Abstract
The field of human genetic variations has progressed rapidly over the past few years. It has added much information and deepened our knowledge and understanding of the diversity of genetic variations in the human genome. This significant progress has been driven mainly by the developments of microarray and next generation sequencing technologies. The array-based methods have been widely used for large-scale copy number variation (CNV) detection in the human genome. The arrival of next generation sequencing technologies, which enabled the completion of several whole genome resequencing studies, has also resulted in a massive discovery of genetic variations. These studies have identified several hundred thousand short indels and a total of thousands of CNVs and other structural variations in the human genome. The discovery of these ‘newer’ types of genetic variations, indels, CNVs and copy neutral variations (inversions and translocations) has also widened the scope of genetic markers in human genetic and disease gene mapping studies. The aim of this review article is to summarize the latest developments in the discovery of human genetic variations and address the issue of inadequate coverage of genetic variations in the current genome-wide association studies, which mainly focuses on common SNPs. Finally, we also discuss the future directions in the field and their impacts on next generation genome-wide association studies.
Similar content being viewed by others
Introduction
Human genetic variations are the differences in DNA sequence within the genome of individuals in populations. Genetic variations in the human genome can take many forms, including single nucleotide changes or substitutions; tandem repeats; insertions and deletions (indels); additions or deletions that change the copies number of a larger segment of DNA sequence; that is, copy number variations (CNVs); other chromosomal rearrangements such as inversions and translocations (also known as copy neutral variations); and copy neutral loss of heterozygosity (LOH) or homozygosity. These genetic variations span a spectrum of sizes from single nucleotides to megabases. Single nucleotide substitutions or alterations, as implied in the terminology, involve a change in a single nucleotide at a particular locus in the DNA sequence, such as restriction fragment length polymorphisms (RFLPs), single nucleotide polymorphisms (SNPs) and single nucleotide indels. On the other extreme, CNVs, inversions, translocations and LOHs encompass larger segments of DNA sequences that range from kilobases to megabases (>1 kb), whereas tandem repeats and indels fall in between the extremes (from >1 bp to 1 kb).
In general, these genetic variations take place naturally in the human genome, and they are the footprints of errors or mistakes that occur in DNA replication during cell division, although external agents, such as viruses and chemical mutagens, can also induce changes in the DNA sequence. The occurrence of each type of genetic variation is mediated by different mechanisms; nonetheless, most of these molecular events or processes are currently unclear and are still being investigated. For example, several mechanisms have been proposed to explain the widespread occurrence of CNVs in the human genome, such as nonallelic homologous recombination and nonhomologous end joining.1 However, for copy neutral LOHs, the homozygosity could have resulted from uniparental isodisomy and autozygosity.2 Regardless of the molecular mechanisms or processes that generated the genetic variations, they can be broadly classified as either somatic or germline variations depending on whether they arose from mitosis or meiosis, respectively.
The field of human genetic variations has advanced considerably over the past five years. It has added much information and deepened our knowledge and understanding of the complexity and diversity of genetic variations in the human genome. In addition to the physical mapping of different types of genetic variations, such as RFLPs in the 1980s,3 tandem repeats in the 1990s,4 and SNPs,5, 6 indels,7 CNVs8, 9, 10 and LOHs2 after the new millennium, the data of their biological functional roles; for example, their effects on or associations with mRNA expression levels, alternative splicing processes and other molecular and regulatory processes have also been accumulating.11, 12, 13, 14 Furthermore, these genetic variations were also found to be associated with various human diseases, including monogenic and complex diseases.14, 15, 16, 17, 18, 19, 20, 21, 22
Presently, research in genetic variations is drawing much attention and effort from the genetics community, as evident from the initiation of the 1000 Genomes Project, which has a major aim to construct the most detailed map of genetic variations in the human genome.23, 24 The non-SNP genetic variations certainly have the potential of becoming the next generation genetic markers in human genetic and disease gene mapping studies. The ‘disease gene mapping’ refers to mapping of genetic loci which may or may not contain genes that are associated with diseases. This review will focus on the discovery of different types of genetic variations and their use as genetic markers in disease gene mapping studies in the past, present and future.
Categories of genetic variations
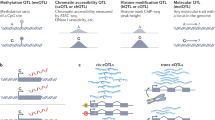
There are issues and problems in categorizing genetic variations into distinct groups, and a clear consensus in defining genetic variations has not been achieved. As a result, the distinction for some of the genetic variations is rather vague at this time. Although SNPs are defined as single nucleotide substitutions, sometimes single nucleotide insertions or deletions also fall under this category (Figure 1a). In general, point mutations include both single nucleotide substitutions and single nucleotide indels, although they are only classified as such when their population frequencies are less than 1%. This is different from polymorphisms, terminology of which is reserved for those genetic variations with population frequencies higher than the arbitrary cutoff of 1% similar to SNPs.
Tandem repeats can be broadly divided into two classes: short tandem repeats (STRs) usually refer to tandem repeats in which the sequence length is eight nucleotides or less, and longer tandem repeats are labeled as variable number tandem repeats (VNTRs; Figure 1b). They are also known as microsatellites and minisatellites, respectively. As such, it is apparent that the distinction between the two classes is solely based on the length of the repeated sequence, but it is only an arbitrary cutoff. The most common types of microsatellites are di-, tri- and tetra-nucleotide repeats. However, repeats of identical nucleotide of several bases or longer in the length; that is, consecutive identical nucleotides in the DNA sequence are known as homopolymer sequences; for example, GGGGG or AAAAA. Although the sequence in the tandem repeats is simple compared with other more complex DNA sequence changes or rearrangements, these simple sequences can be repeated from tens to hundreds of times, thus creating a high heterozygosity or allelic diversity.25, 26
The boundary or distinction between CNVs and indels is even more obscure. In the Database of Genomic Variants (DGV; http://projects.tcag.ca/variation/), deletions and duplications/insertions larger than 1 kb are classified as ‘CNVs’, whereas those between 100 bp to 1 kb are grouped as ‘InDels’. As such, the remaining several hundred thousands of indels in the range of several nucleotides to tens of nucleotides, which were identified in the recent whole genome resequencing experiments, do not currently have their own category.27, 28, 29, 30, 31, 32, 33 For example, Wang et al. (2008)29 found ∼140 000 indels within 1–3 bp in the Han Chinese YH genome, and ∼400 000 indels defined from 1 to 16 bp were also detected in the African NA18507 genome by Bentley et al. (2008).30 Perhaps a new category such as ‘short indels’ needs to be created to fit them in, and those indels between 100 bp to 1 kb should probably be renamed as ‘intermediate indels’ (Figures 1c and d). Similar to SNPs, common CNVs with population frequencies of 1% or higher are known as copy number polymorphisms. However, in some studies, CNVs that are detected in two or more individuals are also considered as copy number polymorphisms.9
However, apart from single nucleotide changes, such as SNPs, all the genetic variations can be broadly grouped under the umbrella of structural variations.34 It is even more confusing when a variety of names are used to describe essentially the same genetic variation. For example, large-scale copy number variants and intermediate-sized variants have been used to describe CNVs before this terminology was introduced.35 Some comparative genomic hybridization array-based studies used chromosomal gains and losses to indicate duplications and deletions, respectively.36 Despite the various categories of genetic variations and terminologies that have been used, it is noteworthy that the definitions or sizes are rather arbitrary. Furthermore, classifications are without biological basis; that is, they are not classified by the mechanisms that mediated their occurrences. Instead, the classification is simply based on the patterns of DNA sequence changes and their sizes. As such, it is more important to describe the characteristics of the genetic variations that are being discovered and identified, rather than be concerned about their respective categories.
The evolution of genetic markers in disease gene mapping
Genetic variations in the human genome are useful as genetic markers for many applications in different areas, such as forensic investigations (for example, genetic or DNA fingerprinting), routine clinical tests (for example, human leucocyte antigen typing for hematopoietic stem cell or organ transplantation), prediction of drug responses or the tailoring of prescription doses (for example, genotyping tests for the SNPs in the thiopurine methyltransferase (TPMT) gene to predict patient responses to 6-mercaptopurine) and population genetics studies (for example, studies of human migration patterns).37, 38, 39, 40 Furthermore, they have also been widely used as genetic markers in disease gene mapping, such as family linkage and genetic association studies to identify the susceptibility loci or genes for monogenic and complex diseases.
Different genetic variations have different characteristics, and their applications are influenced by a number of factors. Tandem repeats such as minisatellites and microsatellites are highly variable or polymorphic in human populations, as such, they have higher allelic states and are more informative than the biallelic genetic markers, such as SNPs. Unlike SNPs in which a single nucleotide substitution will only give rise to two alleles, each repeat in minisatellites and microsatellites is considered as one allelic state. The genetic variations that occur in more than two allelic states are known as multiallelic markers. Owing to their inherent features, tandem repeats have been widely used in genetic fingerprinting and as the genetic markers in linkage studies to locate the chromosomal regions harboring the mutations or genes for monogenic or familial disorders, complex diseases and quantitative traits.41, 42, 43, 44 Although tandem repeats are more informative than SNPs at the individual marker level, their number is far less than the several million SNPs in the human genome. Thus, tandem repeats are not ideal genetic markers for applications that require high marker density or resolution, such as genome-wide association studies (GWASs), in which several hundred thousand of SNPs are needed. In GWAS, a large number of genetic markers are required spanning the whole genome, to achieve comprehensive coverage and adequate statistical power to detect unknown disease variants through linkage disequilibrium (LD).45, 46 In other words, the disease variants would not be detected if no markers in strong LD with them were genotyped.
Apart from the inherent characteristics of genetic variations such as their allelic diversity and abundance in the human genome, their applications are also influenced by technological developments. The rapid advances of high-throughput SNPs genotyping technologies have enabled the genotyping task of several hundreds of thousands to one million SNPs to be done efficiently on thousands of samples in GWAS. On the contrary, no high-throughput method was developed to assay microsatellites on a whole genome scale.47, 48, 49 This technological development, together with their abundance in the human genome, have resulted in SNPs becoming the primary genetic markers used in more than 450 GWAS that have been published to date (A Catalog of Published Genome-Wide Association Studies: http://www.genome.gov/26525384). In fact, almost all the GWAS have used the commercially available whole genome SNPs genotyping arrays from Illumina (San Diego, CA, USA), Affymetrix (Santa Clara, CA, USA).
In the past, researchers had relied solely on RFLPs and tandem repeats as the genetic markers in disease gene mapping studies. The RFLPs were used in linkage studies before the discovery of tandem repeats. Since the availability of the linkage map for microsatellites, RFLPs were mainly used as the genetic markers in candidate gene association studies, in which PCR–RFLP genotyping assay was commonly applied. However, microsatellites were widely used as the genetic markers in linkage studies.41, 42, 43, 44 These genetic variations have been used as the markers in human genetic studies for more than 20 years until the completion of the Human Genome Project 50 and the finding of millions of SNPs by the International SNP Map Working Group and other studies.5, 6 Thereafter, SNPs became the primary markers in genetic association studies, and also replaced microsatellites in some linkage studies.
Although SNPs have been studied in detail over the past decade, a comparable progress in the studies of other genetic variations, such as indels, CNVs and LOHs has not been achieved. In fact, CNVs had only started gaining some attention from the genetics community when the finding of several hundreds of deletions and duplications was first reported in 2004.51, 52 Similarly, no large-scale attempt was made to identify indels until 2006, in which a study found several hundreds of thousands of indels in the human genome.7 The commonness of LOHs or homozygosity regions in the genomes of outbred populations was also under appreciated until the first report appeared in 2006.2 However, the richness of genetic variations in the human genome has recently been further corroborated by the several whole genome resequencing studies, revealing plenty of new SNPs, indels, CNVs and other structural variations.27, 28, 29, 30, 31, 32, 33 The technological developments have facilitated and accelerated the process of identifying genetic variations, especially with the arrival of next generation sequencing technologies, which have made whole genome resequencing and the 1000 Genomes Project feasible.53, 54, 55
In recent years, many studies have been done to directly examine the associations of CNVs with complex diseases using SNP genotyping arrays. These studies have yielded some exciting results for several diseases, such as schizophrenia and autism.56, 57, 58 Therefore, it further supports the use of CNVs as genetic markers to uncover new susceptibility loci for future disease association studies. Interestingly, genome-wide homozygosity mapping approaches have also been applied to dissect the genetic basis of complex diseases and have successfully identified a number of susceptibility loci for schizophrenia.22 Conversely, short indels have not been directly interrogated in GWAS, but how much they can be tagged indirectly through LD by the SNPs in genotyping arrays is unclear. Unlike CNVs and homozygosity mapping, which can be studied by SNPs genotyping arrays, no high-throughput method has been designed and developed to investigate short indels on a genome-wide scale. Direct detection and interrogation of short indels requires sequencing-based methods as demonstrated in the whole genome resequencing studies. As a result they cannot be used effectively as genetic markers in GWAS at the time.
In the following sections, we will discuss the genetic variations and markers in the past (RFLPs and tandem repeats), present (SNPs) and future (CNVs, indels, inversions, translocations and LOHs). The use of ‘past, present and future’ genetic variations is only a ‘time concept’, to illustrate the time of their discoveries and the time when they are most commonly used as genetic markers. For example, RFLPs and tandem repeats were mainly discovered in 1980s and 1990s, so they are considered as the past genetic variations or markers, but this does not mean that they are totally obsolete nowadays or that they are no longer used in human genetic studies. However, although the commonness of CNVs, indels and LOHs in the human genome have already been reported several years ago, they are considered as future genetic variations or markers because they have yet to be ‘intensively and completely’ studied or discovered in the human genome. In addition, so far these newer genetic variations have not been widely used as markers in disease gene mapping.
Past
Restriction fragment length polymorphisms
The RFLPs are single nucleotide substitutions that alter the cutting sites of restriction enzymes. They were one of the earliest genetic markers used in disease gene mapping. The genetic linkage map of RFLPs was constructed in the 1980s.59 The use of RFLPs as genetic markers is based on their ability to create or eliminate the cutting sites of restriction enzymes to distinguish between two alleles. With the invention of the molecular technique PCR, alleles of RFLPs are usually determined by PCR-based methods, such as PCR–RFLP.
In PCR–RFLP assay, one set of probes or PCR primers (forward and reverse primers) are designed to amplify the DNA sequence that contains the RFLP. The PCR amplicons are then followed by restriction enzyme digestion and gel electrophoresis to separate the digestion products. As an example to illustrate the principle of the PCR–RFLP method, the PCR amplicons of G allele will be cut by the restriction enzyme but not for the C allele (a G>C substitution), assuming that there is only one cutting site in the PCR amplicon. Therefore, if all the PCR amplicons remain intact after restriction enzyme digestion (appearing as a single band in gel electrophoresis), this result shows the presence of two C alleles and the genotype is the homozygote CC. Conversely, all the PCR amplicons will be digested by the restriction enzyme for the homozygous GG genotype (two bands in gel electrophoresis for which the sizes are smaller than the PCR amplicon size), and a mixture of three bands suggests the presence of both alleles (Figure 2).
One of the major limitations of using RFLPs as genetic markers is that single nucleotide alterations do not necessarily alter the cutting sites of restriction enzymes. In other words, those single nucleotide substitutions that are not digested by restriction enzymes cannot be studied by PCR–RFLP method. As a result, their numbers are limited. Furthermore, PCR–RFLP is a tedious, laborious and low-throughput genotyping method. Nevertheless, PCR–RFLP has still been widely used in disease gene mapping studies at least before the arrival and feasibility of SNPs genotyping arrays or other higher throughput genotyping methods, such as MassARRAY iPLEX, Invader and SNPlex genotyping assays.60, 61, 62 As RFLPs are single nucleotide substitutions, thus they are actually a subset of SNPs.
Tandem repeats
In addition to RFLPs, the earliest genetic markers also included tandem repeats. The more widespread distribution of microsatellites (>100 000) in the human genome and their higher allelic diversity than RFLPs have made them to be commonly used as the genetic markers in linkage studies for monogenic disorders and complex diseases. Similarly, microsatellite also out-performed VNTRs in terms of their numbers, where there are only a few thousand VNTRs in the human genome.26 The availability of the genetic linkage map of microsatellites has resulted in the immense success of linkage studies in identifying genes for monogenic disorders.4 In contrast, only limited success was achieved in dissecting the genetic basis of complex disease using linkage analysis. For complex diseases, the linkage regions identified were mostly irreproducible and inconsistent, and so far, only a handful disease associated genes, such as CARD15/NOD2 (Crohn's disease), PTPN22 (type-1 diabetes), TCF7L2 (type-2 diabetes) and STAT4 (rheumatoid arthritis and systematic lupus erythematosus), were identified through linkage and positional cloning strategies.63, 64, 65, 66
The failure of linkage studies in interrogating the genetic basis of complex diseases is not due to the inappropriateness of the genetic markers (microsatellites) used to locate the genomic regions that harbor the disease genes, but is instead attributable to the study design. Linkage mapping is a powerful and effective approach to detect rare and highly penetrant mutations, and is best suited for diseases that segregate according to Mendelian inheritance. In contrast, complex diseases are characterized by genetic heterogeneity (multiple genetic variants with incomplete penetrance), and the phenotypes are consequences of complex interactions of genetic factors and environmental exposures.67
The arrival of high-throughput SNP genotyping technologies and the ease of genotyping thousands of SNPs in a microarray have also replaced the use of microsatellites in some linkage studies.68, 69, 70, 71 In classical family linkage studies, a few hundred microsatellites are already sufficient to cover the whole genome. However, this number can be substituted by about 10 000 SNPs to provide a comparable or even greater amount of genetic information.72, 73 The need for a significantly larger number of SNPs is because of their lower heterozygosity as opposed to multiallelic genetic markers. Although microsatellite is more informative at the individual marker level, this can be superseded by a large number of SNPs.
Undoubtedly, microsatellites have been widely used in genome-wide linkage studies, but not in GWAS for complex diseases. Hitherto, there are only a few studies that have genotyped microsatellites in GWAS, and they have adopted a pooling strategy of DNA samples to reduce the amount of genotyping work.74, 75 This is mainly due to the need of genotyping a substantially larger number of microsatellites in GWAS (∼20 000–30 000 markers) compared with linkage studies (∼500 markers). The need for a larger number of microsatellites in GWAS is due to the weaker LD in unrelated individuals, as compared with family members in which there are only a limited number of recombination events. In addition, a larger sample size is also needed in GWAS to achieve adequate statistical power to detect genetic variants with modest effect sizes for complex diseases. Finally, there is a lack of high-throughput methods to assay microsatellites, and this is one of the major reasons that microsatellites have decreased in popularity in the SNP era. However, evidence is now increasing to support the potential functional roles of tandem repeats (tri-nucleotide repeats) and their variation could be associated with human complex diseases. Therefore, they should be reconsidered in the future genetic association.16, 76
Present
Single nucleotide polymorphisms
The completion of the Human Genome Project is a major scientific development in human genomics and biomedical sciences. The reference DNA sequence has provided the basis for studying genetic variations in the human genome among individuals in populations. While the Human Genome Project was about to be completed, genetic variations in particular SNPs were also being uncovered. In 2001, the International SNP Map Working Group identified 1.42 million SNPs in the human genome.5 Currently, more than 17 million SNPs in human genome have been documented in the dbSNP. As a large number of SNPs has been reported, it is unavoidable that some of the entries in the database are actually errors or artifacts rather than ‘genuine SNPs’. In fact, a false positive rate of 15–17% was estimated for dbSNP.77 Therefore, large scale validation in population-based studies would be necessary and important to authenticate them. To bridge this gap of information, the International HapMap Project was conceived in 2003 with the aim to validate several million SNPs in the dbSNP, to obtain the SNP and genotype frequencies information, as well as to study their correlation or LD patterns in populations of European, Asian and African ancestry. These populations are the US Utah population with Northern and Western European ancestry (CEU), Han Chinese from Beijing (CHB), Japanese from Tokyo (JPT) and Yoruba from Ibadan, Nigeria (YRI).78
In general, a SNP is defined as a single nucleotide substitution at one particular locus in the DNA sequence and this mutational event generates two alleles. To distinguish this from a point mutation, the frequency of the minor allele of a SNP has to be at least 1% in any population. Common SNPs are usually defined as those with minor allele frequency >5% and approximately 7 million of the SNPs in the human genome are common.79 Therefore, for single nucleotide substitutions, where their population frequencies are yet to be determined, strictly, they should be labeled as single nucleotide variations (SNVs) to minimize confusion.77 As a substantial fraction of entries in the dbSNP has not been validated in population-based studies, one has to bear in mind that not all the entries in the dbSNP are necessarily SNPs, as the name of database implies. As such, the several hundred thousand ‘new SNPs’ identified by whole genome resequencing studies27, 28, 29, 30, 31, 32, 33 should probably be considered as ‘new SNVs’ instead, until their population frequency information is available (Figure 3a). The distinction between SNPs and SNVs should be emphasized to avoid misleading.
(a) The proportion of new SNPs identified in whole genome resequencing studies. (b) The proportion of new indels identified in whole genome resequencing studies. *89,679 insertions up to 3bp, 124,024 deletions up to 11bp, 12,826 larger indels. 67% of small indels in dbSNP (i.e. insertions up to 3bp and deletions up to 11bp). **Approximately 0.4 million indels were identified and it was reported that about half of the indels are corroborated by entries in dbSNP
Single nucleotide polymorphisms are the most abundant type of genetic variation in the human genome in terms of their number. They occur at an interval of about one SNP in every kilobase of DNA sequence throughout the genome when the DNA sequences of any two individuals are compared. This is approximately equivalent to 3 million SNPs being carried by each individual genome. Therefore, the DNA sequence of any two genomes is estimated to be about 99.9% identical, and the 0.1% genetic variations that are mainly comprised of SNPs, are believed to be responsible for the phenotypic differences, such as physical traits (for example, height, hair and eye colors), disease susceptibility and drug responses, among individuals in populations. However, the finding of thousands of CNVs that collectively encompass hundreds of megabases of the genome8, 9, 10 and the numerous short indels that are identified by whole genome resequencing studies27, 28, 29, 30, 31, 32, 33 have thrown doubts to the estimation of ‘99.9% similarity’. The DNA sequences of individuals within and between populations are genetically more diverse and varied than previously thought.
Most of the SNPs are predicted to be neutral without functional effects and due to their abundance in the human genome; SNPs have become useful genetic markers in GWAS compared with other genetic variants such as microsatellites. In addition to the finding of a myriad of SNPs, some early reports have also documented the correlation patterns among the SNPs in parts of the human genome.80, 81, 82 However, no large-scale effort was undertaken to study the LD patterns in the whole genome until the initiation of the International HapMap Project. So far, a total of >3 million SNPs have been genotyped and validated in the Phase I and Phase II of the project.83, 84
The huge number of SNPs has also created a formidable task in genotyping because it is not technically feasible and cost effective to genotype several million of SNPs in a GWAS even with the latest genotyping technologies. Fortunately, SNPs are not completely independent of each other; instead they are correlated, as has been demonstrated by the International HapMap project. The existence of LD significantly reduces the number of SNPs that needs to be genotyped in a GWAS. The indirect association approach of GWAS is dependent on surrogate markers to locate disease variants through LD. As shown in the International HapMap Project and other published data, about half a million SNPs are already adequate to capture most of the SNPs that have been genotyped in the HapMap Project. However, the genome coverage of commercially genotyping arrays is population dependent. For example, Illumina HumanHap550 Beadchip, which contained ∼550 000 tagging SNPs, achieved genome coverage of 87 and 83% in CEU and CHB+JTP populations, respectively, but it was only 50% in YRI.85, 86, 87
The International HapMap Project has created a useful and valuable resource for GWAS. Furthermore, the availability of HapMap data has also driven the rapid developments in genotyping arrays, in which the data are used to guide the tagging SNPs selection. As the Phase I HapMap was completed in 2005, a number of genotyping arrays has been designed and introduced into the market, and the newer arrays have significantly improved in genome coverage and are also designed for CNVs detection, such as the Illumina Human 1 M Beadchip and Affymetrix 6.0 SNP Arrays.49 Hence, the International HapMap Project was a key and essential component in making the GWAS a feasible approach.
Around the turn of millennium, there were also some intense debates about the genetic architecture of complex diseases.88 It was polarized into two opposing models: the common-disease common-variant (CD/CV) versus multiple rare variant or common-disease rare-variant hypothesis.89 However, the CD/CV model formed the basis of the International HapMap Project; it was clearly shown in the Phase I HapMap, in which common SNPs have become the main focus. Over one million SNPs with minor allele frequency >5% were genotyped in 270 DNA samples collected from the four populations. Even in the Phase II HapMap, common SNPs remained as the focus; however, SNPs within minor allele frequency of 1–5% were also chosen to be genotyped.83, 84 As the HapMap data was used to develop commercial genotyping arrays, the SNP selection has been largely influenced by the CD/CV hypothesis. Therefore, the current GWAS are mainly interrogating the association of common SNPs with various complex diseases and traits.
The reason that the CD/CV model trumped the opposing model was also due to the technologies that were available at that time. Sanger dideoxynucleotide sequencing did not allow the survey of rarer SNPs or point mutations in the whole genome to be carried out efficiently. With the arrival of next generation sequencing technologies, whole genome sequencing is practical now, but still prohibitively expensive to be done in a large sample set for association studies. Instead, targeted sequencing of certain regions identified by GWAS, as well as exomes, is more feasible at the moment.90, 91 This approach has been advocated by genetics community as a temporary alternative to searching for rarer SNPs before we reach the goal of 1000 dollars per genome, enabling thousands of cases and controls to be sequenced. In contrast, the convenient high-throughput genotyping platforms have enabled an efficient interrogation of several hundred thousand to one million SNPs directly throughout the genome, which eventually captured almost all the SNPs in the International HapMap Project indirectly. Furthermore, it is more affordable to genotype (rather than to sequence) the whole genome of several thousand cases and controls for a statistically powerful association study.
Future
Copy number variations
The term CNV was first introduced in 2006, and it is generally defined as additions or deletions in the number of copies of a particular segment of DNA (larger than 1 kb in length) when compared with a reference genome sequence.35 The commonness of CNVs in the human genome was under-appreciated until the first reports in 2004. The findings have also stimulated a lot of enthusiasm and interest in the research of genetic diversity in the human populations and resulted in a series of effort to detect CNVs in different populations. The number of publications of CNVs studies has indeed increased greatly over the past few years.
In contrast to SNPs that have already been relatively well-cataloged in the dbSNP, and well-studied by the International HapMap Project, a lot more remains unclear for other types of genetic variations and to what extent they are present in the human genome. Although the ubiquity of CNVs in the human genome was reported several years ago, and many more have since been found, most of the studies used array-based detection methods that have relatively poor sensitivity compared with sequencing-based approaches.8, 9, 36, 92, 93, 94, 95 These array-based methods include bacterial artificial chromosome clones and oligonucleotides comparative genomic hybridization arrays and SNPs genotyping arrays. These methods are not sensitive enough to detect smaller sizes of CNVs that are less than 50 kb in size due to the limitations in array density or resolution.96 However, the number of smaller CNVs is estimated to be more abundant than the larger CNVs in the human genome.97
The poor sensitivity of array-based methods becomes apparent when their results are to be compared with the sequencing studies. The number of CNVs found in most of the array-based studies was in the range of tens to several hundred per genome on average, which is several fold lesser than the numbers that were reported in the whole genome resequencing studies. In each of the studies, several thousands of CNVs have been found;29, 30, 31, 32 for example, Ahn et al. identified 2920 deletions and 963 insertions in the Korean SJK genome. Although the improvements in SNPs density and inclusion of copy number probes in newer genotyping arrays, such as Illumina Human 1 M Beadchip and Affymetrix 6.0 SNP Arrays, have undoubtedly increased the performance of array-based methods to detect CNVs, the methods overall still suffer from poor sensitivity to detect CNVs smaller than 5–10 kb.9, 98 This was again clearly shown in the findings from whole genome resequencing studies. For example, a total of 2682 structural variations (dominated by deletions) were detected in the Han Chinese YH genome with a median length of about 0.5 kb.29 In contrast, the median length found by array-based methods was in the range of tens to hundreds of kilobases depending on the resolution of the arrays. This indicates that sequencing-based methods have much higher sensitivity to detect smaller CNVs. This also suggests that the overall larger number of CNVs found in whole genome resequencing studies was attributed to the better sensitivity in detecting more CNVs of smaller sizes. In addition, it is worthwhile noting that if the arbitrary cutoff of 1 kb is applied here, at least half of the reported CNVs by Wang et al.29 should be labeled as indels. This further illustrates the problems in classifying CNVs and indels into distinct categories.
Indels
In addition to CNVs, the several whole genome resequencing studies also identified hundreds of thousands of short indels.27, 28, 29, 30, 31, 32 The numbers reported in each study are not directly comparable, because the analyses, detection methods and criteria used are different between the studies. For example, for the two Korean genomes, the number of indels found in one study is twice another one. Ahn et al.32 identified 342 965 indels within a size range of −29 to +14 bp, whereas Kim et al.31 only found 170 202 indels within −29 to +5bp. Collectively these studies have uncovered plenty of short indels in the human genome. Moreover, the number of indels found is likely to represent only a fraction of the total number of indels in the human genome, because a rather narrow size range was defined in each of the studies. In summary, the several whole genome resequencing studies have further revealed the richness of genetic variations in the human genome and their numbers are more abundant than previously expected.
It is estimated that there are 1.6–2.5 million indels in human populations. However, no large-scale attempt was made to identify indels until 2006, in which a study identified 415 436 indels with about equal numbers of insertions and deletions.7 The sizes of these indels ranged from 1 bp to ∼10 kb (which span the ‘1 kb boundary’), thus suggesting that the dataset is actually a mixture of indels and CNVs. In addition, the study also found over 148 000 indels located within known genes and several thousands of them are found in the promoter regions and exons of genes. This means that these indels could potentially alter gene expression levels or affect protein structure or function. Similarly in the whole genome resequencing studies, several hundreds of indels were also found to overlap with coding sequences.28, 31 Despite some differences in the number of indels found in each study that overlapped with coding sequences, these studies have provided evidence to support their putative functional roles and also underscores the importance of investigating them in disease association studies. The discovery effort for indels is not keeping pace with that of SNPs, as indels have not been well cataloged in the dbSNP. This can be clearly shown from the proportion of new indels found in the whole genome resequencing studies; about 50% or more of the identified indels are not in dbSNP. In contrast, less than 30% of the SNPs identified in the studies are new (Figures 3a and b).
Though findings from whole genome resequencing studies have broadened our knowledge in human genetic variation, all of them only sequenced one individual genome, rendering them unable to investigate the population genetics of the identified genetic variants, such as frequencies and LD patterns. This piece of information is crucial and would be needed for future disease association studies. Moving towards this goal, and to accelerate the process of discovery of various genetic variations in the human genome, the 1000 Genomes Project was conceived and initiated in 2008. This project is currently on-going and the aim is to eventually sequence at least 1000 individual genomes from different populations worldwide. The ultimate goal is to build a useful resource of human genetic variations for future disease association studies. The availability of these resources and the genetic variations maps will certainly drive the technological development of new microarrays or other high-throughput methods to capture the non-SNP genetic variations in the near future, and it will bring another revolution to the genetic studies of complex diseases.
Copy neutral variations—inversions and translocations
The discovery of CNVs in the human genome of healthy populations has advanced rapidly over the last few years. However, an equivalent progress has not been seen for the detection of copy neutral variations; this is largely due to the lack of a powerful and efficient method for a genome-wide discovery of inversions and translocations. Unlike CNVs that can be studied by microarrays, the detection of copy neutral variations usually requires sequencing-based methods, and the high-throughput sequencing technologies that have only recently been made more accessible. In addition, inversions and translocations are technically more difficult to detect. A relatively slower progress in the studies of copy neutral variations is evident from the data entries recorded in the DGV, in which more than 29 000 CNVs and nearly 20 000 indels have been reported in the database, whereas less than a thousand inversions have been found, and no data is available for translocations in the DGV at the moment. However, one should be cautious with this interpretation because the numbers are not proportions. As the total number of CNVs, indels and inversions in the human genome is still unknown, therefore, the proportions of these genetic variations that have been discovered are also unknown. The data in the DGV are so far derived from the results of 35 studies using array-based and sequencing-based detection methods, and other approaches. In fact, more than this number of studies have been performed and published for CNVs detection in various populations; but not all their results have been cataloged in the DGV. As such, it is apparent that the entries in the database are still far from complete.
Most of the CNV data were generated by array-based methods (comparative genomic hybridization and SNP arrays), in which the signal intensity information is used to detect deletions and duplications, which relied on differences in signal intensities. As a result, these methods are unsuitable for detecting inversions and translocations (also known as balanced chromosomal rearrangements) because they do not lead to gain or loss of chromosomal or DNA segments. Rather, several different strategies and approaches have been taken to try to identify inversions in the human genome. For example, Feuk et al.99 discovered regions that are inverted between the chimpanzee and human genomes by performing comparative analysis of their DNA sequence assemblies. In the study, they identified about 1600 putative regions of inverted orientation in the genomes that covered >150 megabases of DNA sequence. The inverted regions are distributed throughout the genomes and span the sizes from 23 bp to 62 Mb in length. A number of inverted regions were also selected to be validated by using PCR and fluorescence in situ hybridization, and out of the 23 experimentally validated inversion regions, 3 of them were found to be polymorphic (>1%) in a panel of human samples, and were known as inversion polymorphisms.
However, a statistical method has also been developed to identify large inversion polymorphisms using high-density SNP genotyping data in which it is based on unusual LD patterns. The method was developed to detect chromosomal regions that are inverted in a majority of the chromosomes in a population with respect to the reference human genome sequence. Although this method has worked using the International HapMap Project data to detect inversion polymorphisms, it has not been widely used by other studies. In any case, this study was able to identify 176 inversions ranging from 200 kb to several megabases in length using the Phase I data. However, their results were not placed in the DGV.100 This, together with the study by Feuk et al. (2005)99, also provided some supporting evidence that a considerable portion of their detected inversions were flanked by highly homologous repeats or segmental duplications. This suggests that segmental duplications could be the favorite spots mediating the chromosomal rearrangements that generate inversions.
The breakthrough in the discovery of inversions was credited to the development of a sequencing-based method known as paired-end mapping, and the concurrent advances in next generation sequencing technologies. The paired-end mapping method also contributed greatly to the mapping of CNVs in the human genome. In the paired-end mapping method, both ends of the DNA fragments with known sizes would be sequenced and then aligned to the human reference genome. The principle of the paired-end mapping to detect various structural variations is simple in theory; it is based on the discordances in size or orientation of the DNA fragments that are to be aligned to the reference genome. When both ends of the DNA fragments that map to the reference genome show discordances in terms of size, this is an indication for deletion and insertion, whereas discordances in orientation suggests the presence of inversion.101
The power of this method to detect inversions was first demonstrated in the study by Tuzun et al.102 by sequencing the fosmid paired-end sequences. The study successfully identified 56 inversion breakpoints. The same strategy of fosmid clones sequencing was also used by Kidd et al.103 to detect structural variations in eight individual genomes, and a total of 224 inversions were also identified. However, this study is only the preliminary phase of a larger project that will eventually construct and sequence the fosmid clone libraries (∼40 kb inserts) prepared from the genomic DNA of 48 unrelated females, and bacterial artificial chromosome clone libraries (∼150 kb inserts) from 14 unrelated males in the International HapMap Project.104 Therefore, more inversions are expected to be discovered when the project is finished. The fosmid paired-end sequencing work of these studies was completed by traditional Sanger sequencing methods.
The first proof-of-concept study using next generation sequencing technologies in paired-end mapping to detect structural variations was published in 2007.105 In the study, libraries of 3-kb fragments for two female samples from the International HapMap Project were prepared and sequenced by Roche 454 sequencing, and they found 1297 structural variations, including 122 inversions. Using the same approach, hundreds of inversions were also uncovered by whole genome resequencing studies; for example, 91 and 415 inversions were detected in the African NA18507 genome and Korean SJK genome, respectively.32, 106 Although the progress in the discovery of inversions is moving at a slower pace than CNVs, there is already evidence to support their roles in human diseases.107, 108
Loss of heterozygosity and homozygosity
Copy neutral LOH defines a continuous stretch of DNA sequence without heterozygosity. It is different from a single copy deletion which could also lead to the absence of heterozygosity. More specifically, extended homozygosity is essentially copy neutral LOH, but it encompasses a large region of at least 1 Mb. Again, the distinction between the two categories is solely based on the length of DNA sequence without heterozygosity. Currently, there is no consensus on the definition of extended homozygosity. Previous studies have focused on homozygosity regions larger than 1 Mb, so the true level of homozygosity in the human genome could be underestimated.2, 109
The information regarding the extent of LOHs in the human genome is even less compared with indels and CNVs, but their potential impact on complex diseases could also be as much as other genetic variations. Although the biomedical significance of regions of homozygosity to complex diseases remains largely unexplored, some schizophrenia studies have already shown significant differences in homozygosity regions between cases and controls in a genome-wide study.22 More importantly is that the study has demonstrated the feasibility of using the homozygosity mapping approach to identify susceptibility loci and genes for complex diseases. This also highlights the need to further investigate and catalog the extent of LOH and homozygosity in the human genome. Similar to other genetic variations, LOHs definitely have the potential of being the genetic markers in future GWAS. Although homozygosity mapping has not been widely applied for most of the complex diseases, this approach is commonly used to interrogate the genetic basis of cancers to identify cancer-associated genes.110, 111
The ubiquity of homozygosity in the genomes of outbred populations has not been well documented. Previously, only a few studies reported an abundance of homozygosity in the human genome with frequent occurrence in genomic regions with extensive LD and low recombination rates.2, 109 Three widely discussed possibilities that led to the commonness of homozygosity are parental consanguinity, uniparental disomy and autozygosity. One previous study had demonstrated that the number of homozygosity regions increased markedly in the offspring of consanguineous marriages.112 However, this is unlikely in outbred populations in which parental consanguinity is rare.
Uniparental disomy can be divided into two types: uniparental isodisomy and uniparental heterodisomy. Only the former situation can cause homozygosity as the child inherits two identical copies of a chromosome segment from only one parent.113 This is also an unlikely explanation for the abundant homozygosity given that uniparental disomies are rare genetic abnormalities that can cause severe and rare genomic disorders, such as Prader–Willi Syndrome and Angelman Syndrome. This assumption is further supported by previous research that found extended homozygosity to be generally not due to genetic abnormalities.114 Using this reductionist approach, autozygosity seems to be the most likely process responsible for the commonness of homozygosity in the human genome. Autozygosity is a situation in which common ancestral haplotypes are inherited from both parents. Hence, extended homozygosity seems likely to have occurred as a result of common haplotypes, present in high frequencies in the population, which are passed on by chance from both parents to the child. This is further supported by previous findings of no excess apparent deviation from Mendelian transmission in extended homozygosity.109, 114
The future genetic variations map
The significance of the 1000 Genomes Project for future disease association studies is tremendous. Although SNPs have been widely used as the genetic markers in GWAS to search for disease variants, evidence has started accumulating to suggest that (common) SNPs alone are unlikely to account for all the heritable risk of complex diseases. Concurrently, the amount of data showing the associations of CNVs with complex diseases has been growing.19, 20, 21 Similarly, the importance of rare variants in complex diseases is also being recognized.56, 90, 115, 116 This implies that future disease association studies need to interrogate non-SNP and rare genetic variations as well, and for this to be feasible, a detailed catalog of human genetic variations is a prerequisite. Common SNPs are well documented in the dbSNP, but rarer SNPs (or lower frequency SNPs) are still under-represented in the database and the information of indels and structural variations is far from complete.
Unlike the whole genome resequencing studies of individual genomes, the 1000 Genomes Project is a large scale population-based sequencing study that enables studies of the population properties of genetic variations and their LD patterns. This information will be required to design next generation genotyping arrays to select surrogate markers that are not only able to tag for SNPs, but also to efficiently to capture indels and CNVs as well. This development will certainly widen the scope of genetic variations interrogated in GWAS. In fact, data have shown that CNVs could be tagged by SNPs through LD,9, 10, 117 but a detailed and in-depth investigation of their LD patterns can only be done when most of the SNPs, indels, CNVs and other genetic variations have been identified. In-depth studies of LD among different genetic variations is important, as the finding of the 20-kb deletion located upstream of the IRGM gene for Crohn's disease has demonstrated the efficiency of using SNPs as surrogate markers to identify non-SNP genetic variants.118 Other examples include the finding of a 45-kb deletion that is in perfect LD with BMI-associated SNPs in NEGR1.119
It is less likely that the number of indels and CNVs will reach several millions similar to the SNPs, but the total number of nucleotides encompassed by these genetic variations has already far exceeded that of the SNPs. Given their abundance in the human genome as found by the whole genome resequencing studies, their total nucleotide composition and functional impact on gene expression levels,11, 120, 121 they could potentially account for some or even a substantial portion of the inherited risk of complex diseases.
A comprehensive interrogation of genetic variations is essential because GWAS is an indirect approach to identify disease variants; therefore, its success is dependent on whether surrogate markers that are in strong LD with the disease variants are included in the studies. The LD information between SNPs, indels, CNVs and other genetic variations is valuable because it is more efficient to interrogate or capture indels and CNVs through LD by genotyping a number of SNPs, rather than by locating the probes within the copy number variable regions and detecting them through signal intensity differences. If the number or fraction of ‘untaggable’ indels and CNVs is considerable, then other high-throughput methods or microarrays can be developed to complement the content of next generation SNPs genotyping arrays. Besides driving the development of more efficient genotyping arrays to interrogate SNPs and non-SNP genetic variations, the data from the 1000 Genomes Project will also accelerate the fine mapping work in the regions identified by GWAS and improve the imputation powers because a much more complete reference set of genetic variations will be available for imputing.
The current status of GWAS
Genome-wide association study is a comprehensive and biologically agnostic approach to searching for unknown disease variants, and as demonstrated in more than 450 studies, this strategy has been very successful in identifying new genetic loci for various human complex traits. Most of the genes and loci that have been identified are not previously thought to be associated with their respective diseases.122, 123, 124, 125 More importantly, the GWAS findings have also provided new insights into the molecular pathways of complex diseases even when most of the disease causative variants remain to be discerned from the neighboring correlated markers. For example, the three new genes that have been linked to Crohn's disease: IL23R, ATG16L1 and IRGM have highlighted the importance of interleukin-23 receptor and autophagy pathways underlying the pathophysiology of this chronic inflammatory bowel disease.126, 127 Notably, GWAS have been making some significant advances in our understanding and knowledge of the genetic basis of human complex diseases compared with the pre-GWAS approaches (that is, the candidate gene association and linkage studies).
Most of the risk alleles that have been identified by GWAS are common (allele frequency >5%) and confer small effect sizes (odds ratio <1.5).17, 18 However, this observation is not really reflecting the true allelic frequency spectrum of complex diseases. This is because for any given sample size, association studies have higher statistical power to find associations with common SNPs. The other reason is that the rarer SNPs (allele frequency <5%) are not well-covered either directly or indirectly through LD by the markers in Illumina and Affymetrix genotyping arrays, so they remain unexplored for disease association. The design of GWAS and SNPs selection in commercial genotyping arrays have been largely driven by the CD/CV hypothesis.
Due to their small effect sizes, collectively the identified risk alleles only explain a small portion of the total inherited risk for the diseases. For example, all the type-2 diabetes risk alleles that are identified by GWAS cumulatively only account for ∼5% of the heritability, and similarly for other diseases, only a small proportion of the heritability was accounted for.128 The unexplained or missing heritability has been a major concern in the field, leading to the skepticism of the promise of GWAS to fully decipher the genetic basis of complex diseases. Nevertheless, it is noteworthy that GWAS have only interrogated a fraction of the total genetic variations in the human genome.
The genetic architecture of complex diseases remains elusive; it is unclear how much each type of genetic variation contributes to inherited risk and the relative proportion of rare versus common variants. If non-SNP genetic variants or rarer SNPs constitute most of the genetic component of complex diseases, then GWAS using the current genotyping arrays would be likely to miss them, simply because they are not covered directly by the genotyping arrays. How much they can be tagged through LD by the markers on the arrays still needs further investigation. Regardless, it is important to continue investigating other genetic variations to discover additional disease associated variants to explain the heritability.
Inadequate coverage of genetic variations in GWAS
All the GWAS rely heavily on the commercial genotyping arrays from Illumina and Affymetrix to comprehensively genotype several hundred-thousand of common SNPs. These genotyping arrays have near complete coverage of the >3 million SNPs genotyped by the International HapMap Project in CEU and CHB+JPT populations.85, 86, 87 The HapMap Project SNPs are either genotyped directly or tagged indirectly through LD with one or more SNPs on the arrays. Nevertheless, the HapMap SNPs are only a subset of the entire collection in the dbSNP, and currently there are more than 10 million SNPs cataloged in the database. More than half of the SNPs in dbSNP have not been studied for association with complex diseases directly and the number of these SNPs that are covered indirectly through LD by the genotyping arrays is unclear. It is noteworthy that the current GWAS only investigate a portion of the SNPs and the non-SNP genetic variations are likely not well studied for disease associations.
Furthermore, SNPs are not the only type of genetic variation in the human genome. Although the roles of non-SNP genetic variations in disease susceptibility remain largely unexplored, associations of CNVs with complex diseases such as schizophrenia, autism, autoimmune disorders, HIV infection and cancers have already been established from both candidate gene and genome-wide approaches.56, 115, 129, 130, 131, 132 The amount of evidence is expected to increase in the near future, when we have a better understanding of the characteristics of non-SNP genetic variations and a more comprehensive map of them constructed upon the completion of 1000 Genomes Project, and when more efficient and accurate methods are available to detect and study them. One major limitation of the current GWAS using the commercial genotyping arrays is that it covers only a portion of the total genetic variations, thus a substantial false negative rate is likely due to incomplete interrogation of all the genetic variations for disease association. For future studies, the focus should be directed on studying other genetic variations that have not yet been interrogated by the GWAS, such as tandem repeats, indels, inversions and CNVs, although it is highly dependent on the development of the technologies and methods of detection and analysis.
It is also obvious from the results of GWAS that the common SNPs are unable to account for the total inherited risk of a complex disease. However, it is not clear how much heritability can be attributed to rarer SNPs (<1–5%) at the time. Rarer SNPs are not well-covered by the GWAS or the genotyping arrays, as a result, they have not been intensively studied for disease association. Fortunately, the current genotyping arrays seem to work fine for detecting rare CNVs for diseases.56, 115 The evidence linking complex diseases and traits to multiple rare variants has also been growing; for example, for schizophrenia,56, 115 high-density lipoprotein cholesterol level133, 134 and type-1 diabetes.90 This implies that the rare variants (both SNP and non-SNP) should not be neglected in future studies. Sequencing approaches will improve their detection, and consequently offer a better understanding of the genetic architecture of complex diseases. The advances in sequencing technologies enable researchers to study a wider spectrum of genetic variants compared with genotyping methods.
Conclusions
The ultimate goal of GWAS is to correlate the genotype with disease phenotype, and to identify all the genetic variations that are associated with the diseases. To achieve this, most of the genetic variations in the human genome have to be first identified. It is essential to identify and validate all the genetic variations in the human genome in population-based studies, and catalog them properly in databases, so they can be used as the genetic markers for future disease association studies. Currently, we are moving towards these goals with the on-going 1000 Genomes Project, and only with the availability of a very detailed and near complete map of all genetic variations will it be feasible to perform a truly comprehensive search for the disease causing variants throughout the human genome.
References
Hastings, P. J., Lupski, J. R., Rosenberg, S. M. & Ira, G. Mechanisms of change in gene copy number. Nat. Rev. Genet. 10, 551–564 (2009).
Gibson, J., Morton, N. E. & Collins, A. Extended tracts of homozygosity in outbred human populations. Hum. Mol. Genet. 15, 789–795 (2006).
Botstein, D., White, R. L., Skolnick, M. & Davis, R. W. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am. J. Hum. Genet. 32, 314–331 (1980).
Weissenbach, J., Gyapay, G., Dib, C., Vignal, A., Morissette, J., Millasseau, P. et al. A second-generation linkage map of the human genome. Nature 359, 794–801 (1992).
Sachidanandam, R., Weissman, D., Schmidt, S. C., Kakol, J. M., Stein, L. D., Marth, G. et al. A map of human genome sequence variation containing 1.42 million single nucleotide polymorphisms. Nature 409, 928–933 (2001).
Haga, H., Yamada, R., Ohnishi, Y., Nakamura, Y. & Tanaka, T. Gene-based SNP discovery as part of the Japanese Millennium Genome Project: identification of 190 562 genetic variations in the human genome. Single-nucleotide polymorphism. J. Hum. Genet. 47, 605–610 (2002).
Mills, R. E., Luttig, C. T., Larkins, C. E., Beauchamp, A., Tsui, C., Pittard, W. S. et al. An initial map of insertion and deletion (INDEL) variation in the human genome. Genome Res. 16, 1182–1190 (2006).
Redon, R., Ishikawa, S., Fitch, K. R., Feuk, L., Perry, G. H., Andrews, T. D. et al. Global variation in copy number in the human genome. Nature 444, 444–454 (2006).
McCarroll, S. A., Kuruvilla, F. G., Korn, J. M., Cawley, S., Nemesh, J., Wysoker, A. et al. Integrated detection and population-genetic analysis of SNPs and copy number variation. Nat. Genet. 40, 1166–1174 (2008).
Conrad, D. F., Pinto, D., Redon, R., Feuk, L., Gokcumen, O., Zhang, Y. et al. Origins and functional impact of copy number variation in the human genome. Nature 464, 704–712 (2010).
Stranger, B. E., Forrest, M. S., Dunning, M., Ingle, C. E., Beazley, C., Thorne, N. et al. Relative impact of nucleotide and copy number variation on gene expression phenotypes. Science 315, 848–853 (2007).
Gilad, Y., Rifkin, S. A. & Pritchard, J. K. Revealing the architecture of gene regulation: the promise of eQTL studies. Trends Genet. 24, 408–415 (2008).
Fraser, H. B. & Xie, X. Common polymorphic transcript variation in human disease. Genome Res. 19, 567–575 (2009).
Usdin, K. The biological effects of simple tandem repeats: lessons from the repeat expansion diseases. Genome Res. 18, 1011–1019 (2008).
Haberman, Y., Amariglio, N., Rechavi, G. & Eisenberg, E. Trinucleotide repeats are prevalent among cancer-related genes. Trends Genet. 24, 14–18 (2008).
Hannan, A. J. Tandem repeat polymorphisms: modulators of disease susceptibility and candidates for ‘missing heritability’. Trends Genet. 26, 59–65 (2010).
Altshuler, D., Daly, M. J. & Lander, E. S. Genetic mapping in human disease. Science 322, 881–888 (2008).
Hindorff, L. A., Sethupathy, P., Junkins, H. A., Ramos, E. M., Mehta, J. P., Collins, F. S. et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc. Natl. Acad. Sci. USA 106, 9362–9367 (2009).
Wain, L. V., Armour, J. A. & Tobin, M. D. Genomic copy number variation, human health, and disease. Lancet 374, 340–350 (2009).
Stankiewicz, P. & Lupski, J. R. Structural variation in the human genome and its role in disease. Annu. Rev. Med. 61, 437–455 (2010).
Zhang, F., Gu, W., Hurles, M. E. & Lupski, J. R. Copy number variation in human health, disease, and evolution. Annu. Rev. Genomics Hum. Genet. 10, 451–481 (2009).
Lencz, T., Lambert, C., DeRosse, P., Burdick, K. E., Morgan, T. V., Kane, J. M. et al. Runs of homozygosity reveal highly penetrant recessive loci in schizophrenia. Proc. Natl. Acad. Sci. USA 104, 19942–19947 (2007).
Kaiser, J. A plan to capture human diversity in 1000 genomes. Science 319, 395 (2008).
Kuehn, B. M. 1000 Genomes Project promises closer look at variation in human genome. JAMA 300, 2715 (2008).
Schlötterer, C. The evolution of molecular markers—just a matter of fashion? Nat. Rev. Genet. 5, 63–69 (2004).
Nakamura, Y. DNA variations in human and medical genetics: 25 years of my experience. J. Hum. Genet. 54, 1–8 (2009).
Levy, S., Sutton, G., Ng, P. C., Feuk, L., Halpern, A. L., Walenz, B. P. et al. The diploid genome sequence of an individual human. PLoS Biol. 5, e254 (2007).
Wheeler, D. A., Srinivasan, M., Egholm, M., Shen, Y., Chen, L., McGuire, A. et al. The complete genome of an individual by massively parallel DNA sequencing. Nature 452, 872–876 (2008).
Wang, J., Wang, W., Li, R., Li, Y., Tian, G., Goodman, L. et al. The diploid genome sequence of an Asian individual. Nature 456, 60–65 (2008).
Bentley, D. R., Balasubramanian, S., Swerdlow, H. P., Smith, G. P., Milton, J., Brown, C. G. et al. Accurate whole human genome sequencing using reversible terminator chemistry. Nature 456, 53–59 (2008).
Kim, J. I., Ju, Y. S., Park, H., Kim, S., Lee, S., Yi, J. H. et al. A highly annotated whole-genome sequence of a Korean individual. Nature 460, 1011–1015 (2009).
Ahn, S. M., Kim, T. H., Lee, S., Kim, D., Ghang, H., Kim, D. S. et al. The first Korean genome sequence and analysis: full genome sequencing for a socio-ethnic group. Genome Res. 19, 1622–1629 (2009).
Pushkarev, D., Neff, N. F. & Quake, S. R. Single-molecule sequencing of an individual human genome. Nat. Biotechnol. 27, 847–852 (2009).
Feuk, L., Carson, A. R. & Scherer, S. W. Structural variation in the human genome. Nat. Rev. Genet. 7, 85–97 (2006).
Freeman, J. L., Perry, G. H., Feuk, L., Redon, R., McCarroll, S. A., Altshuler, D. M. et al. Copy number variation: new insights in genome diversity. Genome Res. 16, 949–961 (2006).
de Stahl, T. D., Sandgren, J., Piotrowski, A., Nord, H., Andersson, R., Menzel, U. et al. Profiling of copy number variations (CNVs) in healthy individuals from three ethnic groups using a human genome 32 K BAC-clone-based array. Hum. Mutat. 29, 398–408 (2008).
Tamaki, K. & Jeffreys, A. J. Human tandem repeat sequences in forensic DNA typing. Leg. Med. (Tokyo) 7, 244–250 (2005).
Petersdorf, E. W. HLA matching in allogeneic stem cell transplantation. Curr. Opin. Hematol. 11, 386–391 (2004).
Karas-Kuzelicki, N. & Mlinaric-Rascan, I. Individualization of thiopurine therapy: thiopurine S-methyltransferase and beyond. Pharmacogenomics 10, 1309–1322 (2009).
HUGO Pan-Asian SNP Consortium. Mapping human genetic diversity in Asia. Science 326, 1541–1545 (2009).
Feng, B. J., Huang, W., Shugart, Y. Y., Lee, M. K., Zhang, F., Xia, J. C. et al. Genome-wide scan for familial nasopharyngeal carcinoma reveals evidence of linkage to chromosome 4. Nat. Genet. 31, 395–399 (2002).
Bakker, S. C., van der Meulen, E. M., Buitelaar, J. K., Sandkuijl, L. A., Pauls, D. L., Monsuur, A. J. et al. A whole-genome scan in 164 Dutch sib pairs with attention-deficit/hyperactivity disorder: suggestive evidence for linkage on chromosomes 7p and 15q. Am. J. Hum. Genet. 72, 1251–1260 (2003).
Garner, C. P., Ding, Y. C., Steele, L., Book, L., Leiferman, K., Zone, J. J. et al. Genome-wide linkage analysis of 160 North American families with celiac disease. Genes Immun. 8, 108–114 (2007).
López, S., Buil, A., Ordoñez, J., Souto, J. C., Almasy, L., Lathrop, M. et al. Genome-wide linkage analysis for identifying quantitative trait loci involved in the regulation of lipoprotein a (Lpa) levels. Eur. J. Hum. Genet. 16, 1372–1379 (2008).
Wang, W. Y., Barratt, B. J., Clayton, D. G. & Todd, J. A. Genome-wide association studies: theoretical and practical concerns. Nat. Rev. Genet. 6, 109–118 (2005).
Hirschhorn, J. N. & Daly, M. J. Genome-wide association studies for common diseases and complex traits. Nat. Rev. Genet. 6, 95–108 (2005).
Matsuzaki, H., Dong, S., Loi, H., Di, X., Liu, G., Hubbell, E. et al. Genotyping over 100 000 SNPs on a pair of oligonucleotide arrays. Nat. Methods 1, 109–111 (2004).
Steemers, F. J., Chang, W., Lee, G., Barker, D. L., Shen, R. & Gunderson, K. L. Whole-genome genotyping with the single-base extension assay. Nat. Methods 3, 31–33 (2006).
Ragoussis, J. Genotyping technologies for genetic research. Annu. Rev. Genomics Hum. Genet. 10, 117–133 (2009).
International Human Genome Sequencing Consortium. Finishing the euchromatic sequence of the human genome. Nature 431, 931–945 (2004).
Sebat, J., Lakshmi, B., Troge, J., Alexander, J., Young, J., Lundin, P. et al. Large-scale copy number polymorphism in the human genome. Science 305, 525–528 (2004).
Iafrate, A. J., Feuk, L., Rivera, M. N., Listewnik, M. L., Donahoe, P. K., Qi, Y. et al. Detection of large-scale variation in the human genome. Nat. Genet. 36, 949–951 (2004).
Shendure, J. & Ji, H. Next-generation DNA sequencing. Nat. Biotechnol. 26, 1135–1145 (2008).
Mardis, E. R. Next-generation DNA sequencing methods. Annu. Rev. Genomics Hum. Genet. 9, 387–402 (2008).
Metzker, M. L. Sequencing technologies—the next generation. Nat. Rev. Genet. 11, 31–46 (2010).
International Schizophrenia Consortium. Rare chromosomal deletions and duplications increase risk of schizophrenia. Nature 455, 237–241 (2008).
Glessner, J. T., Wang, K., Cai, G., Korvatska, O., Kim, C. E., Wood, S. et al. Autism genome-wide copy number variation reveals ubiquitin and neuronal genes. Nature 459, 569–573 (2009).
Cook, E. H. Jr. & Scherer, S. W. Copy-number variations associated with neuropsychiatric conditions. Nature 455, 919–923 (2008).
Donis-Keller, H., Green, P., Helms, C., Cartinhour, S., Weiffenbach, B., Stephens, K. et al. A genetic linkage map of the human genome. Cell 51, 319–337 (1987).
De la Vega, F. M., Lazaruk, K. D., Rhodes, M. D. & Wenz, M. H. Assessment of two flexible and compatible SNP genotyping platforms: TaqMan SNP genotyping assays and the SNPlex genotyping system. Mutat. Res. 573, 111–135 (2005).
Olivier, M. The invader assay for SNP genotyping. Mutat. Res. 573, 103–110 (2005).
Ragoussis, J., Elvidge, G. P., Kaur, K. & Colella, S. Matrix-assisted laser desorption/ionisation, time-of-flight mass spectrometry in genomics research. PLoS Genet. 2, e100 (2006).
Hugot, J. P., Chamaillard, M., Zouali, H., Lesage, S., Cézard, J. P., Belaiche, J. et al. Association of NOD2 leucine-rich repeat variants with susceptibility to Crohn's disease. Nature 411, 599–603 (2001).
Bottini, N., Musumeci, L., Alonso, A., Rahmouni, S., Nika, K., Rostamkhani, M. et al. A functional variant of lymphoid tyrosine phosphatase is associated with type I diabetes. Nat. Genet. 36, 337–338 (2004).
Grant, S. F., Thorleifsson, G., Reynisdottir, I., Benediktsson, R., Manolescu, A., Sainz, J. et al. Variant of transcription factor 7-like 2 (TCF7L2) gene confers risk of type 2 diabetes. Nat. Genet. 38, 320–323 (2006).
Remmers, E. F., Plenge, R. M., Lee, A. T., Graham, R. R., Hom, G., Behrens, T. W. et al. STAT4 and the risk of rheumatoid arthritis and systemic lupus erythematosus. N. Engl. J. Med. 357, 977–986 (2007).
Hirschhorn, J. N. Genetic approaches to studying common diseases and complex traits. Pediatric Res. 57, 74–77 (2005).
Kemp, Z., Carvajal-Carmona, L., Spain, S., Barclay, E., Gorman, M., Martin, L. et al. Evidence for a colorectal cancer susceptibility locus on chromosome 3q21–q24 from a high-density SNP genome-wide linkage scan. Hum. Mol. Genet. 15, 2903–2910 (2006).
Sellick, G. S., Goldin, L. R., Wild, R. W., Slager, S. L., Ressenti, L., Strom, S. S. et al. A high-density SNP genome-wide linkage search of 206 families identifies susceptibility loci for chronic lymphocytic leukemia. Blood 110, 3326–3333 (2007).
Stanford, J. L., FitzGerald, L. M., McDonnell, S. K., Carlson, E. E., McIntosh, L. M., Deutsch, K. et al. Dense genome-wide SNP linkage scan in 301 hereditary prostate cancer families identifies multiple regions with suggestive evidence for linkage. Hum. Mol. Genet. 18, 1839–1848 (2009).
Gao, X., Martin, E. R., Liu, Y., Mayhew, G., Vance, J. M. & Scott, W. K. Genome-wide linkage screen in familial Parkinson disease identifies loci on chromosomes 3 and 18. Am. J. Hum. Genet. 84, 499–504 (2009).
Sellick, G. S., Longman, C., Tolmie, J., Newbury-Ecob, R., Geenhalgh, L., Hughes, S. et al. Genomewide linkage searches for Mendelian disease loci can be efficiently conducted using high-density SNP genotyping arrays. Nucleic Acids Res. 32, e164 (2004).
John, S., Shephard, N., Liu, G., Zeggini, E., Cao, M., Chen, W. et al. Whole-genome scan, in a complex disease, using 11 245 single-nucleotide polymorphisms: comparison with microsatellites. Am. J. Hum. Genet. 75, 54–64 (2004).
Yatsu, K., Mizuki, N., Hirawa, N., Oka, A., Itoh, N., Yamane, T. et al. High-resolution mapping for essential hypertension using microsatellite markers. Hypertension 49, 446–452 (2007).
Tamiya, G., Shinya, M., Imanishi, T., Ikuta, T., Makino, S., Okamoto, K. et al. Whole genome association study of rheumatoid arthritis using 27 039 microsatellites. Hum. Mol. Genet. 14, 2305–2321 (2005).
Kozlowski, P., de Mezer, M. & Krzyzosiak, W. J. Trinucleotide repeats in human genome and exome. Nucleic Acids Res. (2010) [e-pub ahead of print].
Day, I. N. dbSNP in the detail and copy number complexities. Hum. Mutat. 31, 2–4 (2010).
International HapMap Consortium. The International HapMap Project. Nature 426, 789–796 (2003).
Frazer, K. A., Murray, S. S., Schork, N. J. & Topol, E. J. Human genetic variation and its contribution to complex traits. Nat. Rev. Genet. 10, 241–251 (2009).
Daly, M. J., Rioux, J. D., Schaffner, S. F., Hudson, T. J. & Lander, E. S. High-resolution haplotype structure in the human genome. Nat. Genet. 29, 229–232 (2001).
Reich, D. E., Cargill, M., Bolk, S., Ireland, J., Sabeti, P. C., Richter, D. J. et al. Linkage disequilibrium in the human genome. Nature 411, 199–204 (2001).
Gabriel, S. B., Schaffner, S. F., Nguyen, H., Moore, J. M., Roy, J., Blumenstiel, B. et al. The structure of haplotype blocks in the human genome. Science 296, 2225–2229 (2002).
International HapMap Consortium. A haplotype map of the human genome. Nature 437, 1299–1320 (2005).
International HapMap Consortium Frazer, K. A.,, Ballinger, D. G., Cox, D. R., Hinds, D. A., Stuve, L. L., Gibbs, R. A. et al. A second generation human haplotype map of over 3.1 million SNPs. Nature 449, 851–861 (2007).
Barrett, J. C. & Cardon, L. R. Evaluating coverage of genome-wide association studies. Nat. Genet. 38, 659–662 (2006).
Eberle, M. A., Ng, P. C., Kuhn, K., Zhou, L., Peiffer, D. A., Galver, L. et al. Power to detect risk alleles using genome-wide tag SNP panels. PLoS Genet. 3, 1827–1837 (2007).
Li, M., Li, C. & Guan, W. Evaluation of coverage variation of SNP chips for genome-wide association studies. Eur. J. Hum. Genet. 16, 635–643 (2008).
Pritchard, J. K. Are rare variants responsible for susceptibility to complex diseases? Am. J. Hum. Genet. 69, 124–137 (2001).
Schork, N. J., Murray, S. S., Frazer, K. A. & Topol, E. J. Common vs rare allele hypotheses for complex diseases. Curr. Opin. Genet. Dev. 19, 212–219 (2009).
Nejentsev, S., Walker, N., Riches, D., Egholm, M. & Todd, J. A. Rare variants of IFIH1, a gene implicated in antiviral responses, protect against type 1 diabetes. Science 324, 387–389 (2009).
Ng, S. B., Turner, E. H., Robertson, P. D., Flygare, S. D., Bigham, A. W., Lee, C. et al. Targeted capture and massively parallel sequencing of 12 human exomes. Nature 461, 272–276 (2009).
Zogopoulos, G., Ha, K. C., Naqib, F., Moore, S., Kim, H., Montpetit, A. et al. Germ-line DNA copy number variation frequencies in a large North American population. Hum. Genet. 122, 345–353 (2007).
de Smith, A. J., Tsalenko, A., Sampas, N., Scheffer, A., Yamada, N. A., Tsang, P. et al. Array CGH analysis of copy number variation identifies 1284 new genes variant in healthy white males: implications for association studies of complex diseases. Hum. Mol. Genet. 16, 2783–2794 (2007).
Shaikh, T. H., Gai, X., Perin, J. C., Glessner, J. T., Xie, H., Murphy, K. et al. High-resolution mapping and analysis of copy number variations in the human genome: a data resource for clinical and research applications. Genome Res. 19, 1682–1690 (2009).
Itsara, A., Cooper, G. M., Baker, C., Girirajan, S., Li, J., Absher, D. et al. Population analysis of large copy number variants and hotspots of human genetic disease. Am. J. Hum. Genet. 84, 148–161 (2009).
Scherer, S. W., Lee, C., Birney, E., Altshuler, D. M., Eichler, E. E., Carter, N. P. et al. Challenges and standards in integrating surveys of structural variation. Nat. Genet. 39, S7–S15 (2007).
Estivill, X. & Armengol, L. Copy number variants and common disorders: filling the gaps and exploring complexity in genome-wide association studies. PLoS Genet. 3, 1787–1799 (2007).
Cooper, G. M., Zerr, T., Kidd, J. M., Eichler, E. E. & Nickerson, D. A. Systematic assessment of copy number variant detection via genome-wide SNP genotyping. Nat. Genet. 40, 1199–1203 (2008).
Feuk, L., MacDonald, J. R., Tang, T., Carson, A. R., Li, M., Rao, G. et al. Discovery of human inversion polymorphisms by comparative analysis of human and chimpanzee DNA sequence assemblies. PLoS Genet. 1, e56 (2005).
Bansal, V., Bashir, A. & Bafna, V. Evidence for large inversion polymorphisms in the human genome from HapMap data. Genome Res. 17, 219–230 (2007).
Medvedev, P., Stanciu, M. & Brudno, M. Computational methods for discovering structural variation with next-generation sequencing. Nat. Methods 6, S13–S20 (2009).
Tuzun, E., Sharp, A. J., Bailey, J. A., Kaul, R., Morrison, V. A., Pertz, L. M. et al. Fine-scale structural variation of the human genome. Nat. Genet. 37, 727–732 (2005).
Kidd, J. M., Cooper, G. M., Donahue, W. F., Hayden, H. S., Sampas, N., Graves, T. et al. Mapping and sequencing of structural variation from eight human genomes. Nature. 453, 56–64 (2008).
Human Genome Structural Variation Working Group Eichler, E. E., Nickerson, D. A., Altshuler, D., Bowcock, A. M., Brooks, L. D., Carter, N. P. et al. Completing the map of human genetic variation. Nature 447, 161–165 (2007).
Korbel, J. O., Urban, A. E., Affourtit, J. P., Godwin, B., Grubert, F., Simons, J. F. et al. Paired-end mapping reveals extensive structural variation in the human genome. Science 318, 420–426 (2007).
McKernan, K. J., Peckham, H. E., Costa, G. L., McLaughlin, S. F., Fu, Y., Tsung, E. F. et al. Sequence and structural variation in a human genome uncovered by short-read, massively parallel ligation sequencing using two-base encoding. Genome Res. 19, 1527–1541 (2009).
Feuk, L. Inversion variants in the human genome: role in disease and genome architecture. Genome Med. 2, 11 (2010).
Antonacci, F., Kidd, J. M., Marques-Bonet, T., Ventura, M., Siswara, P., Jiang, Z. et al. Characterization of six human disease-associated inversion polymorphisms. Hum. Mol. Genet. 18, 2555–2566 (2009).
Curtis, D., Vine, A. E. & Knight, J. Study of regions of extended homozygosity provides a powerful method to explore haplotype structure of human populations. Ann. Hum. Genet. 72, 261–278 (2008).
Assie, G., LaFramboise, T., Platzer, P. & Eng, C. Frequency of germline genomic homozygosity associated with cancer cases. JAMA 299, 1437–1445 (2008).
Bacolod, M. D., Schemmann, G. S., Wang, S., Shattock, R., Giardina, S. F., Zeng, Z. et al. The signatures of autozygosity among patients with colorectal cancer. Cancer Res. 68, 2610–2621 (2008).
Li, L. H., Ho, S. F., Chen, C. H., Wei, C. Y., Wong, W. C., Li, L. Y. et al. Long contiguous stretches of homozygosity in the human genome. Hum. Mutat. 27, 1115–1121 (2006).
Ting, J. C., Roberson, E. D., Miller, N. D., Lysholm-Bernacchi, A., Stephan, D. A., Capone, G. T. et al. Visualization of uniparental inheritance, Mendelian inconsistencies, deletions, and parent of origin effects in single nucleotide polymorphism trio data with SNPtrio. Hum. Mutat. 28, 1225–1235 (2007).
Curtis, D. Extended homozygosity is not usually due to cytogenetic abnormality. BMC Genet. 8, 67 (2007).
Walsh, T., McClellan, J. M., McCarthy, S. E., Addington, A. M., Pierce, S. B., Cooper, G. M. et al. Rare structural variants disrupt multiple genes in neurodevelopmental pathways in schizophrenia. Science 320, 539–543 (2008).
Bochukova, E. G., Huang, N., Keogh, J., Henning, E., Purmann, C., Blaszczyk, K. et al. Large, rare chromosomal deletions associated with severe early-onset obesity. Nature 463, 666–670.
Hinds, D. A., Kloek, A. P., Jen, M., Chen, X. & Frazer, K. A. Common deletions and SNPs are in linkage disequilibrium in the human genome. Nat. Genet. 38, 82–85 (2006).
McCarroll, S. A., Huett, A., Kuballa, P., Chilewski, S. D., Landry, A., Goyette, P. et al. Deletion polymorphism upstream of IRGM associated with altered IRGM expression and Crohn's disease. Nat. Genet. 40, 1107–1112 (2008).
Willer, C. J., Speliotes, E. K., Loos, R. J., Li, S., Lindgren, C. M., Heid, I. M. et al. Six new loci associated with body mass index highlight a neuronal influence on body weight regulation. Nat. Genet. 41, 25–34 (2009).
Henrichsen, C. N., Vinckenbosch, N., Zöllner, S., Chaignat, E., Pradervand, S., Schütz, F. et al. Segmental copy number variation shapes tissue transcriptomes. Nat. Genet. 41, 424–429 (2009).
Cahan, P., Li, Y., Izumi, M. & Graubert, T. A. The impact of copy number variation on local gene expression in mouse hematopoietic stem and progenitor cells. Nat. Genet. 41, 430–437 (2009).
Mohlke, K. L., Boehnke, M. & Abecasis, G. R. Metabolic and cardiovascular traits: an abundance of recently identified common genetic variants. Hum. Mol. Genet. 17, R102–R108 (2008).
Easton, D. F. & Eeles, R. A. Genome-wide association studies in cancer. Hum. Mol. Genet. 17, R109–R115 (2008).
Lettre, G. & Rioux, J. D. Autoimmune diseases: insights from genome-wide association studies. Hum. Mol. Genet. 17, R116–R121 (2008).
Ku, C. S., Loy, E. Y., Pawitan, Y. & Chia, K. S. The pursuit of genome-wide association studies: where are we now? J. Hum. Genet. 55, 195–206 (2010).
Cho, J. H. The genetics and immunopathogenesis of inflammatory bowel disease. Nat. Rev. Immunol. 8, 458–466 (2008).
Mathew, C. G. New links to the pathogenesis of Crohn disease provided by genome-wide association scans. Nat. Rev. Genet. 9, 9–14 (2008).
Maher, B. The case of the missing heritability. Nature 456, 18–21 (2008).
Sebat, J., Lakshmi, B., Malhotra, D., Troge, J., Lese-Martin, C., Walsh, T. et al. Strong association of de novo copy number mutations with autism. Science 316, 445–449 (2007).
Hollox, E. J., Huffmeier, U., Zeeuwen, P. L., Palla, R., Lascorz, J., Rodijk-Olthuis, D. et al. Psoriasis is associated with increased beta-defensin genomic copy number. Nat. Genet. 40, 23–25 (2008).
Gonzalez, E., Kulkarni, H., Bolivar, H., Mangano, A., Sanchez, R., Catano, G. et al. The influence of CCL3L1 gene-containing segmental duplications on HIV-1/AIDS susceptibility. Science 307, 1434–1440 (2005).
Shlien, A. & Malkin, D. Copy number variations and cancer susceptibility. Curr. Opin. Oncol. 22, 55–63 (2010).
Cohen, J. C., Kiss, R. S., Pertsemlidis, A., Marcel, Y. L., McPherson, R. & Hobbs, H. H. Multiple rare alleles contribute to low plasma levels of HDL cholesterol. Science 305, 869–872 (2004).
Romeo, S., Pennacchio, L. A., Fu, Y., Boerwinkle, E., Tybjaerg-Hansen, A., Hobbs, H. H. et al. Population-based resequencing of ANGPTL4 uncovers variations that reduce triglycerides and increase HDL. Nat. Genet. 39, 513–516 (2007).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Ku, C., Loy, E., Salim, A. et al. The discovery of human genetic variations and their use as disease markers: past, present and future. J Hum Genet 55, 403–415 (2010). https://doi.org/10.1038/jhg.2010.55
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2010.55
Keywords
This article is cited by
-
Artificial Intelligence Techniques for the effective diagnosis of Alzheimer’s Disease: A Review
Multimedia Tools and Applications (2023)
-
Establishment and characterization of patient-derived xenograft of a rare pediatric anaplastic pleomorphic xanthoastrocytoma (PXA) bearing a CDC42SE2-BRAF fusion
Scientific Reports (2023)
-
Leveraging Mann–Whitney U test on large-scale genetic variation data for analysing malaria genetic markers
Malaria Journal (2022)
-
New neural network classification method for individuals ancestry prediction from SNPs data
BioData Mining (2021)
-
Genomic medicine on the frontier of precision medicine
Journal of Diabetes & Metabolic Disorders (2021)