Abstract
Mutations in the voltage-gated chloride/proton antiporter ClC-5 gene, CLCN5, are associated with Dent’s disease, an X-linked renal tubulopathy. Our interest is to identify and characterize disease-causing CLCN5 mutations, especially those that alter the splicing of the pre-mRNA. We analyzed the CLCN5 gene from nine unrelated Spanish Dent’s disease patients and their relatives by DNA sequencing. Pre-mRNA splicing analysis was performed by RT-PCR. Seven new mutations were identified, consisting of three missense mutations (C219R, F273L, and W547G), one splice-site mutation (IVS-2A > G), one deletion (976delG), and two non-sense mutations (Y140X and W314X). We found that missense mutation W547G also led to increased expression of a new alternative isoform lacking exons 10 and 11 that was expressed in several human tissues. In addition, we describe another novel CLCN5 splicing variant lacking exon 11 alone, which was expressed only in human skeletal muscle. We conclude that missense mutation W547G can also alter the expression levels of a CLCN5 mRNA splicing variant. This type of mutation has not been previously described in the CLCN5 gene. Our results support the importance of a routine analysis at the pre-mRNA level of mutations that are commonly assumed to cause single amino acids alterations.
Similar content being viewed by others
Introduction
Dent’s disease (OMIM 300009) is an X-linked renal tubulopathy characterized by low molecular weight proteinuria (LMWP), hypercalciuria, nephrocalcinosis/nephrolithiasis, and progressive renal failure. The disease may also be associated with aminoaciduria, phosphaturia, glycosuria, kaliuresis, uricosuria, impaired urinary acidification, and can be complicated by rickets or osteomalacia in some patients (Thakker 2000). Inactivating mutations in the CLCN5 gene cause Dent’s disease (Lloyd et al. 1996). Recently, this disorder has also been associated with mutations in the OCRL1 gene (OMIM #300555) that encodes a phosphatidylinositol-4,5-biphosphate-5-phosphatase and is correlated with Lowe syndrome (Hoopes et al. 2005).
CLCN5 encodes the voltage-gated chloride/proton antiporter ClC-5 (746 amino acids). A recently identified splice variant with an additional 70 amino acids at the intracellular amino-terminus has been detected only at the mRNA level (Ludwig et al. 2003; Jentsch et al. 2005). ClC-5 is a dimer with two identical subunits, each of which contains a pore and has an internal structural repeat composed of 18 α-helices (A–R) in antiparallel orientation (see Fig. 1). Helices H, I, P, and Q are involved in the formation of the interface between both subunits (Dutzler et al. 2002), and helices D, F, N, and R form the chloride selectivity filter (Dutzler et al. 2002). The carboxy terminus of ClC-5 includes two CBS (cystathionine β-synthase) domains, which constitute an energy-sensing module binding ATP, a PY sorting signal involved in trafficking to acidic endosomes, and a potential protein binding module, PDZ (Schwake et al. 2001; Scott et al. 2004; Hryciw et al. 2006).
Schematic representation of the ClC-5 monomer structure, showing the exonic mutations identified in this study. Cylinders depict helices A–R, with the cytoplasmic region below, and the extracellular region above. Regions involved in the chloride selectivity filter are shown in light grey. The CBS (cystathionine β-synthase) domains and the PY motif are also indicated
In the human nephron, ClC-5 is expressed in the proximal tubule, the thick ascending limb, and the intercalated cells of the collecting duct. ClC-5 has been localized in subapical endosomes with vacuolar H+-ATPase, suggesting that it may play a role in the ion transport mechanism that facilitates acidification within endosomes (Jentsch et al. 2005). Endosomes are part of the receptor-mediated endocytic pathway that transports proteins such as albumin, and therefore a ClC-5 dysfunction could explain the LMWP in Dent’s disease. It is noteworthy that ClC-5 knockout mice exhibit a phenotype similar to that of Dent’s disease, with LMWP associated with impaired proximal tubule reabsorption of proteins (Jentsch et al. 2005). Another possible role that has been suggested for ClC-5 is mediating the protein–protein interactions that are involved in the assembly of the endocytic complex (Hyrciw et al. 2003).
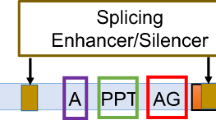
Different types of disease-causing mutations in the CLCN5 gene have been reported (Lloyd et al. 1996; Hoopes et al. 1998; Morimoto et al. 1998; Yamamoto et al. 2000; Carballo-Trujillo et al. 2003; Claverie-Martin et al. 2003; Ludwig et al. 2005). Most of these mutations are single nucleotide substitutions that are scattered throughout the coding sequence of the gene. Point mutations are commonly assumed to cause single amino acids alterations or truncations in the encoded proteins. However, increasing evidence shows that mutations that affect pre-mRNA processing cause a considerable number of genetic diseases and that exonic mutations may also alter the splicing pattern of the pre-mRNA (Pagani and Baralle 2004). Non-sense, missense, and silent mutations, as well as coding single-nucleotide polymorphisms and even intronic variants can inactivate genes by inducing exon skipping (Cartegni et al. 2002). Many splicing mutations affect standard consensus splicing signals, but recent studies have shown that there are mutations that alter other elements needed for regulation of the splicing process, i.e., exonic and intronic splicing enhancers or silencers (Cartegni et al. 2002). Therefore, it has been suggested that, in order to determine the full consequences of a mutation in disease, each mutation should be functionally assessed for its potential effect on splicing (Pagani and Baralle 2004). Until now, only a few CLCN5 mutations affecting pre-mRNA splicing have been described, and only in some cases has the effect been confirmed by mRNA analysis (Morimoto et al. 1998; Yamamoto et al. 2000; Claverie-Martin et al. 2003, 2005; Forino et al. 2004).
In this study, we analyzed the CLCN5 gene in nine Spanish patients with Dent’s disease. We identified seven new mutations consisting of two nonsense mutations, three apparently missense mutations, one deletion and one splice-site mutation. Our results showed that one of the missense mutations also affects the pre-mRNA splicing pattern. In addition, we characterized two novel CLCN5 mRNA splicing variants.
Subjects and methods
Patients
Nine unrelated patients diagnosed with Dent’s disease, from different geographic areas of Spain (Table 1), and 21 of their relatives were investigated after giving informed consent. All patients had LMWP and hypercalciuria, four had nephrocalcinosis and/or nephrolithiasis, and two suffered from rickets/osteopenia. Venous blood samples were obtained from patients and from affected and unaffected family members for mutational analysis of the CLCN5 gene. The protocol was approved by the Ethics Committees of all participating centers.
DNA sequence analysis
DNA was extracted from whole blood by using a DNA Blood Mini kit (Qiagen, Hilden, Germany), and all CLCN5 coding exons and non-coding exons 1a and 1b, including their intronic flanking sequences, were amplified by PCR as described previously (Carballo-Trujillo et al. 2003; Ludwig et al. 2003). Additionally, the OCRL1 gene was screened for mutations in the two patients without apparent mutations in the CLCN5 gene using methods described elsewhere (Hoopes et al. 2005). Thermocycling was performed on a GeneAmp PCR system 9700 (Applied Biosystems, Foster City, CA). The amplified fragments were directly sequenced on both strands using a Big DyeTerminator kit v 3.1 Cycle Sequencing kit and run on an ABI PRISM 310 Genetic Analyzer (Applied Biosystems). The primers used for sequencing were the same primers used for the PCR amplification. DNA mutations were determined by comparison to reference sequences (GenBank accession numbers X91906 for CLCN5 and AL138745 and AL022162 for the 5′ and 3′ ends of OCRL1, respectively), and confirmed by sequencing additional independent amplification products. In addition, in order to demonstrate that mutation C219R was not a common polymorphism in the general population, we designed a primer that annealed adjacent to the affected nucleotide, creating a restriction site for BsmAI only if the mutation was present (5′-GCCCTCTAGTGCACGTGTCT-3′), and amplified a 122 bp fragment with the reverse primer used previously for exon 6. A restriction fragment length polymorphism (RFLP) screen of 100 chromosomes from normal unrelated controls (20 males and 40 females) was performed following the manufacturer's recommendations (not shown).
Pre-mRNA splicing analysis
Total RNA was isolated from whole blood samples using the PAXgene Blood RNA System (Qiagen), digested with RNase-free DNase I (Clontech, Palo Alto, CA) and reverse transcribed with random hexamers using a First-strand cDNA synthesis kit (Roche, Basel, Switzerland). One microliter of cDNA was then used as a template for PCR amplifications to study the effect of the mutations on pre-mRNA splicing. Two segments of CLCN5 cDNA, from exon 2 to exon 5 and from exon 8 to exon 12, respectively, were amplified using primers and conditions described previously (Claverie-Martin et al. 2005).
Alternative isoforms of CLCN5 were studied by RT-PCR using total RNA isolated from whole blood of a healthy individual and poly-A mRNA (BD Biosciences Clontech, Palo Alto, CA) from normal human tissues (kidney, lung, pancreas, liver, brain, and skeletal muscle), and from mouse kidney. Poly-A mRNA (200 ng) was reverse transcribed as described above. Amplifications were carried out using previously described conditions (Claverie-Martin et al. 2005). To study the coding region in the splicing variant, a primer (5′-ATTCCAAGCAATCGCCTAAG-3′) was designed to specifically detect skipping of exons 10 and 11. After testing the specificity of the primer, it was used along with primer CLCN5cDN 2F (5′-GACTTCTTGGAGGAGCCAATC-3′) (Claverie-Martin et al. 2005) to amplify the complete coding region (starting in exon 2) in all tissues. The program for this amplification was 94°C for 2 min, followed by 40 cycles each consisting of three steps, 94°C for 30 s, 60°C for 45 s, and 72°C for 60 s, with a final extension at 72°C for 5 min. In all cases, bands were excised and purified for sequencing using a gel extraction kit (Qiagen). The primers used for sequencing were the same primers used for PCR amplification.
Results and discussion
Nine patients presenting with LMWP and hypercalciuria and their family members were analyzed for possible mutations in the CLCN5 gene (Fig. 2; Table 1). The sequenced DNA revealed the presence of seven single nucleotide mutations not previously reported (Fig. 2; Table 1): one splicing mutation and six exonic variants, consisting of two nonsense mutations, three apparently missense mutations and one deletion (Table 1). Two patients presenting with symptoms resembling Dent’s disease did not show any mutation in CLCN5, as has been observed by others (Morimoto et al. 1998; Hoopes et al. 2005). We also performed a sequence analysis of the OCRL1 gene in these two patients, but no mutations were found (not shown).
Segregation of CLCN5 mutations. Pedigrees of the respective Spanish families are shown with phenotypic findings in males (squares) and females (circles) shown as follows: filled circle female carrier, filled square affected male, open circle unaffected female, and open square unaffected male. Relatives not included in the DNA sequence analysis are indicated by black arrows
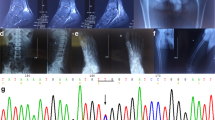
We identified a missense mutation, W547G, which is located at the beginning of exon 10 and is predicted to lead to the substitution of a tryptophan residue in helix Q (Fig. 1), which is involved in the formation of the interface between both ClC-5 subunits. Residue W547 is conserved throughout mammalian ClC-5 proteins but not in the ClC family (results not shown). There is increasing evidence that missense mutations also alter pre-mRNA splicing, and we are especially interested in characterizing this type of mutation in the CLCN5 gene. Therefore, we examined the RNA from blood of this patient by RT-PCR analysis and found that, after amplifying from exon 8 to exon 12, two different splicing products were found: the expected 985-bp product from normal pre-mRNA processing together with a shorter product of 369 bp of similar intensity suggestive of an exon skipping event. Sequencing of the 369-bp fragment confirmed that it lacked exons 10 and 11, and contained exons 9 and 12 precisely spliced together (Fig. 3a). This leads to a change in the open reading frame, generating 11 new codons (DCLESLPKRMC) at the 3′ end of residue 511, followed by a premature termination codon. However, mRNA molecules carrying premature stop codons are expected to be degraded through nonsense-mediated mRNA decay, resulting in decreased expression of the protein. The normal spliced mRNA would be translated into a mutant protein with a glycine instead of a tryptophan at position 547. Therefore, the phenotype of this patient would be due to the combined effect of decreased ClC-5 expression and the presence of a mutant ClC-5 protein. To determine if the spliced form lacking exons 10 and 11 was present in normal tissues, we PCR amplified cDNAs from human kidney, muscle, brain, pancreas, liver, lung, and lymphocytes. This experiment revealed that the alternative product was present in all human tissues examined, albeit in small amounts (Fig. 3b). This product was not detected in mouse kidney when we used either human primers (not shown) or mouse-specific primers (Fig. 3b). In addition, a 768-bp band corresponding to the skipping of exon 11 alone, was observed in human skeletal muscle (Fig. 3b). In an attempt to characterize the 5′ coding part of this alternative CLCN5 mRNA, we amplified the complete coding region spanning from exon 2 to the junction between exons 9 and 12, in all human tissues tested. RT-PCR analysis showed no evidence of additional splicing products other than the skipping of exons 10 and 11 (not shown). The loss of these two exons predicts a truncated protein without the transmembrane helices O, P and Q (P and Q are involved in the formation of the interface between both subunits), the cytoplasmic helix R (a component of the chloride selectivity filter) and the carboxy-terminus of the ClC-5 protein.
RT-PCR analysis and DNA sequence of the cDNA products. a Effect of mutation W547G in CLCN5 pre-mRNA splicing in the patient’s lymphocytes. Lanes P and C show the products from the patient and a normal control, respectively. b Expression pattern of CLCN5 isoforms from different human tissues. Poly-A mRNA from mouse kidney and human genomic DNA, gDNA, were used as controls. c Effect of mutation IVS2-2A > G in CLCN5 pre-mRNA splicing in the patient’s lymphocytes. Lanes P, M and C show the products from the patient, the patient’s mother and a normal control, respectively. Artifacts generated in the RT-PCR are indicated by a star
Our results suggest that mutation W547G leads to an alteration in the expression levels of the alternative isoform lacking exons 10 and 11 (Fig. 3a). The mechanism by which exons 10 and 11 are skipped simultaneously is not known. It has been reported that some mutations alter cis-elements needed for correct splicing (Cartegni et al. 2002). We analyzed mutation W547G using ESE-finder (Cartegni et al. 2002) and found that it does not predict the disruption of any exonic splicing enhancer (ESE). However, ACESCAN2 (Yeo et al. 2005) analysis showed that the T to G substitution leads to the creation of four overlapping exonic splicing silencers (ESS) elements (AAGGGG, AGGGGG, GGGGGT, and GGGGTG) within the exon 10 sequence. ESSs are known to suppress the recognition of exons in splice site definition. Recent analysis of alternative splicing processes in Drosophila and humans supports the hypothesis that splice-site recognition is more efficient across the intron, and that alternative splicing is less likely for exons flanked by short introns (Fox-Walsh et al. 2005). Notably, splice-site recognition across the intron results in enhanced inclusion of exons with weak splice sites. By using either GENESCAN (http://www.genes.mit.edu/GENSCAN.html) or the splice sites predictor (http://www.fruitfly.org/seq_tools/splice.html) we found that CLCN5 exon 11 has a low matching score compared to the consensus acceptor site sequences (data not shown). This, together with the small size of intron 10 (155 bp) could explain the simultaneous skipping of exons 10 and 11 in the presence of the W547G mutation. The creation of the new ESS elements may decrease exon 10 incorporation into the mature mRNA, and the subsequent loss of cis elements that may reside within exon 10 could disable the splicing machinery for cross-intron recognition of exon 11, resulting in increased skipping of both exons. In summary, the missense mutation W547G can also alter the expression levels of a CLCN5 mRNA splicing variant. This type of mutation has not been previously described in the CLCN5 gene. Previously, Yamamoto et al. (2000) reported a CLCN5 missense mutation, D601V, which also results in the creation of a new donor splice site and leads to the skipping of part of exon 10.
Another two new missense mutations were found in this study: C219R and F273L. Mutation C219R, located on exon 6, affects ClC-5 helix F (Fig. 1), which is one of the helices involved in the formation of the chloride selectivity filter, and would lead to a change in the amino acid polarity. Ludwig et al. (2005) used functional studies in Xenopus oocytes to demonstrate that the C221R missense mutation, also located in helix F, not only interfered with protein function but also abolished ClC-5 trafficking to the plasma membrane (Ludwig et al. 2005). C219 is a conserved amino acid among mammalian ClC-5 homologues, but it is not located in the conserved domain GREGP, which is found at the amino terminus of helix F and is directly involved in anion binding (Estevez and Jentsch 2002), or in regions of high homology described for the ClC family (Dutzler et al. 2002). The third missense mutation, F273L, is located at the beginning of exon 8 and introduces a change in the loop between helix H and I (Fig. 1). Residue F273 is conserved between mammalian ClC-5 proteins and also in the ClC family. These changes in conserved amino acids probably affect essential properties of the protein.
In one of our patients, we identified a mutation located at the acceptor splice site of intron 2, IVS2-2A > G, which disrupted the highly conserved AG dinucleotide sequence. As expected, RT-PCR analysis of the patient’s RNA showed an aberrant CLCN5 pre-mRNA splicing product of 340 bp instead of the normal 440-bp product (Fig. 3c). Sequence analysis of the smaller fragment revealed skipping of the entire exon 3, which results in a change in the reading frame that introduces two new codons (VR) and a premature termination codon in exon 4. RNA from the patient’s mother was also analyzed and showed both normal and aberrant products (Fig. 3c). Simultaneous expression of both splicing forms in the mother can be explained by the fact that the multigene domain in Xp11.21-p11.22, where CLCN5 gene is located, has been shown to escape X inactivation (Miller and Willard 1998).
We also identified two new nonsense mutations: Y140X, which is located in exon 5 and predicts a disruption of helix D, one of the transmembrane domains, and W314X, found in exon 8, which introduces a disruption in the loop between helix I and J (Fig. 1). A novel single-base deletion, 976delG in codon 325 of exon 8, was also identified, resulting in a frame-shift leading to the addition of 32 new amino acids (AYLVVCGEHCYSAQTLPGVGSERPPSWASILL) and the premature termination of translation at codon 358. These three mutations would be expected to inactivate channel function, either by truncation or by degradation through nonsense-mediated mRNA decay.
In conclusion, we identified seven new CLCN5 mutations that cause structural alterations in the ClC-5 protein and are likely to result in a loss of function. Missense mutation W547G was shown to alter the pre-mRNA splicing pattern. In addition to expanding the spectrum of CLCN5 mutations associated with this renal tubulopathy, our study reveals two new CLCN5 splicing variants, and supports the importance of a routine analysis at the RNA level of mutations that are commonly assumed to cause single amino acids alterations. The ideal source of RNA to study the effect of mutations on CLCN5 pre-mRNA splicing would be kidney biopsies from patients. However, as this source is not always available, one can use RNA obtained from lymphocytes, which has been shown to be valid for this type of study (Forino et al. 2004; Claverie-Martin et al. 2005).
References
Carballo-Trujillo I, Garcia-Nieto V, Moya-Angeler FJ, Anton-Gamero M, Loris C, Mendez-Alvarez S, Claverie-Martin F (2003) Novel truncating mutations in the ClC-5 chloride channel gene in patients with Dent’s disease. Nephrol Dial Transplant 18:717–723
Cartegni L, Chew SL, Krainer AR (2002) Listening to silence and understanding nonsense: exonic mutations that affect splicing. Nat Rev Genet 3:285–298
Claverie-Martin F, Gonzalez-Acosta H, Flores C, Anton-Gamero M, Garcia-Nieto V (2003) De novo insertion of an Alu sequence in the coding region of the CLCN5 gene results in Dent’s disease. Hum Genet 113:480–485
Claverie-Martin F, Flores C, Anton-Gamero M, Gonzalez-Acosta H, Garcia-Nieto V (2005) The Alu insertion in the CLCN5 gene of a patient with Dent’s disease leads to exon 11 skipping. J Hum Genet 50:370–374
Dutzler R, Campbell EB, Cadene M, Chait BT, MacKinnon R (2002) X-ray structure of a ClC chloride channel at 3.0 Å reveals the molecular basis of anion selectivity. Nature 415:287–294
Estevez R, Jentsch TJ (2002) CLC chloride channels: correlating structure with function. Curr Opin Struct Biol 12:531–539
Forino M, Graziotto R, Tosetto E, Gambaro G, D’Angelo A, Anglani F (2004) Identification of a novel splice site mutation of CLCN5 gene and characterization of a new alternative 5′ UTR end of ClC-5 mRNA in human renal tissue and leukocytes. J Hum Genet 49:53–60
Fox-Walsh KL, Dou Y, Lam BJ, Hung SP, Baldi PF, Hertel KJ (2005) The architecture of pre-mRNAs affects mechanisms of splice-site pairing. Proc Natl Acad Sci USA 102:16176–16181
Hoopes RR Jr, Hueber PA, Reid RJ Jr, Braden GL, Goodyer PR, Melnyk AR, Midgley JP, Moel DI, Neu AM, VanWhy SK, Scheinman SJ (1998) CLCN5 chloride-channel mutations in six new North American families with X-linked nephrolithiasis. Kidney Int 54:698–705
Hoopes RR Jr, Shrimpton AE, Knohl SJ, Hueber P, Hoppe B, Matyus J, Simckes A, Tasic V, Toenshoff B, Suchy SF, Nussbaum RL, Scheinman SJ (2005) Dent disease with mutations in OCRL1. Am J Hum Genet 76:260–267
Hryciw DH, Wang YH, Devuyst O, Pollock CA, Poronnik P, Guggino WB (2003) Cofilin interacts with ClC-5 and regulates albumin uptake in proximal tubule cell lines. J Biol Chem 278:40169–40176
Hryciw DH, Ekberg J, Ferguson C, Lee A, Wang D, Parton RG, Pollock CA, Yun CC, Poronnik P (2006) Regulation of albumin endocytosis by PSD95/Dlg/ZO-1 (PDZ) scaffolds. Interaction of Na+–H+ exchange regulatory factor-2 with ClC-5. J Biol Chem 281:16068–16077
Jentsch TJ, Maritzen T, Zdebik AA (2005) Chloride channel diseases resulting from impaired transepithelial transport or vesicular function. J Clin Invest 115:2039–2046
Lloyd SE, Pearce SH, Fisher SE, Steinmeyer K, Schwappach B, Scheinman SJ, Harding B, Bolino A, Devoto M, Goodyer P, Rigden SP, Wrong O, Jentsch TJ, Craig IW, Thakker RV (1996) A common molecular basis for three inherited kidney stone diseases. Nature 379:445–449
Ludwig M, Waldegger S, Nuutinen M, Bokenkamp A, Reissinger A, Steckelbroeck S, Utsch B (2003) Four additional CLCN5 exons encode a widely expressed novel long CLC-5 isoform but fail to explain Dent’s phenotype in patients without mutations in the short variant. Kidney Blood Press Res 26:176–184
Ludwig M, Doroszewicz J, Seyberth HW, Bokenkamp A, Balluch B, Nuutinen M, Utsch B, Waldegger S (2005) Functional evaluation of Dent’s disease-causing mutations: implications for ClC-5 channel trafficking and internalization. Hum Genet 117:228–237
Miller AP, Willard HF (1998) Chromosomal basis of X chromosome inactivation: identification of a multigene domain in Xp11.21-p11.22 that escapes X inactivation. Proc Natl Acad Sci USA 95:8709–8714
Morimoto T, Uchida S, Sakamoto H, Kondo Y, Hanamizu H, Fukui M, Tomino Y, Nagano N, Sasaki S, Marumo F (1998) Mutations in CLCN5 chloride channel in Japanese patients with low molecular weight proteinuria. J Am Soc Nephrol 9:811–818
Pagani F, Baralle FE (2004) Genomic variants in exons and introns: identifying the splicing spoilers. Nat Rev Genet 5:389–396
Schwake M, Friedrich T, Jentsch TJ (2001) An internalization signal in ClC-5, an endosomal Cl-channel mutated in Dent’s disease. J Biol Chem 276:12049–12054
Scott JW, Hawley SA, Green KA, Anis M, Stewart G, Scullion GA, Norman DG, Hardie DG (2004) CBS domains form energy-sensing modules whose binding of adenosine ligands is disrupted by disease mutations. J Clin Invest 113:274–284
Thakker RV (2000) Pathogenesis of Dent’s disease and related syndromes of X-linked nephrolithiasis. Kidney Int 57:787–793
Yamamoto K, Cox JP, Friedrich T, Christie PT, Bald M, Houtman PN, Lapsley MJ, Patzer L, Tsimaratos M, Van’T Hoff WG, Yamaoka K, Jentsch TJ, Thakker RV (2000) Characterization of renal chloride channel (CLCN5) mutations in Dent’s disease. J Am Soc Nephrol 11:1460–1468
Yeo GW, Van Nostrand E, Holste D, Poggio T, Burge CB (2005) Identification and analysis of alternative splicing events conserved in human and mouse. Proc Natl Acad Sci USA 102:2850–2855
Acknowledgments
We are grateful to the patients and their relatives for their participation in this study. This investigation was supported by grants PI 04/2620 from Fondo de Investigación Sanitaria and PI 57/04 from Fundación Canaria de Investigación y Salud FUNCIS to F.C.M. and V.G.N. C.F. had a postdoctoral fellowship from FUNCIS.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ramos-Trujillo, E., González-Acosta, H., Flores, C. et al. A missense mutation in the chloride/proton ClC-5 antiporter gene results in increased expression of an alternative mRNA form that lacks exons 10 and 11. Identification of seven new CLCN5 mutations in patients with Dent’s disease. J Hum Genet 52, 255–261 (2007). https://doi.org/10.1007/s10038-007-0112-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-007-0112-y
Keywords
This article is cited by
-
Genetics and phenotypic heterogeneity of Dent disease: the dark side of the moon
Human Genetics (2021)
-
Complexity of the 5′UTR region of the CLCN5gene: eleven 5′UTR ends are differentially expressed in the human kidney
BMC Medical Genomics (2014)
-
Dent’s disease: clinical features and molecular basis
Pediatric Nephrology (2011)