. Phosphorylated eIF2α-Ser51 has a high affinity for eIF2Bɛ, but instead of promoting GEF function, the increased affinity reduces the GTP/GDP exchange function, and thus inhibits eIF2B activity.23 We previously showed that certain mutants in eIF2Bɛ lead to stronger association with eIF2α, perhaps explaining the lower GEF activity of complexes containing these mutants.13 We examined whether these missense mutations affect eIF2Bɛ binding to either total eIF2 or phosphorylated eIF2. Our data showed that the heteropentamer containing mutant eIF2Bɛ had binding similar to the wild type, so these mutations are unlikely abolish regulation of eIF2B by eIF2α.
The phenotype of VWM varies from a severe antenatal form to a mildest adult-onset variant. More than 120 eIF2Bα–ɛ mutations have been found in VWM since 2001 (Maletkovic et al.11). Nonsense mutations are rare and observed only in heterozygotes, associated with a missense mutation allele. The correlation between eIF2B genotype and phenotype is not clear. It is reported that the disease severity is not correlated with either the type of the subunit mutated, or the position of the mutation within the protein.29 Recently p.Arg113His in the ɛ and p.Glu213Gly in the β subunits have been associated with milder phenotypes, and a relationship between severe phenotype with early infantile onset and p.Arg195His in the ɛ has been found.29, 30 One hypothesis is that severe phenotypes occur in the presence of homozygotic mutations in highly conserved regions, with mutations in non-conserved amino acids leading to mild phenotype. However, the genotype–phenotype correlation is not consistent.
All patients in our study had the classical phenotype, with disease onset between 2 and 6 years of age. However, even within this phenotype, patients differed from each other in age of onset, disease progression and clinical manifestations. Horzinski et al.27 found a weak correlation between GEF activity measured in the eIF2B-mutated lymphoblastoid cell line and age at disease onset. Patient 2, with compound heterozygous mutations p.Arg269X/p.Cys335Ser, showed the most severe phenotype. Disease symptoms began as early as 2 years of age, with mild developmental delay even before disease onset. As p.Arg269X is almost functional null, the allele with p.Cys335Ser was the only copy available to produce eIF2Bɛ in this patient. Although as a non-conserved amino acid, eIF2Bɛ-Cys335Ser allele resulted in a significant decrease in eIF2B GEF activity, at only half the activity of wild type, and may not provide enough protection from the severe phenotype. GEF activity was also reported to be significant low (39±7% of control) in a patient's lymphoblast with p.Arg269X heterozygous mutation.27 Patient 1, with an onset at around 3 years of age, harbored the heterozygous genotype p.Asn376Asp/p.Ser447Leu. p.Ser447Leu was reported in a patient with the most severe antenatal form of VWM, who had a compound heterozygous genotype with another missense mutation, thus may have contributed to the severe phenotype.31 eIF2Bɛ-Asp376Asp, which had GEF activity that was approximately 80% of wild type, is probably not related to a severe classical phenotype in the classical form. Patient 3 showed the mildest phenotype, with disease symptoms beginning after 6 years of age, and progressing relatively slowly. This patient harbored the mutations p.Asp62Val/p.Arg339Pro. p.Arg339Pro is reported to lead to a significant reduction in GEF activity to approximately 30% of wild type,13 while we found only slightly reduced activity for p.Asp62Val. Therefore, p.Asp62Val is probably related to the milder phenotype in this patient. Like p.Arg269X, p.Ser610-Asp613del is nearly a functional null, so it is not surprising that this mutation occur in a compound heterozygous manner in patient 4.
In summary, our functional study on eIF2Bɛ mutations recently identified in Chinese VWM patients provides further evidence toward an understanding of the functional defects that result from different mutations. Although we were unable to determine the mechanisms by which the three missense mutations impaired eIF2Bɛ function, we found that they caused at least partial loss of the GEF activity, when part of the eIF2B complex. They seem to impair the intrinsic activity of the eIF2B holocomplex, pointing to interactions between the C-terminal catalytic domain and the more N-terminal regions of eIF2Bɛ in which these mutations occur. Complicated mechanisms, like extensive cross-talk between different domains of eIF2Bɛ and other subunits, and phosphorylation and dephosphorylation of eIF2B and eIF2, appear to be required to regulate eIF2B function. GEF activity of eIF2B may be a preliminary indicator for assessing the severity of functional impairment, and may correlate with phenotype severity in some cases.
References
Schiffmann, R. & van der Knaap, M. S. The latest on leukodystrophies. Curr. Opin. Neurol. 17, 187–192 (2004).
Vermeulen, G., Seidl, R., Mercimek-Mahmutoglu, S., Rotteveel, J. J., Scheper, G. C. & van der Knaap, M. S. Fright is a provoking factor in vanishing white matter disease. Ann. Neurol. 57, 560–563 (2005).
van der Knaap, M. S., Pronk, J. C. & Scheper, G. C. Vanishing white matter disease. Lancet. Neurol. 5, 413–423 (2006).
Leegwater, P. A., Pronk, J. C. & van der Knaap, M. S. Leukoencephalopathy with vanishing white matter: from magnetic resonance imaging pattern to five genes. J. Child. Neurol. 18, 639–645 (2003).
Leegwater, P. A., Vermeulen, G., Könst, A. A., Naidu, S., Mulders, J., Visser, A. et al. Subunits of the translation initiation factor eIF2B are mutant in leukoencephalopathy with vanishing white matter. Nat. Genet. 29, 383–388 (2001).
van der Knaap, M. S., Leegwater, P. A., Könst, A. A., Visser, A., Naidu, S., Oudejans, C. B. et al. Mutations in each of the five subunits of translation initiation factor eIF2B can cause leukoencephalopathy with vanishing white matter. Ann. Neurol. 51, 264–270 (2002).
Scheper, G. C., Proud, C. G. & van der Knaap, M. S. Defective translation initiation causes vanishing of cerebral white matter. Trends Mol. Med. 12, 159–166 (2006).
Scheper, G. C., van der Knaap, M. S. & Proud, C. G. Translation matters: protein synthesis defects in inherited disease. Nat. Rev. Genet. 8, 711–723 (2007).
Scali, O., Di Perri, C. & Federico, A. The spectrum of mutations for the diagnosis of vanishing white matter disease. Neurol. Sci. 27, 271–277 (2006).
Pronk, J. C., van Kollenburg, B., Scheper, G. C. & van der Knaap, M. S. Vanishing white matter disease: a review with focus on its genetics. Ment. Retard. Dev. Disabil. Res. Rev. 12, 123–128 (2006).
Maletkovic, J., Schiffmann, R., Gorospe, J. R., Gordon, E. S., Mintz, M., Hoffman, E. P. et al. Genetic and clinical heterogeneity in eIF2B-related disorder. J. Child. Neurol. 23, 205–215 (2008).
Wu, Y., Pan, Y., Du, L., Wang, J., Gu, Q., Gao, Z. et al. Identification of novel EIF2B mutations in Chinese patients with vanishing white matter disease. J. Hum. Genet. 54, 74–77 (2009).
Li, W., Wang, X., van der Knaap, M. S. & Proud, C. G. Mutations linked to leukoencephalopathy with vanishing white matter impair the function of the eukaryotic initiation factor 2B complex in diverse ways. Mol. Cell. Biol. 24, 3295–3306 (2004).
Hiyama, T. B., Ito, T., Imataka, H. & Yokoyama, S. Crystal structure of the alpha subunit of human translation initiation factor 2B. J. Mol. Biol. 392, 937–951 (2009).
Pavitt, G. D. & Proud, C. G. Protein synthesis and its control in neuronal cells with a focus on vanishing white matter disease. Biochem. Soc. Trans. 37 (Part 6), 1298–1310 (2009).
Richardson, J. P., Mohammad, S. S. & Pavitt, G. D. Mutations causing childhood ataxia with central nervous system hypomyelination reduce eukaryotic initiation factor 2B complex formation and activity. Mol. Cell. Bio. 24, 2352–2363 (2004).
van Kollenburg, B., Thomas, A. A., Vermeulen, G., Bertrand, G. A., van Berkel, C. G., Pronk, J. C. et al. Regulation of protein synthesis in lymphoblasts from vanishing white matter patients. Neurobiol. Dis. 21, 496–504 (2006).
Kantor, L., Harding, H. P., Ron, D., Schiffmann, R., Kaneski, C. R., Kimball, S. R. et al. Heightened stress response in primary fibroblasts expressing mutant eIF2B genes from CACH/VWM leukodystrophy patients. Hum. Genet. 118, 99–106 (2005).
van Kollenburg, B., van Dijk, J., Garbern, J., Thomas, A. A., Scheper, G. C., Powers, J. M. et al. Glia-specific activation of all pathways of the unfolded protein response in vanishing white matter disease. J. Neuropathol. Exp. Neurol. 65, 707–715 (2006).
Kantor, L., Pinchasi, D., Mintz, M., Hathout, Y., Vanderver, A. & ElroyStein, O. A point mutation in translation initiation factor 2B leads to a continuous hyper stress state in oligodendroglial-derived cells. PLoS. One. 3, e3783 (2008).
van der Knaap, M. S., Barth, P. G., Gabreëls, F. J., Franzoni, E., Begeer, J. H., Stroink, H. et al. A new leukoencephalopathy with vanishing white matter. Neurology 48, 845–855 (1997).
Fabian, J. R., Kimball, S. R., Heinzinger, N. K. & Jefferson, L. S. Subunit assembly and guanine nucleotide exchange activity of eukaryotic initiation factor-2B expressed in Sf9 cells. J. Biol. Chem. 272, 12359–12365 (1997).
Pavitt, G. D. eIF2B, a mediator of general and gene-specific translational control. Biochem. Soc. Trans. 33, 1487–1492 (2005).
Kubica, N., Jefferson, L. S. & Kimball, S. R. Eukaryotic initiation factor 2B and its role in alterations in mRNA translation that occur under a number of pathophysiological and physiological conditions. Prog. Nucleic. Acid. Res. Mol. Biol. 81, 271–296 (2006).
Welsh, G. I., Miller, C. M., Loughlin, A. J., Price, N. T. & Proud, C. G. Regulation of eukaryotic initiation factor eIF2B: glycogen synthase kinase-3 phosphorylates a conserved serine which undergoes dephosphorylation in response to insulin. FEBS Lett. 421, 125–130 (1998).
Gomez, E. & Pavitt, G. D. Identification of domains and residues within the epsilon subunit of eukaryotic translation initiation factor 2B (eIF2Bepsilon) required for guanine nucleotide exchange reveals a novel activation function promoted by eIF2B complex formation. Mol. Cell. Biol. 20, 3965–3976 (2000).
Horzinski, L., Huyghe, A., Cardoso, M. C., Gonthier, C., Ouchchane, L., Schiffmann, R. et al. Eukaryotic initiation factor 2B (eIF2B) GEF activity as a diagnostic tool for EIF2B-related disorders. PLoS. One. 4, e8318 (2009).
Wang, X. & Proud, C. G. A novel mechanism for the control of translation initiation by amino acids, mediated by phosphorylation of eukaryotic initiation factor 2B. Mol. Cell. Biol. 28, 1429–1442 (2008).
Fogli, A., Schiffmann, R., Bertini, E., Ughetto, S., Combes, P., Eymard-Pierre, E. et al. The effect of genotype on the natural history of eIF2B-related leukodystrophies. Neurology 62, 1509–1517 (2004).
Fogli, A. & Boespflug-Tanguy, O. The large spectrum of eIF2B-related diseases. Biochem. Soc. Trans. 34 (Part 1), 22–29 (2006).
van der Knaap, M. S., van Berkel, C. G., Herms, J., van Coster, R., Baethmann, M., Naidu, S. et al. eIF2B-related disorders: antenatal onset and involvement of multiple organs. Am. J. Hum. Genet. 73, 1199–1207 (2003).
Acknowledgements
We are grateful to the patients’ families. This work was funded by National Key Research Project ‘11–5’ (2006BAI05A07), National Key Research Project ‘973’, (2007CB5119004), Natural Science Foundation of China (30772355, 30872793), Natural Science Foundation of Beijing (7082093), and program for New Century Excellent Talents in University.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Leng, X., Wu, Y., Wang, X. et al. Functional analysis of recently identified mutations in eukaryotic translation initiation factor 2Bɛ (eIF2Bɛ) identified in Chinese patients with vanishing white matter disease. J Hum Genet 56, 300–305 (2011). https://doi.org/10.1038/jhg.2011.9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2011.9