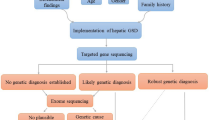
Abstract
Glycogen storage disease type III (GSD III) is an autosomal recessive inborn error of metabolism caused by mutations in the glycogen debranching enzyme amylo-1,6-glucosidase gene, which is located on chromosome 1p21.2. GSD III is characterized by the storage of structurally abnormal glycogen, termed limit dextrin, in both skeletal and cardiac muscle and/or liver, with great variability in resultant organ dysfunction. The spectrum of AGL gene mutations in GSD III patients depends on ethnic group. The most prevalent mutations have been reported in the North African Jewish population and in an isolate such as the Faroe Islands. Here, we present the molecular and biochemical analyses of 22 Tunisian GSD III patients. Molecular analysis revealed three novel mutations: nonsense (Tyr1148X) and two deletions (3033_3036del AATT and 3216_3217del GA) and five known mutations: three nonsense (R864X, W1327X and W255X), a missense (R524H) and an acceptor splice-site mutation (IVS32-12A>G). Each mutation is associated to a specific haplotype. This is the first report of screening for mutations of AGL gene in the Tunisian population.
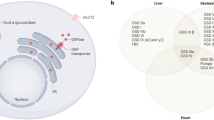
Similar content being viewed by others
Introduction
Glycogen storage disease type III (GSD III) is an autosomal recessive metabolic disorder. It is caused by the mutation of the glycogen debranching enzyme gene (called AGL: amylo-1,6-glucosidase), located on chromosome 1p21.2.1 This gene compound by 35 exons that span a genomic sequence of ∼85 kb.2 It encodes for a single 160-kDa enzyme having two independent catalytic activities: oligo1,4-glucotransferase (EC 2.4.1.25) and amylo-1,6-glucosidase (EC 3.2.1.33). The glycogen-binding site is located in the carboxyl terminal of the protein.3 This enzyme is expressed mainly in liver, skeletal and cardiac muscles. There are different subtypes of the disease depending on the loss of liver and/or muscles activities.4, 5 In 78% of cases, patients lack activity in both their liver and muscle (GSD IIIa subtype), in 15% they lack activity only in liver (GSD IIIb subtype) and in rare cases, selective loss of only one of the two debranching activities (glucosidase or transferase) results in type IIIc or IIId, respectively. These different subtypes result in diverse clinical phenotypes with heterogenic features.6
Molecular analysis of GSD III has been performed in several ethnic populations and over 60 different AGL mutations have been reported to date (Human Gene Mutation Database; http://www.hgmd.cf.ac.uk/ac/gene.php?gene=AGL). In most cases, there is a genetic heterogeneity with a large spectrum of mutations in the same population such as Japanese, Italian and Caucasian populations.7, 8, 9 In populations with high rate of consanguinity, recurrent mutations have been reported, such as the “4455delT” in North African Jewish population,10 “R408X” in the Faroe Islands and “W1327X” in Egyptian and Turkish population.11
Major manifestations of GSD III are hypoglycemia (which may spontaneously improve and disappear with age), hepatomegaly with elevated transaminases, hyperlipidemia, hyperuricemia, skeletal myopathy and cardiomyopathy with increased creatine phosphokinase. There is no specific treatment for GSDs, but diet therapy with nocturnal nasogastric tube feeding and cornstarch, which improves symptoms especially hypoglycemia, reduces the liver size, and resumption overall growth and development.12
In Tunisia, where the ratio of consanguineous marriages still high, the prevalence and recurrence of AGL mutations in GSD III disease is still also unknown. Therefore, we report here, a large clinical, biochemical and genetic study of 22 Tunisian patients affected by GSD III. In addition to the mutations already described in the literature, we found three novels AGL changes. We also analyzed the haplotypes associated to these mutations in order to establish a possible correlation genotype–phenotype.
Materials and methods
Population
This study was approved by local ethics committees and performed with the patients’ and their families’ informed consent.
A total of 22 Tunisian GSD IIIa patients belonging to 20 unrelated families were investigated, originated from different regions of Tunisia (Figure 1).
For each patient, a complete clinical examination was performed followed by a blood sampling for erythrocytes glycogen content and amylo1,6-glucosidase activity measurements. Genomic DNA was extracted from peripheral blood leukocytes using WIZARD® Genomic DNA Purification Kit (Promega, Madison, WI, USA).
Erythrocytes glycogen content
Glycogen concentration was measured in erythrocytes by a colorimetric method.13
Measurement of amylo1, 6-glucosidase activity
Debranching enzyme presents two different activities allowing the two-step debranching of a phosphorylase limit dextrin: a transferase activity (1,4-α-D-glucan: 1,4-α-D-glucan 4-α-D-glycosyltransferase; EC 2.4.1.25) and a hydrolytic activity acting on the α-1,6 linkage at the branch point (amylo-l,6-glucosidase; EC 3.2.1.33).14
Leukocytes were isolated from heparinized blood after lysing the erythrocytes in isotonic ammonium chloride. Leukocyte debranching enzyme was determined fluorimetrically by coupling the production of glucose from “phosphorylase-limit dextrin” to the hexokinase and glucose S-phosphate dehydrogenates reactions. The formation of NADH was measured continuously by a fluorimeter in 440- and 460-nm fluorescent light. The reaction mixture contains 40 mM Tris-acetate buffer (pH=7.0), 1 mM EDTA (pH=7.0), 2 mM MgCl2, 0.22 mM NADP+, 0.2% phosphorylase-limit dextrin, 2 mM ATP, 200 mU hexokinase, 100 mU glucose phosphate dehydrogenase, 250–500 μg leukocyte extract and 2.5 mM NaF. The final pH was 6.4 and the total volume of 0.9 ml. The reaction was carried out at 25±2 °C.15
Genotyping of polymorphic markers
Four microsatellite markers CA/TG flanking the AGL gene in chromosome 1 were amplified.16
They were amplified using 150 ng of genomic DNA in 25 μl PCR reaction containing 5 μl PCR buffer, 1.5 mM MgCl2, 20 mM primer pair mix and 1 U AmpliTaq DNA polymerase (Promega). Amplification conditions were 7 min at 95 °C followed by 35 cycles of 30 s at 94 °C; 30 s at 58 °C; and 30 s at 72 °C, and a final extension for 30 min at 72 °C. PCR products were loaded on an ABI Prism 310 automated genetic analyzer (Applied Biosystem, Foster City, USA). Data were analyzed using the GeneScan and Genotyper softwares (Applied Biosystem). The primer sequences are: AmG24AC (F: 5′-TCTTCATCAAGATGTATAACAATAAAG-3′ and R: 5′-TTTTTCTGGTTTCCCAGATT-3′), D1S2671 (F: 5′-AGAAGATCAACTACCCAAAGAA-3′ and R: 5′-CTCTGCTGCGACTCCA-3′), D1S1658 (F: 5′-GCCATGTCTATTTAATTAGAGTGC-3′ and R: 5′-TGCGTCGAGACTGCAATATA-3′) and AmG29AC (F: 5′-CTGAGGTGGCAGGATCACTT-3′ and R: 5′-TCTCCTGGGGGTGTGTGTAT-3′). Marker informations are available at UCSC Genome Database (http://genome.ucsc.edu/).
Sequence analysis of the AGL gene
Twenty-eight DNA fragments containing all the coding exons (from 3 to 35) and their flanking intron–exon junctions were amplified. The purified fragments were sequenced, using the same PCR primers and the Big Dye Terminator v1.1 on an automated 310 sequencing system (Applied Biosystem, http://www.appliedbiosystems.com). The samples were analyzed with the ABI Sequence Navigator software. Each new mutation was checked and controlled by a new independent amplification and a direct sequencing for each positive sample. To confirm mutations, all sequencing reactions were carried out using the forward and the reverse primers.
The nucleotides of AGL complementary DNA were numbered according to AGL isoform 1 (GenBank accession no. NM_000642). Briefly, PCR was performed in 1.5 mM MgCl2, using 1 U Recombinant Taq DNA polymerase (Invitrogen, Carlsbad, CA, USA) with amplification program beginning by denaturation at 94 °C for 5 min, then 35 cycles of 94 °C denaturing for 30 s, 60 °C annealing for 30 s, 72 °C extension for 30 s and a final step of 7 min at 72 °C. The amplified DNA was purified by Charge Switch® PCR Clean-Up kit (Invitrogen) and then sequenced with the Big Dye terminator cycle sequencing kit. After a cleaning by Wizard®MagneSil™ Sequencing Reaction Clean-Up System kit (Promega), the samples were electrophoresed on an ABI Prism 310 genetic analyzer and analyzed by Sequence Navigator software (Applied Biosystem).
Results
Clinical
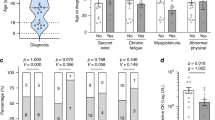
As part of an ongoing study on glycogen storage disease type III, we analyzed 22 cases diagnosed with GSD III (9 boys and 13 girls, sex ratio=0. 57; Table 1). They belong to 20 different families: 12 were originated from Mahdia, 2 from the north of Tunisia (one lives in Beja and the other in Kef) 2 from Sousse, 2 from Zaghouan, one from Gabes of and the last is from Nabeul (Figure 1; Table 1).
Their ages vary between 1 and 14 years with an average of 4.17 years (Table 1). Onset occurred generally before 2 years of age (80% of cases).
Twenty percent of these patients suffer from muscle involvement with cramps, weakness, dyspnea, causing difficulties in daily-life activities.
Biochemical analysis
Laboratory parameters showed raised plasma creatine phosphokinase in seven cases (59%).
There was also an increase in alanine aminotranferease with an average value of 336.9 (22–1072) U 1−1 and aspartate aminotransferase with an average value of 457.45 (112.1246) U 1−1 to varying degrees.
Intra-erythrocytic glycogen was markedly increased with an average of 1350 μg per gHb (⩽200 μg per gHb).
Patients had a negative or null debranching enzyme activity in leukocytes. The debranching enzyme activity determined by difference between the rates of phophorylase limit dextrin and glycogen hydrolysis led to negative or zero values in the 18 patients studied ((phophorylase limit dextrin)–(glycogen)) hydrolysis mU per mg cell protein (Table 1).
Haplotypes and mutations analysis
Genotype analysis of the 22 GSD III subjects showed six different homozygous haplotypes who segregate with the disease and two heterozygote haplotypes (Table 2).
The direct sequencing of AGL gene for these patients allowed the detection of eight different mutations prospectively corresponding to eight particular haplotypes. Three were novel and five were recurrent. All the new mutations were not found in 50 normal Tunisian controls (100 chromosomes).
Based upon haplotype analysis of the AGL gene, we showed that each mutation found was always associated with a specific haplotype (Table 3).
The most frequent mutation identified in this study was the W1327X, found at a homozygote status in 12 patients originated from “Mahdia”. It was also found at the heterozygote status in patient number 22 who comes from the south of Tunisia, associated with R524H (1571G>A in exon 13) (Table 1). The others mutations were: R864X (C2590T in exon 21) identified in two patients, W255X (G765A in exon 7) and IVS32 –12A>G in intron 32.
The three novel mutations were (1) c.3216-3217 del GA (exon 25) found in two patients coming from the region of Sousse. This mutation creates a premature stop codon leading to the interruption of the translation in codon position 1119. (2) c.3033_3036 del AATT (exon 24) in one patient, which creates a premature stop codon at position 1037 and (3) Y1148X (3444C>A transition in exon 27; Table 1).
Discussion
The diagnosis of GSD III needs the combination of clinical, biochemical and molecular investigations. Broad spectrum of mutations affecting AGL gene, makes genetic studies difficult, especially that this gene is very large. A deeper study of a specific population would find some peculiarities shared by all or some groups of this population inherited from a founder effect, generation after generation. Later, these features will be targeted in the routine analysis conferring a gain in time and money.
In this vein, we started a complete study of a Tunisian population and tried to compare our results with other ethnic groups, looking for differences and similarities.
In this study, we investigate 22 GSD III patients (9 boys and 13 girls). Most of them have manifested fasting hypoglycemic and/or hepatomegaly. These manifestations were the predominant features.17 Hypoglycemia is rare in neonates but often manifests after 2 years of age, when parents reduce feeding frequency.18 Generally, hypoglycemia is the primary clinical manifestation of GSD III.19 It is caused by the defect of glycogenolysis due to the deficient activity of debranching enzyme. Only a small portion of the glycogen stored in the liver can be available for glucose homeostasis. As a result, patients may experience significant hypoglycemia, especially after a relatively short fast.6 Hepatic symptoms may be so mild that, in absence of hypoglycemia, the diagnosis could be not suspected until adulthood, when the patient first manifests signs of neuromuscular affection.20, 21
Skeletal and cardiac muscle involvement occurs much more later.17 It is detected by the elevation of plasma creatine kinase and transaminase (alanine aminotranferease and aspartate aminotransferase).22 This, reflect cytolysis caused by the cellular accumulation of abnormal structured glycogen.23
All theses clinical and biological parameters are not specific for type III glycogenosis. They can be present in other metabolic diseases specially type I glycogenosis. However, some particularities lead to make the difference.24 Indeed, transaminases show large elevation in GSD III compared with the other diseases.25 Also, the intra-erythrocyte glycogen elevation, after several hours fasting, is in favor of type III GSD.
Only measurement of debranching enzyme activity and genetic analysis can confirm the diagnosis.
In this study, we found that most of our patients showed negative or null enzymatic activity in leukocytes, which confirms that the use of these cells is widely sufficient and does not need liver or muscle biopsies.
Allelic and genetic heterogeneity found in our population can be explained by the big number of migratory/invasive flows affected North Africa in the past and could have generated genetic admixtures among North African populations, notably, Phoenician presence for 12 centuries from eleventh century BC to 146 AC; Roman settlement up to the fifth century AC; Vandals’ and Byzantines’ invasions in 455 AC and 533 AC respectively; Arab expansion into North Africa during seventh century AC; migration waves of Arab tribes during the eleventh century AC; and finally Ottoman settlement from 1515 AC up to the time of French colonization in 1830 AC.26, 27, 28
From the five mutations that we found (Table 1), three will be explained W1327X, IVS32–12A>G and R864X. The W1327X was found in 12 patients originated from Mahdia. These patients shared a common haplotype that led us to hypothesize a founder effect, with a common ancestor. Such founder mutation is of great interest for genetic counseling of glycogen storage disease. In fact, genotyping study can be used in the screening of AGL gene deficiency before targeting the causative mutation by direct sequencing.
This mutation is also prevalent in Turkish, Egyptian, Caucasian and German–Ukraine patients.11, 29 Turkish and Egyptian patients’ haplotype are different, suggesting that W1327X can be a recurrent mutation in various ethnic groups.30 The IVS32–12A>G mutation found in one patient (number 13), has been previously reported in Japanese population.31 This mutation causes retention of the 11 bp intronic sequence in the debrancher mRNA. It is predicted to result in a truncated enzyme with loss of 112 carboxyl-terminal amino acids as a result of premature termination. Okubo et al. demonstrated that this mutation is responsible for GSD type IIIb, which is an exception because, normally, only mutations situated in exon 3 can give rise to this form of the disease. Haplotyping studies demonstrated that this mutation occurred independently in Japanese and Chinese patients and also in our patient (Table 2).32
The R864X nonsense mutation, found in two patients (number 15 and 16), has been shown to account for ∼10% of GSD III Caucasian patients.9, 33 This nonsense homozygote mutation leads to premature stop codon, a tranqued protein termination, which completely abolishes the enzyme activity.
Similarly, the two others heterozygote mutations: nonsense mutation W255X in exon 7 is predicted to cause a truncated enzyme lacking the glycogen-binding site in the carboxyl terminal34 and R524H mutation, located in exon 13, is also known to alter the catalytic function of the transferase domain.35
The three novel mutations, Y1148X, 3033_3036del AATT and 3216-3217del GA, were predicted to lead to a premature stop codon, respectively, at positions 1148, 1037 and 1119 and abolishing the enzyme activity. They are responsible for typical GSD III.
These results are consistent with the fact that the majority of the mutations reported are nonsense mutations, deletions, insertions and splicing mutations.36
The genetic heterogeneity of AGL gene is even higher, considering all the reported GSD III mutations in different ethnic groups. The mutations cover the whole gene length and any ‘‘hot spot’’ region does not seem to exist.
The GSD III disease is known to lack almost invariably clear links between the genotype and clinical phenotype. Therefore, it is not possible for each case to reliably relate the specific severity of the symptoms to the corresponding genetic defect.37, 38 The common element that these clinical histories shared is the importance of a suitable dietary regimen subsequent to an early diagnosis. It is well known that keeping hypoglycemic episodes under control and providing a protein-rich diet might be beneficial to these patients, although the latter assertion is yet to be formally proven.
In summary, we identified eight AGL mutations, including three novel ones. Our molecular study concerning GSD III patients of different ethnic ancestry showed haplotype heterogeneity of the AGL mutations.
This is the first AGL gene mutation report for Tunisian population. In addition, this is the first molecular study combined between genotyping of polymorphic markers and direct sequencing. This direct comparative study of AGL gene showed that each haplotype found corresponds to a specific mutation. Knowledge of the basic defect will offer a better basis for genetic counseling of couples at risk and for first trimester prenatal diagnosis by direct genomic analysis. So far, three families have requested and benefited from a prenatal diagnosis.
Accession codes
Change history
26 March 2012
This article has been corrected since advance Online Publication, and a corrigendum is also printed in this issue
References
Bao, Y., Dawson, T. L. Jr & Chen, Y. T. Human glycogen debranching enzyme gene (AGL): complete structural organization and characterization of the 5′ flanking region. Genomics 38, 155–165 (1996).
Chen, Y. T. in The Metabolic and Molecular Bases of Inherited Disease (eds Scriver, C. R. BAL, Sly, W. S. and Valle, D.). pp 1521–1551 (McGraw-Hill, New York, 2001).
Bao, Y., Yang, B. Z., Dawson, T. L. Jr & Chen, Y. T. Isolation and nucleotide sequence of human liver glycogen debranching enzyme mRNA: identification of multiple tissue-specific isoforms. Gene 197, 389–398 (1997).
Ding, J. H., de Barsy, T., Brown, B. I., Coleman, R. A. & Chen, Y. T. Immunoblot analyses of glycogen debranching enzyme in different subtypes of glycogen storage disease type III. J. Pediatr. 116, 95–100 (1990).
Van Hoof, F. & Hers, H. G. The subgroups of type 3 glycogenosis. Eur. J. Biochem. 2, 265–270 (1967).
Coleman, R. A., Winter, H. S., Wolf, B., Gilchrist, J. M. & Chen, Y. T. Glycogen storage disease type III (glycogen debranching enzyme deficiency): correlation of biochemical defects with myopathy and cardiomyopathy. Ann. Intern. Med. 116, 896–900 (1992).
Lucchiari, S., Pagliarani, S., Salani, S., Filocamo, M., Di Rocco, M., Melis, D. et al. Hepatic and neuromuscular forms of glycogenosis type III: nine mutations in AGL. Hum. Mutat. 27, 600–601 (2006).
Okubo, M., Horinishi, A., Takeuchi, M., Suzuki, Y., Sakura, N., Hasegawa, Y. et al. Heterogeneous mutations in the glycogen-debranching enzyme gene are responsible for glycogen storage disease type IIIa in Japan. Hum. Genet. 106, 108–115 (2000).
Shaiu, W. L., Kishnani, P. S., Shen, J., Liu, H. M. & Chen, Y. T. Genotype-phenotype correlation in two frequent mutations and mutation update in type III glycogen storage disease. Mol. Genet. Metab. 69, 16–23 (2000).
Parvari, R., Moses, S., Shen, J., Hershkovitz, E., Lerner, A. & Chen, Y. T. A single-base deletion in the 3′-coding region of glycogen-debranching enzyme is prevalent in glycogen storage disease type IIIA in a population of North African Jewish patients. Eur. J. Hum. Genet. 5, 266–270 (1997).
Aoyama, Y., Ozer, I., Demirkol, M., Ebara, T., Murase, T., Podskarbi, T. et al. Molecular features of 23 patients with glycogen storage disease type III in Turkey: a novel mutation p.R1147G associated with isolated glucosidase deficiency, along with 9 AGL mutations. J. Hum. Genet. 54, 681–686 (2009).
Ismail, H. Glycogen storage disease type III presenting with secondary diabetes and managed with insulin: a case report. Cases J. 2, 6891 (2009).
Sidbury, J B. Jr, Gitzelmann, R. & Fisher, J. The glycogenoses. Further observations on glycogen in erythrocytes of patients with glycogenosis. Helv. Paediatr. Acta 16, 506–516 (1961).
Hers, H. G. & Mathieu, M. (eds) The Mechanism of Action of Amylo-l,6 glucosidase. A Ciba Foundation Symp. London: Churchill 1984.
Brown, D. H., Waindle, L. M. & Brown, B. I. The apparent activity in vivo of the lysosomal pathway of glycogen catabolism in cultured human skin fibroblasts from patients with type III glycogen storage disease. J. Biol. Chem. 253, 5005–5011 (1978).
Gribaa, M., Salih, M., Anheim, M., Lagier-Tourenne, C., H’Mida, D., Drouot, N. et al. A new form of childhood onset, autosomal recessive spinocerebellar ataxia and epilepsy is localized at 16q21-q23. Brain 130 (Pt 7), 1921–1928 (2007).
Bhuiyan, J., Al Odaib, A. N. & Ozand, P. T. A simple, rapid test for the differential diagnosis of glycogen storage disease type 3. Clin. Chim. Acta 335, 21–26 (2003).
Labrune, P., Eberschweiler, P. T., Boudjemline, A. M., Hubert-Buron, A., Petit, F. & Gajdos, V. Natural history of hepatic glycogen storage diseases [article in French]. Presse Med. 37, 1172–1177 (2008).
Oh, S. H., Park, H. D., Ki, C. S., Choe, Y. H. & Lee, S. Y. Biochemical and molecular investigation of two Korean patients with glycogen storage disease type III. Clin. Chem. Lab. Med. 46, 1245–1249 (2008).
Dai, Y. J., Chen, L., Guo, Y. P., Ren, H. T., Zhao, Y. H., Wei, M. et al. Clinical and pathological features of glycogen storage disease type III [article in Chinese]. Zhonghua Yi Xue Za Zhi. 89, 1064–1066 (2009).
Nadaj-Pakleza, A. A., Vincitorio, C. M., Laforet, P., Eymard, B., Dion, E. & Teijeira, S. Permanent muscle weakness in McArdle disease. Muscle Nerve 40, 350–357 (2009).
Labrune, P., Huguet, P. & Odievre, M. Cardiomyopathy in glycogen-storage disease type III: clinical and echographic study of 18 patients. Pediatr. Cardiol. 12, 161–163 (1991).
Maire, I., Baussan, C., Moatti, N., Mathieu, M. & Lemonnier, A. Biochemical diagnosis of hepatic glycogen storage diseases: 20 years French experience. Clin. Biochem. 24, 169–178 (1991).
Shin, Y. S. Glycogen storage disease: clinical, biochemical, and molecular heterogeneity. Semin. Pediatr. Neurol. 13, 115–120 (2006).
Lee, P., Burch, M. & Leonard, J V. Plasma creatine kinase and cardiomyopathy in glycogen storage disease type III. J Inherit Metab Dis. 18, 751–752 (1995).
Bosch, E., Clarimon, J., Perez-Lezaun, A. & Calafell, F. STR data for 21 loci in northwestern. Africa Forensic Sci. Int. 116, 41–51 (2001).
Nebel, A., Landau-Tasseron, E., Filon, D., Oppenheim, A. & Faerman, M. Genetic evidence for the expansion of Arabian tribes into the Southern Levant and North Africa. Am. J. Hum. Genet. 70, 1594–1596 (2002).
Lefevre-Witier, P., Aireche, H., Benabadji, M., Darlu, P., Melvin, K., Sevin, A. et al. Genetic structure of Algerian populations. Am. J. Hum. Biol. 18, 492–501 (2006).
Schoser, B., Glaser, D. & Muller-Hocker, J. Clinicopathological analysis of the homozygous p.W1327X AGL mutation in glycogen storage disease type 3. Am. J. Med. Genet. A 146A, 2911–2915 (2008).
Aoyama, Y., Endo, Y., Ebara, T., Murase, T., Shin, Y. S., Podskarbi, T. et al. Novel AGL mutation in a Turkish patient with glycogen storage disease type IIIa. Pediatr. Int. 52, 145–147 (2010).
Okubo, M., Horinishi, A., Nakamura, N., Aoyama, Y., Hashimoto, M. & Endo, Y. A novel point mutation in an acceptor splice site of intron 32 (IVS32 A-12-->G) but no exon 3 mutations in the glycogen debranching enzyme gene in a homozygous patient with glycogen storage disease type IIIb. Hum. Genet. 102, 1–5 (1998).
Horinishi, A., Okubo, M., Tang, N. L., Hui, J., To, K. F., Mabuchi, T. et al. Mutational and haplotype analysis of AGL in patients with glycogen storage disease type III. J. Hum. Genet. 47, 55–59 (2002).
Shen, J., Bao, Y., Liu, H. M., Lee, P., Leonard, J. V. & Chen, Y. T. Mutations in exon 3 of the glycogen debranching enzyme gene are associated with glycogen storage disease type III that is differentially expressed in liver and muscle. J. Clin. Invest. 98, 352–357 (1996).
Endo, Y., Horinishi, A., Vorgerd, M., Aoyama, Y., Ebara, T., Murase, T. et al. Molecular analysis of the AGL gene: heterogeneity of mutations in patients with glycogen storage disease type III from Germany, Canada, Afghanistan, Iran, and Turkey. J. Hum. Genet. 51, 958–963 (2006).
Nakayama, A., Yamamoto, K. & Tabata, S. Identification of the catalytic residues of bifunctional glycogen debranching enzyme. J. Biol. Chem. 276, 28824–28828 (2001).
Shen, J. J. & Chen, Y. T. Molecular characterization of glycogen storage disease type III. Curr. Mol. Med. 2, 167–175 (2002).
Lucchiari, S., Fogh, I., Prelle, A., Parini, R., Bresolin, N., Melis, D. et al. Clinical and genetic variability of glycogen storage disease type IIIa: seven novel AGL gene mutations in the Mediterranean area. Am. J. Med. Genet. 109, 183–190 (2002).
Lucchiari, S., Donati, M. A., Melis, D., Filocamo, M., Parini, R., Bresolin, N. et al. Mutational analysis of the AGL gene: five novel mutations in GSD III patients. Hum. Mutat. 22, 337 (2003).
Acknowledgements
We thank Mrs Ahlem Msakni, Mrs Sihem Sassi, Mr Hammadi Ben Khelifa and Mr Hedi Laatiri for their excellent technical assistance. This work was supported by the Ministry of High Education, Scientific Research and Technology in Tunisia.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Mili, A., Ben Charfeddine, I., Mamaï, O. et al. Molecular and biochemical characterization of Tunisian patients with glycogen storage disease type III. J Hum Genet 57, 170–175 (2012). https://doi.org/10.1038/jhg.2011.122
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2011.122
Keywords
This article is cited by
-
The biallelic novel pathogenic variants in AGL gene in a chinese patient with glycogen storage disease type III
BMC Pediatrics (2022)
-
Glycogen debranching pathway deduced from substrate specificity of glycogen debranching enzyme
Glycoconjugate Journal (2022)
-
A mutation analysis of the AGL gene in Korean patients with glycogen storage disease type III
Journal of Human Genetics (2014)
-
Neonatal Hypoglycemia
The Indian Journal of Pediatrics (2014)
-
Molecular and biochemical characterization of a novel intronic single point mutation in a Tunisian family with glycogen storage disease type III
Molecular Biology Reports (2013)