Abstract
The myosin VIIA (MYO7A) gene encodes a protein classified as an unconventional myosin. Mutations within MYO7A can lead to both syndromic and non-syndromic hearing impairment in humans. Among different mutations reported in MYO7A, only five led to non-syndromic sensorineural deafness autosomal dominant type 11 (DFNA11). Here, we present the clinical, genetic and molecular characteristics of two large Chinese DFNA11 families with either high- or low-frequency hearing loss. Affected individuals of family DX-J033 have a sloping audiogram at young ages with high frequency are most affected. With increasing age, all test frequencies are affected. Affected members of family HB-S037 present with an ascending audiogram affecting low frequencies at young ages, and then all frequencies are involved with increasing age. Genome-wide linkage analysis mapped the disease loci within the DFNA11 interval in both families. DNA sequencing of MYO7A revealed two novel nucleotide variations, c.652G>A (p.D218N) and c.2011G>A (p.G671S), in the two families. It is for the first time that the mutations identified in MYO7A in the present study are being implicated in DFNA11 in a Chinese population. For the first time, we tested electrocochleography (ECochG) in a DFNA11 family with low-frequency hearing loss. We speculate that the low-frequency sensorineural hearing loss in this DFNA11 family was not associated with endolymphatic hydrops.
Similar content being viewed by others
Introduction
Hearing impairment is a common clinical finding with both genetic and environmental origins.1 MYO7A located at 11q13.5 encodes an unconventional myosin. Mutations in MYO7A are responsible for Usher syndrome type 1B,2, 3 recessively inherited atypical Usher syndrome,4, 5 autosomal recessive (DFNB2) and dominant (DFNA11) non-syndromic sensorineural deafness.6, 7, 8 MYO7A has 49 exons, of which 48 are coding. The predicted protein consists of a polypeptide of 2215 amino acids. MYO7A is expressed in the inner ear, retina, testis, lung and kidney.9 In inner ear hair cells, MYO7A is expressed in stereocilia bundles, cuticular plate and cell body. MYO7A is also detected in both type I and type II hair cells of the semicircular canals and utricle.9, 10 A total of 340 different mutations and 248 protein variants in MYO7A have been reported to date (http://www.umd.be/MYO7A/). Only five MYO7A mutations leading to DFNA11 have been reported: p.delA886-K887-K888 in a Japanese pedigree,7, 11 p.G772R in an American pedigree,12 p.N458I in a Dutch pedigree,13 p.R853C in a German pedigree14 and p.A230V in an Italian pedigree.15 Here, we present the clinical, genetic and molecular characteristics of two large Chinese families with either non-syndromic high- or low-frequency DFNA11. These represent the sixth and seventh DFNA11 families reported to date.
Materials and methods
Families and clinical evaluations
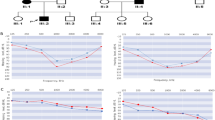
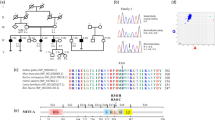
We have ascertained two large multigeneration Chinese families (DX-J033 and HB-S037) with autosomal dominant late-onset progressive non-syndromic sensorineural hearing loss (SNHL). The families DX-J033 and HB-S037 span five generations (Figure 1). Written informed consent was obtained from all participating individuals in accordance with the ethics committee of the Chinese PLA General Hospital. Medical history was obtained by using a questionnaire regarding the following aspects of this condition: subjective degree of hearing loss, age at onset, evolution, symmetry of the hearing impairment, use of hearing aids, presence of tinnitus, pressure in the ears or vertigo, medication, noise exposure, pathological changes in the ear and other relevant clinical manifestations. Otoscopy, physical examination and pure tone audiometry (at frequencies from 250 to 8000 Hz) were performed to identify the phenotype. Immittance testing was applied to evaluate middle-ear pressures, ear canal volumes and tympanic membrane mobility. Electrocochleography (ECochG) was carried out to understand the pathology of the low-frequency hearing loss of family HB-S037. In the ECochG test, short tone bursts with a frequency of 1 kHz were used to evoke cochlear microphonics and alternating polarity clicks were used to measure summating potentials (SPs) and cochlear nerve action potential (APs). Cochlear microphonics and AP detection thresholds, and the SP/AP ratios at the intensity of 90-dB nHL were used as indicators. Computed tomography scan analysis of the probands of two families has been conducted to rule out inner-ear malformations.
Linkage analysis
Blood samples were drawn from 29 and 27 participants from families DX-J033 and HB-S037, respectively. Genomic DNA was extracted from peripheral blood using the Genomic DNA isolation kit (HuaShun, Shanghai, China). A genome-wide scan was performed using 382 fluorescent-labeled microsatellite markers from the ABI Prism Linkage Mapping Set Version 2 (Applied Biosystems, Foster City, CA, USA). Markers for the refinement of the critical interval were taken from the Marshfield chromosome 11 map (http://research.marshfieldclinic.org/genetics). Alleles were assigned using Genescan and Genotyper Software (Applied Biosystems). Linkage analysis was performed using the LINKAGE 5.1 software package (http://linkage.rockefeller.edu/soft/linkage/). The two-point logarithm of odds (LOD) score between the deafness locus and each marker was calculated under a fully penetrant autosomal dominant mode of inheritance, setting the disease allele frequency to 0.0001. The meiotic recombination frequencies were considered to be equal for men and women. Haplotypes were constructed on the basis of known marker orders.
Mutational analysis of MYO7A
Primers were designed to amplify all exons and flanking intronic sequences of the MYO7A gene (NM_000260). All 49 exons and exon–intron boundaries of the MYO7A gene were amplified using standard PCR conditions and were completely sequenced. The primer sequences and precise PCR conditions are available from the authors on request. Bi-directional sequencing was carried out using genomic DNA extracted from two affected and one unaffected member of each family, and was performed using the ABI PRISM Big Dye Terminator Sequencing kit (Applied Biosystems) on a 3100 ABI DNA-sequencer (Applied Biosystems). DNA samples from 100 unrelated Chinese control individuals were also analyzed.
Molecular modeling
To analyze the structural impact of D218N and G671S substitutions in MYO7A, we used the DeepView (Swiss- PdbViewer) program version 3.7 (http://www.expasy.org/spdbv).16, 17 The sequences of wild-type and mutant MYO7A were submitted to the protein homology-modeling server and the 3-D structures were modeled automatedly using PDB ID:1w7jA as template. The images were shown and rendered with PYMOL software (http://www.pymol.org/).
Results
Families and clinical evaluations
In 11 affected subjects from family DX-J033 and 9 from HB-S037 family, hearing impairment was symmetric, with severity varying from mild to profound on their audiograms (Table 1). Affected family members in DX-J033 present with a sloping audiogram at the age of onset at high frequencies, whereas in family HB-S037, affected individuals have an ascending audiogram at age of onset at low frequencies. The progression of hearing loss was modest in the families, affecting all frequencies, with increasing age, resulting in a flat or downward sloping audiogram contour in family DX-J033 and a flat or ascending audiogram contour in family HB-S037. Audiograms of some affected members of families DX-J033 and HB-S037 are shown in Figures 1a and b, respectively. Vestibular symptoms were not reported by any of the affected subjects. All the affected members at the onset of hearing loss have high-frequency tinnitus. Other audiological evaluation of both DX-J033 and HB-S037 family members demonstrated normal immittance testing and bone conduction values equal to the air conduction measurements, thereby suggesting sensorineural hearing impairment. The EcochG test generated an SP/AP ratio within the normal range for two affected individuals, IV-10 and V-1, of family HB-S037 (data not shown). Comprehensive family medical histories and clinical examination of these individuals showed no other clinical abnormalities, including diabetes, cardiovascular diseases, visual problems and neurological disorders. Computer tomography scan analysis of the probands of DX-J033 and HB-S037 families ruled out inner-ear malformations.
Linkage analysis
For family HB-S037, 23 family members from the last three generations were considered informative, and were selected for linkage study. Genomic scanning at 10 cM intervals identified a region on the long arm of chromosome 11 that yielded an LOD score of 3.67 at D11S1314, with exclusion of the remainder of the genome. We then tested additional markers spanning the region, and a maximum two-point LOD score of 4.18 at θ=0 was obtained for markers D11S4081, D11S911 and D11S4166, located within 1 cM of the MYO7A gene. For family DX-J033, evidence of linkage was found for marker D11S4166 with a maximum two-point LOD score of 4.4 at θ=0 on chromosome 11. The family was then genotyped for five additional markers flanking D11S4166 to confirm linkage and for fine mapping of the genetic interval. A maximum two-point LOD score of 4.5 at θ=0 was obtained for marker D11S911. Haplotype analysis placed the locus within a 10 cM genetic interval flanked by markers D11S1314 and D11S901, overlapping with that of MYO7A (Figure 2).
Mutation analysis
We sequenced all coding exons and about 100 bp flanking intronic sequence of MYO7A in two affected and one unaffected member of families DX-J033 and HB-S037. A total of two novel missense mutations were detected in a heterozygous state. The first novel p.D218N mutation identified in family DX-J033 is caused by a G to A transition at nucleotide 652 in exon 7, resulting in the replacement of aspartic acid, a polar/small/charged amino acid, with an uncharged polar amino acid, asparagine, in the motor domain of MYO7A (Figures 3a and c). In family HB-S037, we identified another unreported sequence change, p.G671S, due to a G>A transition at nucleotide position 2011 in exon 17. This substitution results in a replacement of glycine which has a small/nonpolar neutral residue, with serine, a polar/uncharged amino acid. This change occurs within the motor domain of the MYO7A protein (Figures 3a and c).
The MYO7A mutations identified in family DX-J033 and HB-S037. (a) DNA sequence chromatograms showing heterozygous c.652 G>A (p.D218N) and c.2011G>A (p.G671S) in affected individuals compared with homozygous D218 and G671 in controls. (b) Protein alignment showing conservation of residues p.D218N and p.G671S in myosin across nine species: Danio rerio, Gallus gallus, Rattus, Mus musculus, Homo sapiens, Macaca, Sus scrofa, Bos taurus and Drosophila melanogaster. (c) Schematic representation of MYO7A functional regions indicates the location of eight DFNA11 mutations. p.D218N and p.G671S substitutions in DX-J033 and HB-S037 pedigrees are shown in bold. The head region contains ATP and actin binding sites. The tail domain contains four notable regions: (1) IQs represent five light-chain binding repeats; (2) coil indicates a coiled-coil domain that may be involved in dimerization; (3) MyTH4 indicates myosin tail homology-4 domains that are regions conserved between myosins; and (4) talin represents talin-like homology domains that are predicted to bind actin.
Comparison of myosins showed that the two amino acids, D218 and G671 (those with sequence changes p.D218N and p.G671S in our study), remained perfectly conserved among different species and interspecies (Figure 3b). Neither of these mutations in MYO7A was detected in unaffected members of families, nor was either mutation detected in the DNA from 100 unrelated Chinese controls. Table 2 lists the eight polymorphisms identified in the present study, including two synonymous and two non-synonymous substitutions, and four intronic nucleotide substitutions. Four of these have been previously reported as non-pathogenic (http://www.umd.be/MYO7A/).
We have excluded the four novel changes (c.4956G>A, IVS2-19A>T, IVS41-175A>G and IVS42-12T>C) as causative mutations for deafness because they do not create putative cryptic splice sites and/or did not segregate with deafness in the family. We used DeepView to predict the likely functional impact of D218N and G671S on MYO7A. Residue 218 belongs to a β-strand. In close proximity is A230 in a neighboring strand, and I418, belonging to a loop. A missense A230V mutation in the MYO7A protein has been reported previously in a family with an autosomal dominant form of non-syndromic SNHL with vestibular involvement. Affected individuals show a bilateral moderate to profound SNHL at all frequencies.15
The D218N mutation identified in a Chinese DFNA11 family in the present study was implicated in high-frequency hearing loss. A substitution of aspartic acid, a negative charge residue, to neutral asparagine at 218, may have led to loss of hydrogen bonds, thus altering the interchain interactions between the neighboring β-strand, containing residues A230, K231 and D218 (Figures 4a and b). The MYO7A G671 residue is in the region of the myosin head converter domain. The N-terminal, globular domain of myosin interacts with actin filaments, using energy from ATP hydrolysis to generate force and movement. High-resolution structural data show the myosin motor to be comprised of a central core. Extensions to this central core form the actin-binding site and the relay, converter and lever-arm regions.18 Interaction of a portion of the myosin head referred to as the relay loop with the converter domain occurs in all myosin crystal structures examined. The relay loop mediates conformational information between the converter domain, the nucleotide-binding site and the actin-binding site. The relay loop and converter domain are coupled to each other in part via a conserved hydrophobic pocket formed by residues F727 from the converter domain with Y477 and I482 from the relay loop. The model shown in Figures 4c and d demonstrates that the substituted serine side chain projects into the conserved hydrophobic pocket, generating steric hindrance with neighboring amino acid residue Y477.
The functional impact of D218N and G671S mutants on MYO7A predicted by molecular modeling (PDB ID:1w7jA). (a) Residues K231 and wild-type D218, belonging to neighboring beta strands, are shown (b) The area comprising K231 and D218N is represented. (c) Ribbon diagram with the wild-type G671 residue. Only residue numbers for MYO7A are shown. (d) Ribbon diagram containing the G671S mutation. Side-chain interactions resulting in steric hindrance are depicted.
Discussion
In this study, we report the sixth and seventh family in which high- or low-frequency non-syndromic autosomal dominant hearing impairment segregates with the DFNA11 locus. By linkage analysis, we mapped the hearing loss locus in both families on the long arm of chromosome 11 at 11q13.4–q14.1, within a genetic interval that contains the MYO7A gene. By sequence analysis, we identified two novel sequence variations (p.D218N and p.G671S), which are located in the motor domain of the MYO7A protein. The replacement of aspartic acid with asparagine at 218 may have led to alteration of the interchain interactions between the neighboring β-strand containing residues A230, K231 and D218. Glycine to serine substitution at position 671 may disrupt the conserved hydrophobic pocket formed by amino acid side chains in the converter domain and relay loop of the myosin head. The converter region, connecting the lever arm to the motor domain, is thought to be responsible for allowing the lever arm to rotate relatively to the catalytic domain. The rigid relay loop, which provides a link between the active site and the converter domain, as well as actin-binding regions, is believed to be critically involved in the conduction and amplification of structural changes at myosin’s active site to both the lever arm and the actin-binding interface.19 Thus, disruption of the interface between the myosin relay loop and the converter may have perturbed the communication pathway. Further confirmation of the causative nature of the MYO7A G671S mutation would be provided by studies demonstrating functional defects in motility, ATPase activity, lever-arm swing and ability to translocate actin filaments.
The genetic and clinical characteristics of all identified MYO7A mutations associated with DFNA11 are summarized in Table 3. To date, five of seven mutated alleles identified in the DFNA11 mutations are located in the motor domain of the MYO7A gene. There appears to be no obvious correlation between the location of a missense mutation in the MYO7A protein and the resulting phenotype.
In this study, the affected young members of family HB-S037 showed low-frequency SNHL and no vestibular dysfunction. To our knowledge, the low-frequency SNHL is thought to be a specific audiometric configuration indicative of endolymphatic hydrops in DFNA1 families with DIAPH1 mutations and in DFNA6/14/38 families with WFS1 mutations.20, 21 An enlarged SP/AP ratio in ECochG reflects the specific abnormalities associated with endolymphatic hydrops. To understand the pathology of the low-frequency hearing loss of family HB-S037, we tested ECochG for two affected young members of family HB-S037. On the basis of the fact that both of them exhibited bilaterally normal SP/AP ratios, we speculated that the low-frequency SNHL observed in the DFNA11 family was not associated with endolymphatic hydrops.
In summary, here we report clinical, genetic and molecular characteristics of two large Chinese families with DFNA11. For the first time, we tested ECochG in a DFNA11 family with low-frequency hearing loss and suggest that endolymphatic hydrops were not involved in the pathology of low-frequency SNHL in DFNA11. DNA sequencing of MYO7A revealed two novel nucleotide variations, c.652G>A (p.D218N) and c. 2011G>A (p.G671S). These changes are therefore considered to underlie the hearing impairment in the Chinese families DX-J033 and HB-S037. These families carry the sixth to seventh DFNA11 mutations, respectively, described in the literature. These are, however, the first two mutations reported in the Chinese population. The identification of additional disease-causing mutations in DFNA11 further confirms the crucial role of the MYO7A protein in auditory function.
References
Morton, N. E. Genetic epidemiology of hearing impairment. Ann. NY. Acad. Sci. 630, 16–31 (1991).
Weil, D., Blanchard, S., Kaplan, J., Guilford, P., Gibson, F., Walsh, J. et al. Defective myosin VIIA gene responsible for Usher syndrome type 1B. Nat. 374, 60–61 (1995).
Weston, M. D., Kelley, P. M., Overbeck, L. D., Wagenaar, M., Orten, D. J., Hasson, T. et al. Myosin VIIA mutation screening in 189 Usher syndrome type 1 patients. Am. J. Hum. Genet . 59, 1074–1083 (1996).
Liu, X. Z., Hope, C., Walsh, J., Newton, V., Ke, X. M., Liang, C. Y. et al. Mutations in the myosin VIIA gene cause a wide phenotypic spectrum, including atypical Usher syndrome. Am. J. Hum. Genet. 63, 909–912 (1998).
Zina, Z. B., Masmoudi, S., Ayadi, H., Chaker, F., Ghorbel, A. M., Drira, M. et al. From DFNB2 to Usher syndrome: variable expressivity of the same disease. Am. J. Med .Genet . 101, 181–183 (2001).
Liu, X. Z., Walsh, J., Mburu, P., Kendrick-Jones, J., Cope, M. J., Steel, K. P. et al. Mutations in the myosin VIIA gene cause non-syndromic recessive deafness. Nat. Genet. 16, 188–190 (1997a).
Liu, X. Z., Walsh, J., Tamagawa, Y., Kitamura, K., Nishizawa, M., Steel, K. P. et al. Autosomal dominant non-syndromic deafness caused by a mutation in the myosin VIIA gene. Nat. Genet. 17, 268–269 (1997b).
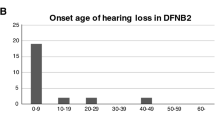
Liu, X. Z. The clinical presentation of DFNB2. Adv. Otorhinolaryngol. 61, 120–123 (2002).
Hasson, T., Heintzelman, M. B., Santos-Sacchi, J., Corey, D. P. & Mooseker, M. S. Expression in cochlea and retina of myosin VIIa, the gene product defective in Usher syndrome type 1B. Proc. Natl . Acad. Sci. USA. 92, 9815–9819 (1995).
Hasson, T., Gillespie, P. G., Garcia, J. A., MacDonald, R. B., Zhao, Y., Yee, A. G. et al. Unconventional myosins in inner-ear sensory epithelia. J. Cell. Biol. 137, 1287–1307 (1997).
Tamagawa, Y., Kitamura, K., Ishida, T., Nishizawa, M., Liu, X. Z., Walsh, J. et al. Sensorineural hearing impairment non-syndromic, dominant DFNA11. Adv. Otorhinolaryngol. 56, 103–106 (2000).
Street, V. A., Kallman, J. C. & Kiemele, K. L. Modifier controls severity of a novel dominant low-frequency MyosinVIIA (MYO7A) auditory mutation. J. Med. Genet. 41, e62 (2004).
Luijendijk, M. W., Van Wijk, E., Bischoff, A. M., Krieger, E., Huygen, P. L., Pennings, R. J. et al. Identification and molecular modelling of a mutation in the motor head domain of myosin VIIA in a family with autosomal dominant hearing impairment (DFNA11). Hum. Genet. 115, 149–156 (2004).
Bolz, H., Bolz, S. S., Schade, G., Kothe, C., Mohrmann, G., Hess, M. et al. Impaired calmodulin binding of myosin-7A causes autosomal dominant hearing loss (DFNA11). Hum. Mutat. 24, 274–275 (2004).
Di Leva, F., D’Adamo, P., Cubellis, M. V., D’Eustacchio, A., Errichiello, M., Saulino, C. et al. Identification of a novel mutation in the myosin VIIA motor domain in a family with autosomal dominant hearing loss (DFNA11). Audiol. Neurootol. 11, 157–164 (2006).
Schwede, T ., Kopp, J ., Guex, N . & Peitsch, M. C. SWISS-MODEL: an automated protein homology- modeling server. Nucleic. Acids. Research. 31, 3381–3385 (2003).
Guex, N . & Peitsch, M. C. SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modelling. Electrophoresis. 18, 2714–2723 (1997).
Geeves, M. A. & Holmes, K. C. Structural mechanism of muscle contraction. Annu. Rev. Biochem. 68, 687–728 (1999).
Houdusse, A. & Sweeney, H. L. Myosin motors: missing structures and hidden springs. Curr. Opin. Struct. Biol. 11, 182–194 (2001).
Bramhall, N. F., Kallman, J. C., Verrall, A. M. & Street, V. A. A novel WFS1 mutation in a family with dominant low frequency sensorineural hearing loss with normal VEMP and EcochG findings. BMC. Med. Genet. 9, 48 (2008).
Fujikawa, T., Noguchi, Y., Ito, T., Takahashi, M. & Kitamura, K. Additional heterozygous 2507A>C mutation of WFS1 in progressive hearing loss at lower frequencies. Laryngoscope. 120, 166–171 (2010).
Acknowledgements
We sincerely thank all family members for their participation and support in this study. These investigations were supported by grants from the National High Technology Research and Development Program of China (863 Program) to Huijun Yuan (2007AA02E466), grants from the Key Project of National Natural Science Foundation of China to Huijun Yuan (81030017) and by funds from the National Natural Science Foundation of China to Xuezhong Liu (30528025).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Sun, Y., Chen, J., Sun, H. et al. Novel missense mutations in MYO7A underlying postlingual high- or low-frequency non-syndromic hearing impairment in two large families from China. J Hum Genet 56, 64–70 (2011). https://doi.org/10.1038/jhg.2010.147
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2010.147
Keywords
This article is cited by
-
First glance at the molecular etiology of hearing loss in French-Canadian families from Saguenay-Lac-Saint-Jean’s founder population
Human Genetics (2022)
-
An effective screening strategy for deafness in combination with a next-generation sequencing platform: a consecutive analysis
Journal of Human Genetics (2016)
-
Genetics of vestibular disorders: pathophysiological insights
Journal of Neurology (2016)
-
Analysis of two Arab families reveals additional support for a DFNB2 nonsyndromic phenotype of MYO7A
Molecular Biology Reports (2014)