Abstract
Hepatic lipase (HL) plays a major role in the regulation of plasma lipids. Several groups seeking to find association between the gene encoding HL (LIPC) and plasma concentrations of high-density lipoprotein cholesterol (HDLc) using various methods and populations have reported conflicting results. We have approached the problem of demonstrating a relationship between the LIPC locus and HDLc by means of haplotype association using four single nucleotide polymorphisms (SNPs) (rs12594375G/A, rs8023503C/T, rs4775047C/T, and rs11634134T/A) located in intron 1 of the LIPC gene in two independent Japanese populations consisting of 2,970 and 1,638 individuals, respectively. Significant association between hyperalphalipoproteinemia and a specific haplotype in this intron was detected in both populations. When HDLc levels among the three haplotypic categories were analyzed [haplotype rs8023503C/rs12594375G (haplotype-1; H1) homozygotes (H1H1), haplotype rs8023503T/rs12594375A (haplotype-2; H2) homozygotes (H2H2), and heterozygotes (H1H2)], HDLc levels were lowest among H1H1 [mean ± standard error (SE) = 58.4 ± 0.4 mg/dl], highest among H2H2 (62.5 ± 0.8 mg/dl), and intermediate among H1H2 (59.2 ± 0.4 mg/dl) (P = 0.00011), indicating that H2 haplotype elevates plasma HDLc levels. This association was validated in the second population (n = 1,638) (P = 0.00070). The results provide convincing evidence that the LIPC locus influences HDL metabolism.
Similar content being viewed by others
Introduction
Genetic mechanisms remain obscure for the majority of human hyperalphalipoproteinemias that involve elevated high-density lipoprotein cholesterol (HDLc) levels in plasma (Glueck et al. 1975, 1977). Rare Mendelian monogenic conditions do exist, however. One of them is an autosomal recessive trait familial deficiency of hepatic lipase (HL) (Hegele et al. 1991a, b). Such patients present with elevated plasma HDLc as well as triglyceride-rich beta very-low-density lipoprotein (VLDL), LDL, and HDL lipoproteins (Carlson et al. 1986). The molecular basis of HL deficiency has been demonstrated biochemically by lack of HL activity in postheparin plasma from affected cases (Carlson et al. 1986; Little et al. 1986). Hegele et al. found several mutations of a gene designated LIPC among patients with familial HL deficiency, establishing a genetic basis for this condition (Hegele et al. 1991a, b). Human LIPC is a large gene spanning more than 60 kb on human chromosome 15q21 and consists of nine exons and eight introns (Cai et al. 1989; Sparkes et al. 1987).
HL is a 477-amino-acid glycoprotein that is synthesized and secreted primarily in the liver. It belongs to a family of enzymes involved in regulation of lipoprotein metabolism in plasma. It catalyzes the hydrolysis of triglycerides of large remnant VLDL, LDL, and nascent HDL1 to form dense HDL(2) (Kinnunen 1984; Jensen et al. 1982). Elevated enzymatic activity of HL has been associated with high HDL levels (Santamarina-Fojo et al. 1998; Dugi et al. 2000; Zambon et al. 2000).
The enzyme also mediates hepatic uptake of HDL cholesteryl esters and contributes to the reverse cholesterol transport system (Lambert et al. 1999). Association between low HL activity and coronary artery disease might reflect a decrease in HL-mediated uptake of lipoproteins by the liver (Brunzell and Deeb 2001). The enzymatic activity of HL varies widely in normal individuals (five- to eightfold), probably under the influence of genetic variation and other environmental factors (Zambon et al. 2003).
Previous attempts to correlate variations of LIPC with HDLc levels in general populations have shown conflicting results. In particular, recent evidence argues both for and against association between a single nucleotide polymorphism (SNP) in the promoter region, −514C/T, and plasma HDLc levels. Some studies have shown a significant association with plasma HDL (Jansen et al. 1997; Couture et al. 2000; Guerra et al. 1997; Hubácek et al. 2001; Jansen et al. 1999; Murtomäki et al. 1997; Ordovas et al. 2002; Yamakawa-Kobayashi et al. 2002; Zambon et al. 2003; Chen et al. 2003; Ji et al. 2002; Ko et al. 2004), but other studies (Vega et al. 1998; Tahvanainen et al. 1998; Grundy et al. 1999; Carr et al. 2002; Fang and Liu 2002; Hegele et al. 1999; Juo et al. 2001; Hong et al. 2000; St-Pierre et al. 2003) have failed to demonstrate it. The disparate data might reflect differences in ethnicity, gender, and/or heterogeneity of the test populations in clinical settings, as well as the fact that most of these studies involved only small population samples. Earlier studies in Japanese groups suffered from selection bias; for example, subjects were chosen from among patients undergoing hemodialysis or postmenopausal hormone replacement therapy.
We approached the problem of defining a possible relationship between the LIPC locus and HDLc levels in plasma by typing multiple SNPs located in intron 1 of LIPC and investigating haplotype association. Here we describe association between a haplotype in intron 1 of LIPC and hyperalphalipoproteinemia in two independent Japanese populations for applying in medical practice.
Materials and methods
Study populations
This investigation was part of an ongoing Molecular Epidemiological Study utilizing the Regional Characteristics of the Twenty-first Century Centers of Excellence (COE) Program in Japan. The first regional population was Takahata (T), a local area of northeastern Japan with a resident population of 15,222 adults over the age of 40 years (men 7,109, women 8,113), as described previously in detail (Konta et al. 2006). In 2004 or 2005, 1,343 men and 1,626 women (total 2,970 residents in regional population T) took part in the program, agreed to join the study, and went through the medical checkup. In this regional population, mean age and body mass index (BMI) with standard error (SE) were 63.0 ± 0.2 years and 23.5 ± 0.1 kg/m2, respectively. Mean values and SD of HDLc in regional population T were 59.1 ± 0.3 mg/dl (Table 1). The second regional population, Funagata (F), consisted of 1,638 individuals in another local area of northeastern Japan; all agreed to join the study. A Caucasian regional population studied elsewhere for ethnic comparison from the state of Utah, USA (U), consisted of a random sample of 488 men and 541 women (total 1,029 individuals).
All works were approved by the institutional ethical committee, and all participants gave written informed consent.
Measurement of plasma lipoproteins
The protocols for lipid and lipoprotein measurement were based on Centers for Disease Control and Prevention (CDC)-approved procedures referred from ARUP National Reference Laboratory, as described previously (Wu et al. 1989). In brief, plasma cholesterol and triglyceride concentrations were assayed enzymatically, and concentrations of HDLc were determined by MgCl2-dextran precipitation methods.
Selection and genotyping of single nucleotide polymorphisms (SNPs)
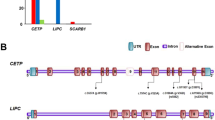
We extracted four SNPs present in intron 1 of the LIPC gene according to the dbSNP database of the National Center for Biotechnology Information (NCBI) (http://www.ncbi.nlm.nih.gov/SNP/) (Fig. 1). Genotyping of these four SNPs (rs12594375, rs8023503, rs4775047, and rs11634134) was performed by Invader assay (Third Wave Technologies, Madison, WI, USA) according to the manufacturer’s protocol (Mein et al. 2000; Lyamichev et al. 1999). The manufacturer provided designed probe sets along with the required reagents. Some genotypes were determined by TaqMan Assay (Applied Biosystems) according to the manufacturer’s protocol using the ABI prism 7900HT (Livak 1999).
Haplotyping, linkage disequilibrium, and statistical analysis
Quantitative associations between genotypes and clinical parameters were analyzed by analysis of variance (ANOVA) with regression analysis as a post hoc test using SPSS version 15.0.1J (SPSS, Chicago, IL, USA). The three genotypic categories of each SNP were indicated, e.g., C/C, C/T, and T/T for rs8041059, with values corresponding to the number of chromosomes possessing a minor allele of the respective SNP. Association was examined among subjects in two Japanese regional populations. Significant association was defined when the given P value of the ANOVA F test was 1% (P < 0.01). Chi-square tests were used to ascertain Hardy–Weinberg equilibrium among genotypes (P > 0.05) using the R2.4.1 package, genetics (http://www.r-project.org/). Linkage disequilibrium (LD) for all possible two-way combinations of the SNPs was tested with D’ by using Haploview Ver.3.3 (Barrett et al. 2005). Haplotype frequencies were calculated by the haplo.stats program, which implements the methods of Schaid et al. (2001). Haplotypes were inferred, and haplotype frequencies were estimated using the modified expectation-maximization (EM) method of haplotype inference included in the haplo.stats program (Schaid et al. 2001). Haplotype-based associations with clinical traits were performed using the haplo.stats program, and output global-score statistics and haplotype-specific scores were derived from generalized linear models.
Results
Almost complete linkage disequilibrium between SNPs rs8023503C/T and rs12594375G/A in intron 1 of LIPC
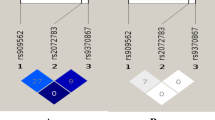
A panel of 2,970 Japanese adult participants was first analyzed in detail for genotypes of four SNPs present in intron 1 of LIPC: rs12594375G/A, rs8023503C/T, rs4775047C/T, and rs11634134T/A. Each of these sites had a minor allele frequency >0.1 in this test population at a call rate >99% (Table 2). No deviation of genotype frequencies from Hardy–Weinberg equilibrium occurred among our subjects in any of the polymorphisms. We analyzed LD between the four SNPs among all 2,970 subjects using Thompson’s method (D′ and r 2). We observed an obvious consistency of genotypes between two SNPs, rs8023503C/T and rs12594375G/A, by detecting almost complete LD (D′ = 0.97 and r 2 = 0.91) (Fig. 2a, b).
Calculating the frequencies of the constructed haplotypes predicted two major haplotypes segregating among the test population. The most frequently observed haplotype, rs8023503C/rs12594375G, designated here as haplotype-1 (H1), constituted 66.9% of the predicted haplotypes (Table 3). The other haplotyp,e rs8023503T/rs12594375A, designated here as haplotype-2 (H2), represented 31.3% of the haplotypes. Because the predicted frequencies of two other haplotypes, rs8023503C/rs12594375A and rs8023503T/rs12594375G, were rare (about 2%), we regarded them as negligible for this analysis.
Association of haplotype 2 with elevated HDLc levels
Association of the determined genotypes and haplotypes to plasma levels of HDLc was analyzed among all 2,970 Japanese participants. ANOVA and linear regression tests identified significant association for each of two SNPs: rs12594375G/A (P = 0.00003) and rs8023503 (P = 0.00010) (Table 4). Similar analysis of the two frequent haplotypes versus HDLc levels in the same individuals revealed a significant association (P = 0.00011). Comparison of HDLc levels among the three haplotypic categories—haplotype-1 homozygotes (H1H1), heterozygotes (H1H2), and haplotype-2 homozygotes (H2H2)—revealed the lowest HDLc levels among H1H1 individuals (mean ± SE = 58.4 ± 0.4 mg/dl), highest for H2H2 (62.5 ± 0.8 mg/dl), and intermediate for H1H2 (59.2 ± 0.4 mg/dl). Possession of haplotype-2 appeared to elevate plasma HDLc.
When subjects were separated into two groups, those who bore at least one haplotype-1 (H1H1 and H1H2) and those who lacked haplotype-1 (H2H2 homozygotes), the former group registered significantly lower plasma HDLc levels than the latter (mean ± SE = 58.8 ± 0.3 mg/dl vs. 62.5 ± 0.8 mg/dl, P = 0.00005) (Fig. 3).
Plasma high-density lipoprotein cholesterol levels (mg/dl) in regional population T (Takahata) (2,970 Japanese adults); P values were calculated by analysis of variance. Histogram bars represent mean values; T-bars above them indicate standard error. The number of individuals analyzed for each haplotype is shown in parentheses under the respective columns
Validation of the association of haplotype-2 with high HDLc in a separate Japanese population
To examine reproducibility of our findings, we chose to analyze genotypes of rs8023503 against HDLc for validation of association in regional population F. ANOVA and logistic regression analysis in these subjects again revealed a significant association between the LIPC allele and HDLc levels; i.e., mean plasma concentrations of HDLc were higher among 173 homozygotic T-allele carriers (mean ± SE = 60.9 ± 1.1 mg/dl) and 735 heterozygotes (60.1 ± 0.5 mg/dl) than among 725 homozygotic C-allele carriers (57.5 ± 0.5 mg/dl) (P = 0.00070) (Fig. 4).
Association of one LIPC polymorphism with high-density lipoprotein cholesterol levels (mg/dl) in regional population F (Funagata); P values were calculated by analysis of variance. Histogram bars represent mean values; T-bars above them indicate standard error. The number of individuals analyzed for each genotype is shown in parentheses under each column
Association of LIPC alleles with HDLc levels in the Caucasian study population was not possible because no variation of the rs8023503 SNP was detectable in the regional population U.
Multivariate analysis of HDLc levels against SNPs in intron 1 of LIPC and other parameters
To examine association with HDLc levels of other factors, we investigated multiple logistic regression analysis including two SNPs (rs12594375 and rs8023503), gender, age, BMI, systolic blood pressure, diastolic blood pressure, and lifestyle (drinking and smoking status) as confounding factor showed that each of these factors was associated with HDLc. There was interaction between HDLc and rs12594375 (P < 0.05), but it was not shown in rs8023503 (P > 0.05). Other factors—BMI, drinking (drinker and nondrinker), and smoking (smoker and nonsmoker), also were associated with HDLc (P < 0.05, respectively).
Discussion
In this study, we found that significant association between HDLc and a specific haplotype in intron 1 of the LIPC gene was detected in both populations. HDL system has received attention in recent years because of its close relationship to the incidence of coronary heart disease. A number of epidemiological investigations have shown an inverse relationship between the incidence of coronary heart disease and high levels of HDLc in plasma (Castelli et al. 1977; Nikkila 1976; Miller and Miller 1975).
Several attempts have been made to examine a potential relationship between HDLc and variation in the LIPC gene, focusing on an SNP in the 5′ flanking region, −514C/T (Jansen et al. 1997). Jansen et al. first reported association of this promoter SNP with HDLc levels in 1997; subsequent studies confirmed the significance of this association (Jansen et al. 1997; Couture et al. 2000; Guerra et al. 1997; Hubácek et al. 2001; Jansen et al. 1999; Murtomäki et al. 1997; Ordovas et al. 2002; Yamakawa-Kobayashi et al. 2002; Zambon et al. 2003; Chen et al. 2003; Ji et al. 2002; Ko et al. 2004; Arai et al. 2005), and rejected (Vega et al. 1998; Tahvanainen et al. 1998; Grundy et al. 1999; Carr et al. 2002; Fang and Liu 2002; Hegele et al. 1999; Juo et al. 2001; Hong et al. 2000; St-Pierre et al. 2003). Although genetic heterogeneity and other confounding factors, such as ethnic differences, are clearly possibilities, the contradictory results might reflect the limitations of analysis using only the −514C/T variation. The exclusive use of this single SNP in every previous study overlooked other regions of the LIPC gene that might have shown associations with HDLc.
In the work reported here, we studied haplotype association by analyzing four SNPs present in intron 1 of LIPC in two independent Japanese adult populations (regional population T and regional population F). By constructing haplotypes and analyzing them for association with plasma lipoprotein levels, we demonstrated a significant association of elevated HDLc levels with one of the major haplotypes constructed from two intronic SNPs that were in almost complete LD. Plasma HDLc levels were higher among haplotype-2 homozygotes than among heterozygotes or haplotype-1 homozygotes. These data implied that variation within the first intron of LIPC might have affected lipoprotein metabolism in these individuals, eventually inducing variation in plasma HDLc levels. Elevated HDLc could be a result of accelerated turnover to mature HDL(2) particles after uptake of HDL particles by the liver, or a reduced liver uptake of HDL. Future analysis of haplotype variation among various ethnicities, ages, and disease states would be required to prove the hypothesis.
Several interpretations are possible for the association with HDLc that we observed for the two SNPs that were in strong LD in our Japanese population. First, one site or the other might play a predominant mechanistic role. Second, both nucleotide changes could be required to impart the functional change that underlies the observed association (i.e., a codominant effect of the two polymorphisms). Third, these markers might themselves be in LD with as yet unmeasured and functional variants that are the true mechanistic basis for the association. These possibilities should be distinguished by functional studies in the future. In addition, multivariate analysis showed that intronic variations of the LIPC gene and other risk factors (BMI, smoking, and drinking) were associated with HDLc level.
In conclusion, we identified a haplotype association between intronic variations of the LIPC gene and plasma HDLc levels in a Japanese population consisting of 2,970 adults and validated the finding in a second Japanese population. The implied involvement of these genetic variations in HDL metabolism may explain, at least in part, the wide variations in plasma HDLc levels noted among individuals and may contribute to establishment of designs for medical checkup, suitable treatments, and preventive strategies to address low HDL as a risk factor for coronary artery disease.
Abbreviations
- ANOVA:
-
analysis of variance
- CETP:
-
cholesteryl ester transfer protein
- HDLc:
-
high-density lipoprotein cholesterol
- HL:
-
hepatic lipase
- LD:
-
linkage disequilibrium
- LDLc:
-
low-density lipoprotein cholesterol
- LIPC :
-
hepatic lipase gene
- LPL:
-
lipoprotein lipase
- QTL:
-
quantitative trait locus
- RLP:
-
remnant-like particle cholesterol
- SNP:
-
single nucleotide polymorphism
- TC:
-
total cholesterol
- TG:
-
triglyceride
- VLDL:
-
very-low-density lipoprotein
References
Arai H, Yamamoto A, Matsuzawa Y, Saito Y, Yamada N, Oikawa S, Mabuchi H, Teramoto T, Sasaki J, Nakaya N, Itakura H, Ishikawa Y, Ouchi Y, Horibe H, Egashira T, Hattori H, Shirahashi N, Kita T (2005) Polymorphisms in four genes related to triglyceride and HDL-cholesterol levels in the general Japanese population in 2000. J Atheroscler Thromb 12:240–250
Barrett JC, Fry B, Maller J, Daly MJ (2005) Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21:263–265
Brunzell JD, Deeb SS (2001) Familial lipoprotein lipase deficiency, apo C-II deficiency, and hepatic lipase deficiency. In: Scriver CR, Beaudet AL, Sly WS, Valle D, Childs B, Kinzler KW, Vogelstein B (eds) The metabolic and molecular basis of inherited disease. McGraw-Hill, New York, pp 2789–2816
Cai SJ, Wong DM, Chen SH, Chan L (1989) Structure of the human hepatic triglyceride lipase gene. Biochemistry 28:8966–8971
Carlson LA, Holmquist L, Nilsson-Ehle P (1986) Deficiency of hepatic lipase activity in post-heparin plasma in familial hyper-alpha-triglyceridemia. Acta Med Scand 219:435–447
Carr MC, Ayyobi AF, Murdoch SJ, Deeb SS, Brunzell JD (2002) Contribution of hepatic lipase, lipoprotein lipase, and cholesteryl ester transfer protein to LDL and HDL heterogeneity in healthy women. Arterioscler Thromb Vasc Biol 22:667–673
Castelli WP, Doyle JT, Gordon T, Hames CG, Hjortland MC, Hulley SB, Kagan A, Zukel WJ (1977) HDL cholesterol and other lipids in coronary heart disease. The cooperative lipoprotein phenotyping study. Circulation 55:767–772
Chen W, Srinivasan SR, Boerwinkle E, Berenson GS, Bogalusa Heart Study (2003) Hepatic lipase promoter C-514T polymorphism influences serial changes in HDL cholesterol levels since childhood: the Bogalusa Heart Study. Atherosclerosis 169:175–182
Couture P, Otvos JD, Cupples LA, Lahoz C, Wilson PW, Schaefer EJ, Ordovas JM (2000) Association of the C-514T polymorphism in the hepatic lipase gene with variations in lipoprotein subclass profiles: the Framingham Offspring Study. Arterioscler Thromb Vasc Biol 20:815–822
Dugi KA, Amar MJ, Haudenschild CC, Shamburek RD, Bensadoun A, Hoyt RF Jr, Fruchart-Najib J, Madj Z, Brewer HB Jr, Santamarina-Fojo S (2000) In vivo evidence for both lipolytic and nonlipolytic function of hepatic lipase in the metabolism of HDL. Arterioscler Thromb Vasc Biol 20:793–800
Fang DZ, Liu BW (2002) Polymorphism of HL +1075C, but not −480T, is associated with plasma high density lipoprotein cholesterol and apolipoprotein AI in men of a Chinese population. Atherosclerosis 161:417–424
Glueck CJ, Fallat RW, Millett F, Gartside P, Elston RC, Go RC (1975) Familial hyper-alpha-lipoproteinemia: studies in eighteen kindreds. Metabolism 24:1243–1265
Glueck CJ, Gartside PM, Tsang RC, Mellies MJ, Steiner PM (1977) Neonatal familial hyperalphalipoproteinemia. Metabolism 26:469–472
Grundy SM, Vega GL, Otvos JD, Rainwater DL, Cohen JC (1999) Hepatic lipase activity influences high density lipoprotein subclass distribution in normotriglyceridemic men. Genetic and pharmacological evidence. J Lipid Res 40:229–234
Guerra R, Wang J, Grundy SM, Cohen JC (1997) A hepatic lipase (LIPC) allele associated with high plasma concentrations of high density lipoprotein cholesterol. Proc Natl Acad Sci USA 94:4532–4537
Hegele RA, Little JA, Connelly PW (1991a) Compound heterozygosity for mutant hepatic lipase in familial hepatic lipase deficiency. Biochem Biophys Res Commun 179:78–84
Hegele RA, Vezina C, Moorjani S, Lupien PJ, Gagne C, Brun LD, Little JA, Connelly PW (1991b) A hepatic lipase gene mutation associated with heritable lipolytic deficiency. J Clin Endocrinol Metab 72:730–732
Hegele RA, Harris SB, Brunt JH, Young TK, Hanley AJ, Zinman B, Connelly PW (1999) Absence of association between genetic variation in the LIPC gene promoter and plasma lipoproteins in three Canadian populations. Atherosclerosis 146:153–160
Hong SH, Song J, Kim JQ (2000) Genetic variations of the hepatic lipase gene in Korean patients with coronary artery disease. Clin Biochem 33:291–296
Hubácek JA, Waterworth DM, Pitha J, Humphries SE, Talmud PJ, Poledne R (2001) Polymorphisms in the lipoprotein lipase and hepatic lipase genes and plasma lipid values in the Czech population. Physiol Res 50:345–351
Jansen H, Verhoeven AJ, Weeks L, Kastelein JJ, Halley DJ, van den Ouweland A, Jukema JW, Seidell JC, Birkenhäger JC (1997) Common C-to-T substitution at position −480 of the hepatic lipase promoter associated with a lowered lipase activity in coronary artery disease patients. Arterioscler Thromb Vasc Biol 17:2837–2842
Jansen H, Chu G, Ehnholm C, Dallongeville J, Nicaud V, Talmud PJ (1999) The T allele of the hepatic lipase promoter variant C-480T is associated with increased fasting lipids and HDL and increased preprandial and postprandial LpCIII:B: European Atherosclerosis Research Study (EARS) II. Arterioscler Thromb Vasc Biol 19:303–308
Jensen GL, Daggy B, Bensadoun A (1982) Triacylglycerol lipase, monoacylglycerol lipase and phospholipase activities of highly purified rat hepatic lipase. Biochim Biophys Acta 710:464–470
Ji J, Herbison CE, Mamotte CD, Burke V, Taylor RR, van Bockxmeer FM (2002) Hepatic lipase gene -514 C/T polymorphism and premature coronary heart disease. J Cardiovasc Risk 9:105–113
Juo SH, Han Z, Smith JD, Colangelo L, Liu K (2001) Promoter polymorphisms of hepatic lipase gene influence HDL(2) but not HDL(3) in African American men: CARDIA study. J Lipid Res 42:258–264
Ko YL, Hsu LA, Hsu KH, Ko YH, Lee YS (2004) The interactive effects of hepatic lipase gene promoter polymorphisms with sex and obesity on high-density-lipoprotein cholesterol levels in Taiwanese-Chinese. Atherosclerosis 172:135–142
Kinnunen PK (1984) Hepatic endothelial lipase: isolation, some characteristics, and physiological role. In: Borgstrom B, Brockman HL (eds) Lipases. Elsevier Scientific, New York, pp 307–382
Konta T, Hao Z, Abiko H, Ishikawa M, Takahashi T, Ikeda A, Ichikawa K, Takasaki S, Kubota I (2006) Prevalence and risk factor analysis of microalbuminuria in Japanese general population: the Takahata study. Kidney Int 70:751–756
Lambert G, Chase MB, Dugi K, Bensadoun A, Brewer HB Jr, Santamarina-Fojo S (1999) Hepatic lipase promotes the selective uptake of high density lipoprotein-cholesteryl esters via the scavenger receptor B1. J Lipid Res 40:1294–1303
Little JA, Connelly PW (1986) Familial hepatic lipase deficiency. Adv Exp Med Biol 201:253–260
Livak KJ (1999) Allelic discrimination using fluorogenic probes and the 5′ nuclease assay. Genet Anal 14:143–149
Lyamichev V, Mast AL, Hall JG, Prudent JR, Kaiser MW, Takova T, Kwiatkowski RW, Sander TJ, de Arruda M, Arco DA, Neri BP, Brow MA (1999) Polymorphism identification and quantitative detection of genomic DNA by invasive cleavage of oligonucleotide probes. Nat Biotechnol 17:292–296
Mein CA, Barratt BJ, Dunn MG, Siegmund T, Smith AN, Esposito L, Nutland S, Stevens HE, Wilson AJ, Phillips MS, Jarvis N, Law S, de Arruda M, Todd JA (2000) Evaluation of single nucleotide polymorphism typing with invader on PCR amplicons and its automation. Genome Res 10:330–343
Miller GJ, Miller NE (1975) Plasma high-density-lipoprotein concentration and development of ischaemic heart disease. Lancet 1:16–19
Murtomäki S, Tahvanainen E, Antikainen M, Tiret L, Nicaud V, Jansen H, Ehnholm C (1997) Hepatic lipase gene polymorphisms influence plasma HDL levels. Results from Finnish EARS participants. European Atherosclerosis Research Study. Arterioscler Thromb Vasc Biol 17:1879–1884
Nikkila EA (1976) Letter: serum high-density-lipoprotein and coronary heart disease. Lancet 2:320
Ordovas JM, Corella D, Demissie S, Cupples LA, Couture P, Coltell O, Wilson PW, Schaefer EJ, Tucker KL (2002) Dietary fat intake determines the effect of a common polymorphism in the hepatic lipase gene promoter on high-density lipoprotein metabolism: evidence of a strong dose effect in this gene-nutrient interaction in the Framingham Study. Circulation 106:2315–2321
Santamarina-Fojo S, Haudenschild C, Amar M (1998) The role of hepatic lipase in lipoprotein metabolism and atherosclerosis. Curr Opin Lipidol 9:211–219
Schaid DJ, Rowland CM, Tines DE, Jacobson RM, Poland GA (2001) Score tests for association between traits and haplotypes when linkage phase is ambiguous. Am J Hum Genet 70:425–434
Sparkes RS, Zollman S, Klisak I, Kirchgessner TG, Komaromy MC, Mohandas T, Schotz MC, Lusis AJ (1987) Human genes involved in lipolysis of plasma lipoproteins: mapping of loci for lipoprotein lipase to 8p22 and hepatic lipase to 15q21. Genomics 1:138–144
St-Pierre J, Miller-Felix I, Paradis ME, Bergeron J, Lamarche B, Després JP, Gaudet D, Vohl MC (2003) Visceral obesity attenuates the effect of the hepatic lipase −514C>T polymorphism on plasma HDL-cholesterol levels in French-Canadian men. Mol Genet Metab 78:31–36
Tahvanainen E, Syvanne M, Frick MH, Murtomaki-Repo S, Antikainen M, Kesaniemi YA, Kauma H, Pasternak A, Taskinen MR, Ehnholm C (1998) Association of variation in hepatic lipase activity with promoter variation in the hepatic lipase gene. The LOCAT Study Invsestigators. J Clin Invest 101:956–960
Vega GL, Gao J, Bersot TP, Mahley RW, Verstraete R, Grundy SM, White A, Cohen JC (1998) The -514 polymorphism in the hepatic lipase gene (LIPC) does not influence androgen-mediated stimulation of hepatic lipase activity. J Lipid Res 39:1520–1524
Wu LL, Warnick GR, Wu JT, Williams RR, Lalouel JM (1989) A rapid micro-scale procedure for determination of the total lipid profile. Clin Chem 35:1486–1491
Yamakawa-Kobayashi K, Somekawa Y, Fujimura M, Tomura S, Arinami T, Hamaguchi H (2002) Relation of the −514C/T polymorphism in the hepatic lipase gene to serum HDL and LDL cholesterol levels in postmenopausal women under hormone replacement therapy. Atherosclerosis 162:17–21
Zambon A, Deeb SS, Bensadoun A, Foster KE, Brunzell JD (2000) In vivo evidence of a role for hepatic lipase in human apoB-containing lipoprotein metabolism, independent of its lipolytic activity. J Lipid Res 41:2094–2099
Zambon A, Deeb SS, Pauletto P, Crepaldi G, Brunzell JD (2003) Hepatic lipase: a marker for cardiovascular disease risk and response to therapy. Curr Opin Lipidol 14:179–189
Acknowledgments
This study was supported in part by the Twenty-first Century COE (Center of Excellence) program from the Ministry of Education, Culture, Sports, Science, and Technology, Japan.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Iijima, H., Emi, M., Wada, M. et al. Association of an intronic haplotype of the LIPC gene with hyperalphalipoproteinemia in two independent populations. J Hum Genet 53, 193–200 (2008). https://doi.org/10.1007/s10038-007-0236-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-007-0236-0