Abstract
We describe three patients with retinoblastoma, dysmorphic features and developmental delay. Patients 1 and 2 have high and broad forehead, deeply grooved philtrum, thick anteverted lobes and thick helix. Patient 1 also has dolicocephaly, sacral pit/dimple and toe crowding; patient 2 shows intrauterine growth retardation and short fifth toe. Both patients have partial agenesis of corpus callosum. Patient 3 has growth retardation, microcephaly, thick lower lip and micrognathia. Using array-comparative genomic hybridization (CGH), we identified a 13q14 de novo deletion in patients 1 and 2, while patient 3 had a 7q11.21 maternally inherited deletion, probably not related to the disease. Our results confirm that a distinct facial phenotype is related to a 13q14 deletion. Patients with retinoblastoma and malformations without a peculiar facial phenotype may have a different deletion syndrome or a casual association of mental retardation and retinoblastoma. Using array-CGH, we defined a critical region for mental retardation and dysmorphic features. We compared this deletion with a smaller one in a patient with retinoblastoma (case 4) and identified two distinct critical regions, containing 30 genes. Four genes appear to be good functional candidates for the neurological phenotype: NUFIP1 (nuclear fragile X mental retardation protein 1), HTR2A (serotonin receptor 2A), PCDH8 (prothocaderin 8) and PCDH17 (prothocaderin 17).
Similar content being viewed by others
Introduction
Retinoblastoma is the most common intraocular tumor of early childhood, with an incidence of 1/15,000–28,000 live births. Tumor development is caused by inactivation of both alleles of the RB1 gene located in 13q14.2. In 68% of cases RB1 is inactivated by point mutations, in 5% RB1 complete gene deletions have been found, while gross-sized molecular deletions have been found in 10% of cases (Albrecht et al. 2005; Dahiya et al. 2000; Kloss et al. 1991; Lohmann and Gallie 2004; Sampieri et al. 2006). When the deletion involves part of the RB1 surrounding genome it causes a contiguous gene deletion syndrome characterized by retinoblastoma, developmental abnormalities and peculiar facial dysmorphisms. The first author to suggest a specific facial phenotype associated with 13q14 deletion was Motegi in 1983 (Motegi et al. 1983). He described two patients with retinoblastoma and common facial features including prominent eyebrows, broad nasal bridge, bulbous nasal tip, large mouth, thin upper lip and long philtrum (Motegi, et al. 1983). A few years later, he described an additional patient, and was able to improve the clinical definition of the syndrome (Motegi et al. 1987). In 1999, Baud et al. (1999) defined the dysmorphic features of 13q14 deletion syndrome. He described a cohort of 22 patients with the following common dysmorphic abnormalities: high and broad forehead, thick and everted ear lobes, short nose, prominent philtrum and thick everted lower lip (Baud et al. 1999). In 2001, Bojinova et al. (2001) extended the facial phenotype associated with the 13q14 deletion syndrome with the description of additional 13 patients characterized by cranial anomalies, frontal bossing, deeply grooved and long philtrum, depressed and broad nasal bridge, bulbous tip of the nose, thin upper lip, broad cheeks, and large ears and lobules. Afterward, a patient with a X:13 translocation and phenotypic features peculiar to the 13q14 deletion syndrome was described (Dries et al. 2003). Finally, in 2004, a patient with retinoblastoma, pinealoma and mild multiple congenital anomalies/mental retardation syndrome (MCA/MR) and a germline 13q14 deletion were reported (Skrypnyk and Bartsch 2004).
All these reported cases were studied by means of cytogenetic analysis. We investigated using array-based comparative genomic hybridization (array-CGH) three patients with retinoblastoma and MCA/MR. Using the same method, we analyzed an additional patient with isolated retinoblastoma and a previously identified RB1 deletion (Sampieri et al. 2006) to attempt to define a minimal critical region for MCA/MR. Patients were selected among the cohort of retinoblastoma cases collected in the biobank of the Medical Genetics Unit of the University of Siena (http://www.biobank.unisi.it). Here we report an accurate clinical and molecular characterization of these patients.
Materials and methods
Array-CGH analysis
Array-CGH analysis was performed using commercially available oligonucleotide microarrays containing approximately 43,000 60-mer probes (Human Genome CGH Microarray 44B Kit. Agilent Technologies, Santa Clara, California), as previously reported (Pescucci et al. 2006). The average resolution is about 75–100 kb.
Real-time quantitative PCR
Real-time quantitative polymerase chain reaction (PCR) was performed to confirm array-CGH data. We used TaqMan Gene Expression Assays by design (Applied Biosystems, http://www.products.appliedbiosystems.com), which provides a pre-designed primers-probe set for real-time PCR experiments. In order to validate the presence of the 13q deletion in cases 1, 2 and 4, we used the TaqMan probe and primers in exon 17 of RB1, as previously described (Sampieri et al. 2006). For validating the presence of the 7q deletion in case 3, we designed the probe in the BC066990 sequence related to the 7q11.21 locus. Forward primer: 5′-GTG CTG TAG TGC AGA ATG TAA CAA A-3′; reverse primer: 5′-CAG AAA GCC AAG AAT AAC-3′; TaqMan probe: 5′-AGG GTG AAC AAA ACC AGT TGA GTT-3′. PCR was carried out using an ABI prism 7000 (Applied Biosystems) in a 96-well optical plate with a final reaction volume of 50 μl. A total of 100 ng (10 μl) was dispensed in each of the four sample wells for quadruplicate reactions. Thermal cycling conditions included a pre-run of 2 min at 50°C and 10 min at 95°C. Cycle conditions were 40 cycles at 95°C for 15 s and 60°C for 1 min, according to the TaqMan Universal PCR Protocol (ABI). The TaqMan Universal PCR Master Mix and Microamp reaction tubes were supplied by Applied Biosystem. The starting copy number of the unknown samples was determined using the comparative Ct method, as previously described (Livak 1997).
RB1 mutation analysis
Genomic DNA was amplified by PCR. Primers and PCR conditions for single exons and promoter analysis have been described previously (Hogg et al. 1992; Houdayer et al. 2004; Scheffer et al. 2000). PCR products were mixed with an equal volume of formamide, denatured by heating at 95°C for 5 min, followed by immediate chilling on ice. Single-strand conformational polymorphism (SSCP) was performed on a Genephor apparatus (Pharmacia Amersham, Little Braunschweig, Germany) using a GeneGel Excel 12.5/24 Kit (Pharmacia Amersham).
Results
Clinical description
Case 1
Patient number 1, a 1-year and 2-month-old female, is the first and only child of healthy unrelated parents (Fig. 1a). At birth, the mother and father were 28 and 33 years old, respectively. Their family history was unremarkable. No teratogen exposure during pregnancy had been reported. The child was born on term by means of caesarean delivery. Birth weight was 3,130 g (50th percentile), length was 51 cm (50th percentile) and head circumference was 36 cm (>90th percentile). Bilateral retinoblastoma was diagnosed at 5 months of age. At that time, MRI revealed corpus callosum hypoplasia. ABR and ankle ultrasonography were normal. At our first examination (6 months), psychomotor development was slightly delayed. Her weight was 7.850 g (75th percentile), length 68 cm (90th percentile) and head circumference 46 cm (97th percentile) with dolicocephaly. The patient presented scalp anomalies including widely open fontanelles and an alopecic area on the right temporo-parietal region. High and broad forehead, deeply grooved philtrum, and thick anteverted lobes and thick helix were noted. In addition, she showed sacral dimple and clinodactyly of the 5th toe on the left and toe crowding on the right (Fig. 2a). At the age of 11 months, she presented with a relapse in the right eye treated by chemotherapy. Two months later, the right eye was enucleated. In the following months, a relapse in the left eye occurred, which was treated successfully by means of radiotherapy. At our second clinical examination (14 months), her weight was 9 kg (10–25th percentile), length 76–77 cm (50th percentile) and head circumference 49 cm (>97th percentile). Psychomotor delay persisted. Ultrasound cardiac examination was normal (Table 1).
Face and profile views of the patients. a Case 1: patient no. 1 at the age of 1 year 2 months. Frontal view showing high and broad forehead, deeply grooved philtrum. Side view showing dolicocephaly and thick anteverted lobes and helix. b Case 2: patient no. 2 at the age of 2 years 7 months. Frontal view showing hypotelorism, long palpebral fissures, epichantic folds, slight unilateral ptosis and thick and everted lower lip. Thick anteverted lobes and helix are showed on the side view. c Case 3: patient no. 3 at the age of 7 years 6 months. Frontal view showing sparse eyebrows in the medial third broad nasal bridge, bulbous tip of the nose, long philtrum, thick and everted lower lip. Side view showing large ears and micrognathia
Case 2
Patient number 2, a 2-year and 7-month-old boy, is the third-born of healthy and non-consanguineous parents (Fig. 1b). Intrauterine growth retardation was noted at the 36th week of gestation. He was born on term. At birth, his weight was 2,300 g (<3rd percentile), length was 47 cm (10–25th percentile) and OFC was 32 cm (3rd–10th percentile). He demonstrated a deficit of thermoregulation. On the 11th day, he presented with enterococcus sepsis. During the first months of life, the parents noted iris bilateral heterochromia. Right eye retinoblastoma was diagnosed at 10 months of age. At 1 year of age, a MRI was performed and hypoplasia of the corpus callosum was noted. By 1 year and 5 months of age, he suffered a relapse, which was treated by enucleation. At our first examination (2 years of age) his height was 78 cm (<3rd percentile), weight 8,250 kg (<<3rd percentile) and OFC 44 cm (<<3rd percentile). He showed hypotonia and particular facial features including high and broad forehead, deeply grooved philtrum, thick and everted lower lip, thick and everted auricular lobes, and thick helix. Moreover, he had short 5th toe with hypoplastic toenail (Fig. 2b). A second clinical examination 7 months later confirmed growth delay: 80 cm in height (<5th percentile), weight 9.0 kg (<<3rd percentile) and OFC 45 cm (<<3rd percentile). The previously noted facial features were still present. There were no abnormalities of other organs and systems. He reached self-governing deambulation at 2 years and 6 months. Presently, he is able to say only few words and he has no sphincter control. An echocardiogram showed minimum aortic reflux, probably due to the infantile infection. X-rays of hands and toes indicated no abnormalities (Table 1).
Case 3
Patient number 3, a 7-year and 11-month-old female, is the first-born of healthy, non-consanguineous parents (Fig. 3c) at birth, the mother was 35 years and the father 47 years old. After bearing this child, the mother later had a spontaneous abortion. However, this patient does have a 19-year-old maternal half-sister suspected to have Gilles De La Tourette syndrome; the mother had a spontaneous abortion after her birth. During gestation, ultrasound study revealed microcephaly. This patient was born on term and her weight was 2,780 g (10–25th percentile); data on length and OFC are not available. Development has been slightly delayed: she reached self-governing deambulation at 20 months of age and was able to say her first words when she was 2 years old. At 2.5 years of age, her mother noted right leucoria. Unilateral retinoblastoma was diagnosed and treated with eye enucleation. At our first examination (5 years and 7 months of age), her weight was 14 kg (<5th percentile), height 109.5 cm (25–50th percentile) and OFC 41 cm (<<3rd percentile). Physical examination showed sparse eyebrows in the medial third, epichantic folds, broad nasal bridge, bulbous nasal tip, long philtrum, thick and everted lower lip, large ears, micrognathia and cutis marmorata. She also showed a moderate mental retardation (Table 1). A second clinical examination at 7 years and 11 months of age confirmed short stature (115 cm, <5th percentile), the same previously described facial features and microcephaly (OFC 42 cm, <<3rd percentile).
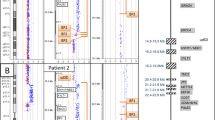
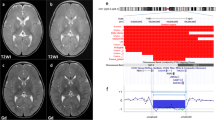
Patients' molecular data. a Array CGH ratio profiles. On the left, chromosome ideogram for each case. On the right, the log2 ratio of the chromosome probes plotted as a function of chromosomal position. Oligos with a value of zero represent equal fluorescence intensity ratio between sample and reference. Each dot represents a single probe (oligo) spotted on the array. Copy number loss shifts the ratio to the left (value of about –2X). b and c Real-time quantitative PCR validation experiment. b RB1 ddCT ratios and standard deviations of two different controls and of patients 1, 2 and 4 and their parents. Patients 1, 2 and 4 and the mother of 4 show a ddCT ratio of about 0.5, indicating the presence of a single copy of RB1 (deletion). The parents of patients 1 and 2 and the father of patient 4, as the controls, show ddCT ratios of about 1.0, which indicates a double copy of the gene. c BC066990 sequence ratios and standard deviations of two different controls and of patient 3 and her parents. The patient and the mother show ddCT ratio of about 0.5, indicating the presence of a single copy of BC066990 sequence (deletion); while the father, like the controls, shows ddCT ratios of about 1.0 indicating a double copy of the sequence
Case 4
Patient number 4, a 5-year and 4-month-old female, is the second child of unrelated parents. At birth, the mother was 20 years and the father 26 years old. Paternal history was unremarkable and the first child of the couple is healthy. The mother and a younger brother, however, have been affected by retinoblastoma. The mother presented with retinoblastoma in the left eye at 11 months of age and enucleation was immediately performed. In the younger brother, the diagnosis was made at 40 days of life. In the second child, patient number 4, the gestation of the proband was unremarkable and no teratogen exposure was reported. Multifocal retinoblastoma in the left eye was diagnosed at 2 years and 7 months. After four cycles of chemotherapy, the tumors showed good regression but three relapses occurred and enucleation was performed. During ophthalmological follow-up, no foci were noted in the right eye. On clinical examination, neither mental retardation nor dysmorphisms have been noted. Isolated unilateral retinoblastoma was the unique clinical sign.
Molecular characterization
Point mutation analysis of promoter and coding sequences of the RB1 gene in the four cases did not reveal any alterations.
Oligonucleotide array-CGH analysis with an averaged spatial resolution of approximately 75 kb was performed on DNA from the four patients. The analysis of ratio profiles in cases 1, 2 and 4 revealed a different sized interstitial deletion in the long arm of chromosome 13. Based on the array findings, the deleted region observed in case 1 was found to extend approximately 19 Mb [46,XXdel(13)(q13.3q21.2)]. The proximal breakpoint is mapped in 13q13.3 (last oligonucleotide present located in 40.34 Mb, first deleted in 40.40 Mb position), while the distal breakpoint is located between 59.29 and 59.36 Mb in 13q21.2 (last oligonucleotide deleted and first present, respectively) (Fig. 3a). The deleted region seen in case 2 is approximately 36 Mb in size [46,Xydel(13)(q14.11q31.1)]. In this case, the proximal breakpoint is mapped in 13q14.11 (last oligonucleotide present located in 43 Mb, first deleted in 43.24 Mb position), while the distal breakpoint is located between 79.27 and 79.80 Mb in 13q31.1 (last oligonucleotide deleted and first present, respectively) (Fig. 3a). In contrast, the array-CGH analysis of case 3 revealed a 200-kb proximal deletion on chromosome 7q [46,XX del(7)(q11.21)](Fig. 3a).
The array-CGH analysis of case 4 allowed us to identify an interstitial deletion of approximately 1.7 Mb [46,XX del(13)(q14.2)]. In this case, both breakpoints are located in 13q14.2 (last oligonucleotide present located in 47.35 Mb, first deleted in 47.44 Mb position while last oligonucleotide deleted in 49.10 Mb position and first oligonucleotide present in 49.17Mb position). In order to confirm array findings, real-time Quantitative PCR experiments were performed in the patients and their parents. In all cases, the deletion was confirmed. Identified deletions of the long arm of chromosome 13 were de novo in patients 1 and 2 and inherited in 4. The 7q microdeletion in patient 3 was inherited from the unaffected mother (Fig. 3b, c).
Discussion
Two distinct syndromes are associated with deletions involving different regions of the long arm of chromosome 13. One of these syndromes is caused by a more distal deletion that involves band q32 and is phenotypically characterized by the presence of severe mental retardation, major malformations and digital anomalies (Brown et al. 1993, 1995). The other one is due to a proximal deletion that involves band q14 and is associated with retinoblastoma and mental retardation. Baud et al. defined the peculiar facial traits of this syndrome that is characterized by anteverted ear lobes, high and broad forehead, and a prominent philtrum (Baud et al. 1999; Motegi et al. 1987, 1983).
The facial features of patients 1 and 2 suggest the 13q14 deletion syndrome described by Baud et al. 1999 (Table 1). In particular, both patients have high and broad foreheads, deeply grooved philtrum, thick and anteverted lobes and thick helices. These characteristics are absent in patient 3. As expected on the basis of clinical examination, array CGH analysis confirmed a 13q14 deletion syndrome in patients 1 and 2 but not in 3. Our findings confirm that 13q14 deletion syndrome is characterized by specific facial features and that this diagnosis may be strongly suspected on a clinical ground even before the genetic test.
A more complex situation is present in case 3. The presence of the small 7q deletion in the unaffected mother suggests that this deletion is not responsible for the clinical phenotype in this patient. In addition, a known copy-number polymorphism (CNP) is located at a distance of 80 kb from our deletion (http://www.projects.tcag.ca/bioxrt/), and the deletion is included in a region of highly homologous duplicated sequences (Sharp et al. 2005). However, three different imprinted regions located on the long arm of chromosome 7 have already been described (http://www.geneimprint.com/site/genes-by-species.Homo+sapiens.imprinted-All). Consequently, additional analyses are necessary to investigate whether the deleted region is imprinted and to definitively rule out the involvement of this small deletion in the clinical phenotype of the patient.
Deletions identified in cases 1 and 2 partially overlap and allowed us to define a minimal critical region for mental retardation and dysmorphic features of about 16 Mb that includes 39 known genes.
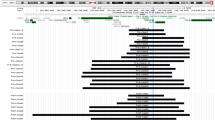
Comparison of this critical region with the 13q14.2 deletion present in case 4 with isolated retinoblastoma allowed us to exclude a central region containing 9 known genes. Consequently, we identified two distinct critical regions, a centromeric sub-region of about 4 Mb and a telomeric one of about 10 Mb. Gene content analysis of the centromeric sub-region showed the presence of 14 known genes (Fig. 4, Table 2). Among them, NUFIP1 is of particular interest due to its putative role in central nervous system (CNS) development and to its preferential brain expression. The NUFIP1 gene encodes for a nucleo-cytoplasmatic RNA binding protein: FMRP interacting protein 1. NUFIP1 interacts with FMRP, the protein disrupted in Fragile X Mental Retardation (Bardoni et al. 2003). In particular, NUFIP1 could be involved in the regulation of local protein synthesis near active synapses in association with FMRP (Bardoni et al. 2003). Due to its role in synaptic plasticity, NUFIP1 could be a good candidate gene for mental retardation in our patients. The HTR2A gene is located within the centromeric deleted sub-region. This gene codifies for a receptor of serotonin. Disturbances in the serotonergic neurotransmission system may be responsible for behavioral disorders (Bruce et al. 2005). In particular, a polymorphism in the HTR2A gene was associated with the remission of attention deficit/hyperactivity disorder (ADHD) (Li et al. 2006). Disruption of serotonin receptor activity may contribute to CNS disorders that have been associated with impaired development.
Gene content of the deleted region (UCSC Genome Browser; http://www.genome.ucsc.edu). a Comparison between deletions observed in cases 1 and 2 and minimal critical region. b Gene content of the two critical sub-regions. Orange rectangles delimit deleted region identified in case 4. Circled genes are discussed in the text
Gene content analysis of the deleted telomeric sub-region showed the presence of 16 known genes (Fig. 4, Table 3). Among them, PCDH8 and PCDH17 may be good candidates for the generation of the neurological phenotype in our patients. These genes belong to the protocadherin gene family and codify for integral membrane proteins, which are thought to function in signaling pathways and in cell adhesion in a CNS-specific manner. The role of Pcdh8 in the nervous system was investigated in rat hippocampus (Yamagata et al. 1999). Antibodies against Pcdh8 attenuate basal synaptic transmission and completely inhibit long-term potentiation in hippocampal slices (Yamagata et al. 1999). The expression and function of the PCDH17 gene is not well known.
To date, all 48 cases with a 13q14 microdeletion reported in the literature have been characterized at the cytogenetic level. This is the first report of characterization at the molecular level, using array-CGH, of patients with retinoblastoma and mental retardation, and a critical region for mental retardation is defined. Further experiments are necessary to narrow this critical region and to dissect the syndrome, thus identifying the gene(s) responsible for the neurological phenotype in these patients.
References
Albrecht P, Ansperger-Rescher B, Schuler A, Zeschnigk M, Gallie B, Lohmann DR (2005) Spectrum of gross deletions and insertions in the RB1 gene in patients with retinoblastoma and association with phenotypic expression. Hum Mutat 26(5):437–445
Bardoni B, Willemsen R, Weiler IJ, Schenck A, Severijnen LA, Hindelang C, Lalli E, Mandel JL (2003) NUFIP1 (nuclear FMRP interacting protein 1 is a nucleocytoplasmic shuttling protein associated with active synaptoneurosomes. Exp Cell Res 289(1):95–107
Baud O, Cormier-Daire V, Lyonnet S, Desjardins L, Turleau C, Doz F (1999) Dysmorphic phenotype and neurological impairment in 22 retinoblastoma patients with constitutional cytogenetic 13q deletion. Clin Genet 55(6):478–82
Bojinova RI, Schorderet DF, Addor MC, Gaide AC, Thonney F, Pescia G, Nenadov-Beck M, Balmer A, Munier FL (2001) Further delineation of the facial 13q14 deletion syndrome in 13 retinoblastoma patients. Ophthalmic Genet 22(1):11–18
Brown S, Gersen S, Anyane-Yeboa K, Warburton D (1993) Preliminary definition of a “critical region” of chromosome 13 in q32: report of 14 cases with 13q deletions and review of the literature. Am J Med Genet 45(1):52–59
Brown S, Russo J, Chitayat D, Warburton D (1995) The 13q-syndrome: the molecular definition of a critical deletion region in band 13q32. Am J Hum Genet 57(4):859–866
Bruce KR, Steiger H, Joober R, Ng Ying Kin NM, Israel M, Young SN (2005) Association of the promoter polymorphism -1438G/A of the 5-HT2A receptor gene with behavioral impulsiveness and serotonin function in women with bulimia nervosa. Am J Med Genet B Neuropsychiatr Genet 137(1):40–44
Dahiya A, Gavin MR, Luo RX, Dean DC (2000) Role of the LXCXE binding site in Rb function. Mol Cell Biol 20(18):6799–6805
Dries D, Baca K, Truss L, Dobin S (2003) Interstitial deletion of 13q and a 13;X chromosome translocation results in partial trisomy 13 and bilateral retinoblastoma. Ophthalmic Genet 24(3):175–180
Hogg A, Onadim Z, Baird PN, Cowell JK (1992) Detection of heterozygous mutations in the RB1 gene in retinoblastoma patients using single-strand conformation polymorphism analysis and polymerase chain reaction sequencing. Oncogene 7(7):1445–1451
Houdayer C, Gauthier-Villars M, Lauge A, Pages-Berhouet S, Dehainault C, Caux-Moncoutier V, Karczynski P, Tosi M, Doz F, Desjardins L, Couturier J, Stoppa-Lyonnet D (2004) Comprehensive screening for constitutional RB1 mutations by DHPLC and QMPSF. Hum Mutat 23(2):193–202
Kloss K, Wahrisch P, Greger V, Messmer E, Fritze H, Hopping W, Passarge E, Horsthemke B (1991) Characterization of deletions at the retinoblastoma locus in patients with bilateral retinoblastoma. Am J Med Genet 39(2):196–200
Li J, Kang C, Wang Y, Zhou R, Wang B, Guan L, Yang L, Faraone SV (2006) Contribution of 5-HT2A receptor gene -1438A > G polymorphism to outcome of attention-deficit/hyperactivity disorder in adolescents. Am J Med Genet B Neuropsychiatr Genet 141(5):473–476
Livak K (1997) ABI Prism 7700 Sequence detection system
Lohmann DR, Gallie BL (2004) Retinoblastoma: revisiting the model prototype of inherited cancer. Am J Med Genet C Semin Med Genet 129(1):23–28
Motegi T, Ikeda K, Watanabe K, Yanagawa Y, Minoda K (1987) Deletion (13)(q13q14.3) with retinoblastoma: confirmation and extension of a recognisable pattern of clinical features in retinoblastoma patients with 13q deletion. J Med Genet 24(11):696–697
Motegi T, Kaga M, Yanagawa Y, Kadowaki H, Watanabe K, Inoue A, Komatsu M, Minoda K (1983) A recognizable pattern of the midface of retinoblastoma patients with interstitial deletion of 13q. Hum Genet 64(2):160–162
Pescucci C, Caselli R, Grosso S, Mencarelli MA, Mari F, Farnetani MA, Piccini B, Artuso R, Bruttini M, Priolo M, Zuffardi O, Gimelli S, Balestri P, Renieri A (2006) 2q24-q31 Deletion: report of a case and review of the literature. Eur J Med Genet
Sampieri K, Hadjistilianou T, Mari F, Speciale C, Mencarelli MA, Cetta F, Manoukian S, Peissel B, Giachino D, Pasini B, Acquaviva A, Caporossi A, Frezzotti R, Renieri A, Bruttini M (2006) Mutational screening of the RB1 gene in Italian patients with retinoblastoma reveals 11 novel mutations. J Hum Genet 51(3):209–216
Scheffer H, Van Der Vlies P, Burton M, Verlind E, Moll AC, Imhof SM, Buys CH (2000) Two novel germline mutations of the retinoblastoma gene (RB1) that show incomplete penetrance, one splice site and one missense. J Med Genet 37(7):E6
Sharp AJ, Locke DP, McGrath SD, Cheng Z, Bailey JA, Vallente RU, Pertz LM, Clark RA, Schwartz S, Segraves R, Oseroff VV, Albertson DG, Pinkel D, Eichler EE (2005) Segmental duplications and copy-number variation in the human genome. Am J Hum Genet 77(1):78–88
Skrypnyk C, Bartsch O (2004) Retinoblastoma, pinealoma, and mild overgrowth in a boy with a deletion of RB1 and neighbor genes on chromosome 13q14. Am J Med Genet A 124(4):397–401
Yamagata K, Andreasson KI, Sugiura H, Maru E, Dominique M, Irie Y, Miki N, Hayashi Y, Yoshioka M, Kaneko K, Kato H, Worley PF (1999) Arcadlin is a neural activity-regulated cadherin involved in long term potentiation. J Biol Chem 274(27):19473–1979
Acknowledgments
This work was supported by grants from Pierfranco e Luisa Mariani Foundation and from Telethon Foundation (GTF05005) to A.R. and by a Grant on Retinoblastoma from Istituto Toscano Tumor (ITT) to A.R.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Caselli, R., Speciale, C., Pescucci, C. et al. Retinoblastoma and mental retardation microdeletion syndrome: clinical characterization and molecular dissection using array CGH. J Hum Genet 52, 535–542 (2007). https://doi.org/10.1007/s10038-007-0151-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-007-0151-4
Keywords
This article is cited by
-
13q mosaic deletion including RB1 associated to mild phenotype and no cancer outcome – case report and review of the literature
Molecular Cytogenetics (2018)
-
Fine mapping of whole RB1 gene deletions in retinoblastoma patients confirms PCDH8 as a candidate gene for psychomotor delay
European Journal of Human Genetics (2013)
-
Genotype–phenotype correlations in patients with retinoblastoma and interstitial 13q deletions
European Journal of Human Genetics (2011)