Abstract
A growing list of membrane-spanning proteins involved in the transport of a large variety of drugs has been recognized and characterized to include peptide and organic anion/cation transporters. Given such an important role of transporter genes in drug disposition process, the role of single-nucleotide polymorphisms (SNPs) in such transporters as potential determinants of interindividual variability in drug disposition and pharmacological response has been investigated. To define the distribution of transporter gene SNPs across ethnic groups, we screened 450 DNAs in cohorts of 250 Korean, 50 Han Chinese, 50 Japanese, 50 African-American and 50 European-American ancestries for 64 SNPs in four transporter genes encoding proteins of the solute carrier family (SLC15A2, SLC22A1, SLC22A2 and SLC22A6). Of the 64 SNPs, 19 were core pharmacogenetic variants and 45 were HapMap tagging SNPs. Polymorphisms were genotyped using the golden gate genotyping assay. After genetic variability, haplotype structures and ethnic diversity were analyzed, we observed that the distributions of SNPs in a Korean population were similar to other Asian groups (Chinese and Japanese), and significantly different from African-American and European-American cohorts. Findings from this study would be valuable for further researches, including pharmacogenetic studies for drug responses.
Similar content being viewed by others
Introduction
Membrane transporters have a critical role in a variety of physiological processes. They maintain cellular and organismal homeostasis by importing nutrients essential for cellular metabolism and exporting cellular waste products and toxic compounds.1 Furthermore, membrane transporters are important in drug response as they provide attractive targets for many commonly used drugs and are major determinants of drug absorption, distribution and elimination.2 Membrane transport proteins share a similar secondary structure characterized by multiple membrane-spanning domains joined by alternating intracellular and extracellular segments.3 Two of the major superfamilies of membrane transport proteins are the ATP-binding cassette transporters and the solute carrier (SLC) transporters, both of which take up neurotransmitters, nutrients, heavy metals and other substrates into cells.4, 5
In this study, we screened for genetic polymorphisms in four SLC transporters, SLC15A2, SLC22A1, SLC22A2 and SLC22A6 as part of a pharmacogenetics project that estimates allele frequency, linkage disequilibrium and haplotype structure of pharmacogenes in a Korean population.
Located on 3q13.33, SLC15A2 is a transporter for peptide-like drugs such as penicillins, cephalosporins and glycyl-sarcosine.6, 7 Genetic variations in SLC15A2 (F350L, S409P and K509R) are associated with varying Km values for glycyl-sarcosine.8
Located on 6q26, organic cation transporters, SLC22A1 and SLC22A2, are members of the SLC22 family. The uptake of metformin in the hepatocytes by SLC22A1 is an essential step for the inhibition of hepatic gluconeogenesis and the glucose-lowering effect of metformin.9 Genetic variations in SLC22A1 (R61C, G401S, M420del, G465R and rs622342) are associated with differences in metformin blood levels, glucose levels after an oral glucose tolerance test and glucose-lowering effect of metformin.2, 9, 10 Among SLC22A2 variants, A270S is associated with differences in metformin blood levels and renal excretion, and R400C and K432Q have significant roles in altered activity of SLC22A2.11, 12
Located on 11q13.1-q13.2, organic anion transporter, SLC22A6, has been shown to transport methotrexate (cytotoxic antimetabolite used for rheumatoid arthritis, psoriasis and cancer of various types) and to be inhibited by non-steroidal anti-inflammatory drugs.13, 14 The R50H variant in SLC22A6 is associated with kinetic differences for the nucleoside phosphonate analogs adefovir, cidofovir and tenofovir.15
Because the genetic variants associated with impaired transport function may have an influence on substrate disposition, it is of importance to identify the polymorphisms and ethnic diversity of the four SLC genes. However, little information is available regarding genetic polymorphisms of the SLC genes in a Korean population or on the ethnic differences among Asian groups regarding genetic polymorphisms related to allele frequencies and haplotype blocks.
To define the distribution of polymorphisms in these transporter genes across ethnic groups, we screened an ethnically diverse collection of 450 genomic DNA samples in cohorts of 250 Korean, 50 Chinese, 50 Japanese, 50 African-American and 50 European-American ancestries for 64 single-nucleotide polymorphisms (SNPs) in SLC15A2, SLC22A1, SLC22A2 and SLC22A6. Of the 64 SNPs, 19 were core pharmacogenetic variants and 45 were HapMap tagging SNPs. Genotyping variants in this DNA collection allowed us to determine the levels and patterns of genetic diversity in different ethnic groups.
Materials and methods
Study subjects
DNAs from 250 unrelated Korean ethnic group individuals were graciously provided by the Center for Genome Science, Korea Centers for Disease Control and Prevention. Other ethnic groups were obtained from a large panel of anonymous, unrelated DNA samples from the Human Variation Panels available from the Coriell Institute (NJ, USA). We specifically used sets of DNA samples obtained from four distinct ethnic groups residing in the USA, including 50 Han Chinese, 50 Japanese, 50 African-American and 50 European-American which is sufficient sample size for the screening of the ethnic diversity.16
SNP selection and genotyping
We selected four SLC transporter genes (SLC15A2, SLC22A1, SLC22A2 and SLC22A6) and their pharmacogenetic core SNPs from PharmGKB (http://www.pharmgkb.org) and PharmaADME (http://www.pharmaadme.org) databases. To examine the whole gene region, we then searched the HapMap database (http://www.hapmap.org) for tagging SNPs (r2 threshold of 0.98) with a minor allele frequency of over 5% in Asian groups (Han Chinese and Japanese).
Genotyping was performed at a multiplex level of using the Golden Gate genotyping system (Illumina Inc., San Diego, CA, USA).17 The genotype quality score for retaining data was set to 0.25. SNPs that could not satisfy the following criteria were excluded: a minimum call rate of 98% and no duplicate error. A total of 64 SNPs were successfully genotyped. Four core SNPs (SLC22A1-420del,9 SLC22A2-T199I and T201M,18 SLC22A6-R454Q19) failed to genotype on the Illumina platform and, therefore, were not included in further analysis.
Statistical analysis
The χ2-tests were used to determine whether individual variants were in equilibrium at each locus in each population (Hardy–Weinberg equilibrium). Pairwise linkage disequilibrium was estimated for the SNPs in each gene using standardized summary statistics20 D′ and r2, which were calculated using HaploView software.21 Haplotype blocks were assigned using the D′ confidence interval algorithm from Gabriel et al., which is implemented in HaploView.21 Fisher's Exact tests for allele-frequency differences in SNPs were conducted in the HelixTree software (Golden Helix Inc., Bozeman, MT, USA).
Results and discussion
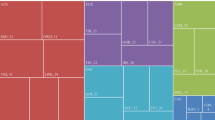
The general haplotype structure of each gene is shown in Figure 1 by ethnicity. Detailed results of all statistical analyses and gene maps for each gene and ethnicity are shown in Figure 2 and Supplementary Data. The allele frequencies and haplotype structures in SLC15A2, SLC22A1 and SLC22A6 were similar across all Asian populations (Korean, Han Chinese and Japanese). Although allele frequencies variants in SLC22A2 were also similar across Asians, haplotype structures were different in each Asian group. Four haplotype blocks were common to three Asian populations and thirteen blocks were ethnic specific. All SNPs were in Hardy–Weinberg equilibrium (P>0.01). Fisher's exact tests revealed statistically significant differences in allele frequencies from African-American and European-American populations compared with Koreans (Figure 2).
Haplotype structure overview of four SLC transporter genes by population. Columns indicate the gene and rows indicate ethnicity. AA, Africa-American; EA, European-American; HC, Han Chinese; JP, Japanese; KR, Korean. A full color version of this figure is available at the Journal of Human Genetics journal online.
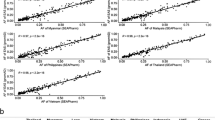
Haplotype blocks, allele frequencies, −log P-values of Hardy–Weinberg equilibrium tests and −log P-values by frequency difference among Korean, Han Chinese, Japanese, African-American and European-American samples; SLC15A2 (a), SLC22A1 (b), SLC22A2 (c) and SLC22A6 (d). Relative physical positions of SNPs are shown at the top of each gene. Core SNPs and identified haplotype blocks are indicated below SNP IDs. SNP statistics, such as minor allele frequency, −log P-values of Hardy–Weinberg equilibrium tests and −log P-values of Fisher's exact tests are shown as line graphs. AA, Africa-American; EA, European-American; HC, Han Chinese; JP, Japanese; KR, Korean.
The haplotype block of SLC15A2 was most common to all ethnicities. Notably, the SLC15A2 haplotype block in a Japanese cohort was more similar to African-American and European-American populations (Figures 1 and 2a). All four pharmacogenetic core SNPs (rs2293616 [A284A], rs2257212 [L350F], rs1143671 [P409S] and rs1143672 [R509K])8 were in absolute linkage disequilibrium (r2=1) (Supplementary Figure 1a). SLC15A2 has been shown to recognize a wide spectrum of pharmacologically active compounds, including β-lactam antibiotics,22 inhibitors of the angiotensin converting enzyme23, 24 and non-peptidic drugs.24 Genetic variations in SLC15A2 (F350L, S409P and K509R) displayed different Km values for glycyl-sarcosine.8 Therefore, individuals with these genetic variations may have altered drug response.
For the SLC22A1 gene, two well-defined haplotype blocks ranging from rs9457840 to 3798167 (26 Kb) and from rs715083 to rs609468 (5 Kb) were inferred from the Asian populations. In contrast, the haplotype structures of African-American and European-American populations were distinctive with fewer and smaller blocks than the Asian ethnicities. An intronic core SNP, rs622342 which has been known to be associated with metformin response in patients with diabetes mellitus10, 25 was not included in the common haplotype blocks in all ethnicities (Figures 1 and 2b). Among nine core SNPs in SLC22A1, rs55918055 (C88R26), rs36103319 (G220V27) and rs4646278 (R287G28) were monomorphic in all ethnicities, and rs12208357 (R61C) and rs34059508 (G465R)2, 9, 10 were monomorphic only in Asians (Figure 2b; Supplementary Figure 1b; Supplementary Table 1). SLC22A2 showed the most complicated haplotype structures in all five populations. There were eight distinct haplotype blocks, none of which was common to each ethnicity. One clearly defined haplotype block in the whole SLC22A2 gene region was observed in European-Americans (Figures 1 and 2c). Among five core SNPs in SLC22A2, rs8177504 (P54S), rs8177507 (M165I), rs8177516 (R400C) and rs8177517 (K432Q)12 were monomorphic in Asian and European-American populations. One polymorphic core SNP, rs316019 (S270A) showed similar allele frequencies in all ethnicities (Figure 2c; Supplementary Figure 1c; Supplementary Table 1). Functional polymorphisms in SLC22A1 and SLC22A2 can influence the clinical effects and pharmacokinetics of metformin, a drug used as a primary therapy for type 2 diabetes mellitus. Homozygous carriers of the low activity SLC22A2 variant, 270S, were reported to have significantly lower rates of renal clearance and higher plasma concentration of metformin than those homozygous for the active variant, 270A.11, 29 On the other hand, low-function SLC22A1 variants including R61C, G401S and G465R have been associated with significantly higher rates of renal clearance of metformin.30 In addition to the effect on metformin pharmacokinetics, low-function SLC22A1 variants have been also associated with significantly decreased glucose-lowering response of metformin.2
For the SLC22A6 gene, one haplotype block ranging from rs4149170 to rs4149172 was obtained in Asian populations, and none was defined in African-American and European-American cohorts (Figures 1 and 2d). One core SNP, rs11568626 (R50H15), was monomorphic in Asian and European-American populations (Figure 2d; Supplementary Figure 1d; Supplementary Table 1). The Km for the nucleoside phosphonate analogs adefovir, cidofovir and tenofovir is decreased in the R50H variant compared with the wild-type.15 Since SLC22A6 has been suggested as the basis of nephrotoxicity of the nucleoside phosphonate analogs,31 individuals carrying variations in the gene such as R50H may respond to these drugs differently.
In pharmacogenetics, screening of known genetic variants between ethnicities is worthwhile for bridging studies and personalized medicine. The increasing pharmacogenetic knowledge such as the discovery of rare mutations that induce unusual enzyme activity and common variants that modify drug response phenotypes invited a more thorough characterization of ethnic differences in allele frequency and haplotype structure across four genes in the SLC family (SLC15A2, SLC22A1, SLC22A2 and SLC22A6). Although a complete characterization of all haplotype structures and allele frequencies in more advanced studies involving many individuals are required, the results of this study provide a basic working understanding of population differences in these four genes that is necessary for further pharmacogenetic studies. In addition, detailed analyses of core SNPs in SLC genes provide a set of haplotype tagging SNPs that economically represent drug responses. For example, one tag SNP may reflect the enzyme activity of SLC15A2 because all four core SNPs, A284A, L350F, P409S and R509K were in absolute linkage disequilibrium (r2=1).
Although we failed to perform functional characterization of the analyzed variants, our results suggest that ethnic differences significantly contribute to individuals' drug responses in relation to the SLC family genes. For example, in our study, minor allele frequencies of four core SNPs in SLC15A2 (A284A, L350F, P409S and R509K8) were 0.21–0.24 in Asians, but 0.47–0.52 in African-Americans and European-Americans. Other HapMap tagging SNPs also showed similar differences in allele frequencies. In addition, the minor alleles of 11 SNPs among 19 core SNPs were not detected in Asian populations. Because these core SNPs are known to be associated with functional changes in SLC family genes, the frequency differences could result in ethnic differences in drug disposition and response.
Together with allele frequencies of pharmacogenetic core variants, haplotype structures representing the functional unit of the gene32 are particularly useful in correlating drug response phenotypes with a specific gene. However, the haplotype blocks identified in this study were statistically inferred without experimental validation of the molecular linkage of SNPs, and analyzing a large genomic region in small number of samples is likely to result in an error rate that may decrease the precision of haplotype prediction.
In conclusion, our findings give a comprehensive overview on genetic variability, haplotype structure and ethnic diversity of four important pharmacogenetic SLC family transporters. The observed differences in frequency in this study would be valuable for further researches, including pharmacogenetic studies for drug responses.
References
Leabman, M. K., Huang, C. C., DeYoung, J., Carlson, E. J., Taylor, T. R., de la Cruz, M. et al. Natural variation in human membrane transporter genes reveals evolutionary and functional constraints. Proc. Natl Acad. Sci. USA 100, 5896–5901 (2003).
Shu, Y., Sheardown, S. A., Brown, C., Owen, R. P., Zhang, S., Castro, R. A. et al. Effect of genetic variation in the organic cation transporter 1 (OCT1) on metformin action. J. Clin. Investig. 117, 1422–1431 (2007).
Adams, G. A. & Rose, J. K. Structural requirements of a membrane-spanning domain for protein anchoring and cell surface transport. Cell 41, 1007–1015 (1985).
Hediger, M. A., Romero, M. F., Peng, J. B., Rolfs, A., Takanaga, H. & Bruford, E. A. The ABCs of solute carriers: physiological, pathological and therapeutic implications of human membrane transport proteins. Introduction. Pflugers Arch. 447, 465–468 (2004).
Hooiveld, G. J., van Montfoort, J. E., Meijer, D. K. & Muller, M. Function and regulation of ATP-binding cassette transport proteins involved in hepatobiliary transport. Eur. J. Pharm. Sci. 12, 13–30 (2000).
Inui, K., Terada, T., Masuda, S. & Saito, H. Physiological and pharmacological implications of peptide transporters, PEPT1 and PEPT2. Nephrol Dial Transplant. 15 (Suppl 6), 11–13 (2000).
Inui, K. I., Masuda, S. & Saito, H. Cellular and molecular aspects of drug transport in the kidney. Kidney Int. 58, 944–958 (2000).
Pinsonneault, J., Nielsen, C. U. & Sadee, W. Genetic variants of the human H+/dipeptide transporter PEPT2: analysis of haplotype functions. J. Pharmacol. Exp. Ther. 311, 1088–1096 (2004).
Shu, Y., Brown, C., Castro, R. A., Shi, R. J., Lin, E. T., Owen, R. P. et al. Effect of genetic variation in the organic cation transporter 1, OCT1, on metformin pharmacokinetics. Clin. Pharmacol. Ther. 83, 273–280 (2008).
Becker, M. L., Visser, L. E., van Schaik, R. H., Hofman, A., Uitterlinden, A. G. & Stricker, B. H. Genetic variation in the organic cation transporter 1 is associated with metformin response in patients with diabetes mellitus. Pharmacogenomics J. 9, 242–247 (2009).
Song, I. S., Shin, H. J., Shim, E. J., Jung, I. S., Kim, W. Y., Shon, J. H. et al. Genetic variants of the organic cation transporter 2 influence the disposition of metformin. Clin. Pharmacol. Ther. 84, 559–562 (2008).
Leabman, M. K., Huang, C. C., Kawamoto, M., Johns, S. J., Stryke, D., Ferrin, T. E. et al. Polymorphisms in a human kidney xenobiotic transporter, OCT2, exhibit altered function. Pharmacogenetics 12, 395–405 (2002).
Takeda, M., Khamdang, S., Narikawa, S., Kimura, H., Hosoyamada, M., Cha, S. H. et al. Characterization of methotrexate transport and its drug interactions with human organic anion transporters. J. Pharmacol. Exp. Ther. 302, 666–671 (2002).
Uwai, Y., Taniguchi, R., Motohashi, H., Saito, H., Okuda, M. & Inui, K. Methotrexate-loxoprofen interaction: involvement of human organic anion transporters hOAT1 and hOAT3. Drug Metab. Pharmacokinet. 19, 369–374 (2004).
Bleasby, K., Hall, L. A., Perry, J. L., Mohrenweiser, H. W. & Pritchard, J. B. Functional consequences of single nucleotide polymorphisms in the human organic anion transporter hOAT1 (SLC22A6). J Pharmacol Exp Ther. 314, 923–931 (2005).
Thompson, D., Stram, D., Goldgar, D. & Witte, J. S. Haplotype tagging single nucleotide polymorphisms and association studies. Hum. Hered. 56, 48–55 (2003).
Oliphant, A., Barker, D. L., Stuelpnagel, J. R. & Chee, M. S. BeadArray technology: enabling an accurate, cost-effective approach to high-throughput genotyping. Biotechniques (Suppl), 56–58, 60–61 (2002).
Kang, H. J., Song, I. S., Shin, H. J., Kim, W. Y., Lee, C. H., Shim, J. C. et al. Identification and functional characterization of genetic variants of human organic cation transporters in a Korean population. Drug Metab. Dispos. 35, 667–675 (2007).
Fujita, T., Brown, C., Carlson, E. J., Taylor, T., de la Cruz, M., Johns, S. J. et al. Functional analysis of polymorphisms in the organic anion transporter, SLC22A6 (OAT1). Pharmacogenet. Genomics 15, 201–209 (2005).
Devlin, B. & Risch, N. A comparison of linkage disequilibrium measures for fine-scale mapping. Genomics 29, 311–322 (1995).
Barrett, J. C., Fry, B., Maller, J. & Daly, M. J. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics (Oxford, England) 21, 263–265 (2005).
Ganapathy, M. E., Brandsch, M., Prasad, P. D., Ganapathy, V. & Leibach, F. H. Differential recognition of beta-lactam antibiotics by intestinal and renal peptide transporters, PEPT 1 and PEPT 2. J. Biol. Chem. 270, 25672–25677 (1995).
Zhu, T., Chen, X. Z., Steel, A., Hediger, M. A. & Smith, D. E. Differential recognition of ACE inhibitors in Xenopus laevis oocytes expressing rat PEPT1 and PEPT2. Pharm. Res. 17, 526–532 (2000).
Rodriguez, L., Batlle, A., Di Venosa, G., MacRobert, A. J., Battah, S., Daniel, H. et al. Study of the mechanisms of uptake of 5-aminolevulinic acid derivatives by PEPT1 and PEPT2 transporters as a tool to improve photodynamic therapy of tumours. Int. J. Biochem. Cell Biol. 38, 1530–1539 (2006).
Becker, M. L., Visser, L. E., van Schaik, R. H., Hofman, A., Uitterlinden, A. G. & Stricker, B. H. Interaction between polymorphisms in the OCT1 and MATE1 transporter and metformin response. Pharmacogenet. Genomics 20, 38–44 (2010).
Kerb, R., Brinkmann, U., Chatskaia, N., Gorbunov, D., Gorboulev, V., Mornhinweg, E. et al. Identification of genetic variations of the human organic cation transporter hOCT1 and their functional consequences. Pharmacogenetics 12, 591–595 (2002).
Shu, Y., Leabman, M. K., Feng, B., Mangravite, L. M., Huang, C. C., Stryke, D. et al. Evolutionary conservation predicts function of variants of the human organic cation transporter, OCT1. Proc. Natl Acad. Sci. USA 100, 5902–5907 (2003).
Sakata, T., Anzai, N., Shin, H. J., Noshiro, R., Hirata, T., Yokoyama, H. et al. Novel single nucleotide polymorphisms of organic cation transporter 1 (SLC22A1) affecting transport functions. Biochem. Biophys. Res. Commun. 313, 789–793 (2004).
Wang, Z. J., Yin, O. Q., Tomlinson, B. & Chow, M. S. OCT2 polymorphisms and in-vivo renal functional consequence: studies with metformin and cimetidine. Pharmacogenet. Genomics 18, 637–645 (2008).
Tzvetkov, M. V., Vormfelde, S. V., Balen, D., Meineke, I., Schmidt, T., Sehrt, D. et al. The effects of genetic polymorphisms in the organic cation transporters OCT1, OCT2, and OCT3 on the renal clearance of metformin. Clin. Pharmacol. Ther. 86, 299–306 (2009).
Ho, E. S., Lin, D. C., Mendel, D. B. & Cihlar, T. Cytotoxicity of antiviral nucleotides adefovir and cidofovir is induced by the expression of human renal organic anion transporter 1. J. Am. Soc. Nephrol. 11, 383–393 (2000).
Drysdale, C. M., McGraw, D. W., Stack, C. B., Stephens, J. C., Judson, R. S., Nandabalan, K. et al. Complex promoter and coding region beta 2-adrenergic receptor haplotypes alter receptor expression and predict in vivo responsiveness. Proc. Natl Acad. Sci. USA 97, 10483–10488 (2000).
Acknowledgements
This work was supported by a grant from Korea National Institute of Food and Drug Safety Evaluation (NIFDS) funded by Korea Food and Drug Administration (KFDA).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Cheong, H., Kim, H., Na, H. et al. Screening of genetic variations of SLC15A2, SLC22A1, SLC22A2 and SLC22A6 genes. J Hum Genet 56, 666–670 (2011). https://doi.org/10.1038/jhg.2011.77
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2011.77
Keywords
This article is cited by
-
Population pharmacogenomics: an update on ethnogeographic differences and opportunities for precision public health
Human Genetics (2022)
-
Genome-wide association study and genomic heritabilities for blood protein levels in Lori-Bakhtiari sheep
Scientific Reports (2021)
-
Tenofovir-associated kidney disease in Africans: a systematic review
AIDS Research and Therapy (2019)
-
Influence of body weight and UGT2B7 polymorphism on varenicline exposure in a cohort of smokers from the general population
European Journal of Clinical Pharmacology (2019)
-
Identification of two novel genes SLC15A2 and SLCO1B3 associated with maintenance dose variability of warfarin in a Chinese population
Scientific Reports (2017)