Abstract
Hereditary hemorrhagic telangiectasia (HHT) is an autosomal dominant disorder causing vascular dysplasias. About 70–80% of HHT patients carries mutations in ENG or ACVRL1 genes, which code for a TGFβ receptor type III and I respectively. Molecular data on a large cohort of Italian HHT patients are presented, discussing the significance of missense and splice site mutations. Mutation analysis in ENG and ACVRL1 genes was performed using single strand conformation polymorphisms (SSCP), denaturing high performance liquid chromatography (DHPLC) and subsequent direct sequencing. Overall, 101 mutations were found, with ACVRL1 involved in 71% of cases. The highest number of mutations (28/101 subjects, 14/76 different mutations referring to both genes) was in ACVRL1, exon 3. Mutation analysis was then extended to a total of 356 family members, and 162 proven to carry the mutation. New polymorphisms were identified in both genes, and evidence that ENG P131L change is not a disease-causing mutation was also provided. An in silico analysis was performed in order to characterize splice-site mutations. These results were compared to other European national studies and data from Italy, France and Spain were consistent for an higher incidence of ACVRL1 mutations.
Similar content being viewed by others
Introduction
Hereditary hemorrhagic telangiectasia (HHT; Mutation Database: http://137.195.14.43/cgi-bin/WebObjects/hht.woa/wa/default) is an autosomal dominant disorder causing vascular dysplasias such as mucocutaneous telangiectases and arterovenous malformations (AVMs). Telangiectases may lead to epistaxes and gastrointestinal bleeding, which may be severe enough to require transfusions. Epistaxes and telangiectases are the most frequent symptoms, present in more than 95% of the patients. AVMs are mostly observed in liver (60%), lungs (18–70%) and brain (6%), and may cause severe life-threatening complications (Lesca et al. 2007). The phenotype is highly variable, even among members of the same family, and the disease displays age-related penetrance, with increased manifestations developing over a lifetime. About 70–80% of HHT patients carry mutations in either of two genes—ENG (OMIM #131195) (HHT1: OMIM 187300) or ACVRL1 (OMIM #601284) (HHT2: OMIM 600376)—which code for a TGFβ receptor type III and I respectively, although David et al. 2007 recently demonstrated that BMP9 is the more effective ligand of ACVRL1. Evidence for a third and a fourth locus on chromosomes 5 and 7 respectively has also been reported (Cole et al. 2005; Bayrak-Toydemir et al. 2006a). Association of the HHT phenotype with juvenile polyposis (JPHT, #175050) and mutations in the MADH4 gene (coding for SMAD4, involved in the TGFβ signaling pathway) have recently been demonstrated as well (Gallione et al. 2004). Moreover, MADH4 mutations may be observed in HHT patients without a prior diagnosis of JP (Gallione et al. 2006).
Several papers have recently been published reporting the results of screening for mutations in ENG and ACVRL1 genes, and more than 200 pathogenetic variations have been reported for each gene.
Many polymorphisms are also present in both genes (HHT Mutation database: www.hhtmutation.org). A peculiar distribution of mutations has been reported: ENG mutations are more frequently found in patients from Northern Europe and the Americas, while Mediterranean populations have a majority of ACVRL1 mutations (Olivieri et al. 2002; Brusgaard et al. 2004; Lesca et al. 2004, 2006; Abdalla et al. 2005; Kjeldsen et al. 2005; Letteboer et al. 2005, 2006; Bayrak-Toydemir et al. 2006b; Fernandez-L et al. 2006; Lenato et al. 2006; Bossler et al. 2006).
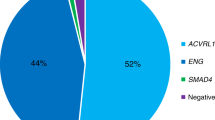
Here we report the results of our screening for mutations in ENG and ACVRL1 genes in a group of 137 Italian HHT patients. Overall, we found a mutation in 101 subjects (76 different mutations), with ACVRL1 involved in 71% of our cases. We also identified several new polymorphisms in both genes, and provide evidence that ENG P131L change is not a disease-causing mutation.
Materials and methods
Patients
We screened a total of 137 index cases; 123 with a known family history, and 14 sporadic cases (family SLL in which the ENG P131L was found is not included in the figures). For each family, we selected a single case in whom the diagnosis was “definite”, according to the diagnostic criteria suggested by the International HHT Advisory board, known as “Curaçao criteria” (Shovlin et al. 2000). After the identification of a mutation, all other available relatives were also studied.
In 117 of them the diagnosis was made by EB in Crema (CR) and by FP, LS, SC and CD in Pavia, and the clinical data were shared between both centres. In 20 cases, the diagnoses were made in other centres who were certainly informed about the diagnostic criteria. Six families were of Croatian, Dutch, Hungarian, Turkish and Indian origin. Detailed data as to the geographical origin of the Italian families were also recorded.
Molecular analysis
After informed consent (prior to inclusion in the study), DNA was extracted from 3 to 7 ml of peripheral blood from each subject, using standard protocols.
Exons (including intron-exon boundaries) 1–13 of ENG gene and 1–10 of ACVRL1 gene were amplified by PCR using the primers reported in the GDB database (Human Genome Database. The Official World-Wide Database for the Annotation of the Human Genome. http://www.gdb.org) (ENG 1–12, including 9a and 9b) or published by Berg et al. 1997 (ACVRL1 2–10). Exon 13 of ENG was split into three different PCRs in order to completely cover this region. Primers for these PCRs were designed by C. O. using the software on the web PRIMER3INPUT (Access date is May 25th, 2007 for all these softwares, http://frodo.wi.mit.edu/cgi-bin/primer3/primer3_www.cgi) and are available on request, as are the amplification protocols for the 26 different PCRs.
Mutation screening was performed by SSCP (ACVRL1 gene) and/or DHPLC (ACVRL1 and ENG genes) analysis. At least two different SSCP were performed, using different polyacrilammide concentration (from 6 to 8%, acrilammide bisacrilammide ratio: 19:1; 29:1; 37,5:1) as suggested by Ravnic-Glavač et al (1994). The DHPLC was performed on a WAVE DNA fragment analysis system (Transgenomic, Cheshire, UK). The temperature of the oven for optimal heteroduplex separation at partial DNA sequence was obtained with the Wavemaker 4.0 software (Transgenomic). The DHPLC optimal temperatures are available on request.
Each time an abnormal pattern was found, direct sequencing was performed using the Big Dye terminator method combined with Taq FS (cycle sequencing reaction). Sequence was analysed on a ABI-PRISM 3700 DNA analyser (Applied Biosystems).
Each mutation was confirmed at least by a second sequence. When an unpublished missense mutation was found, we analysed 50 control DNAs and, when possible, we confirmed the cosegregation of the mutation with the disease in the family. We also checked if the amino acid was conserved in orthologous genes.
When a mutation was found in an “index case”, all other subjects belonging to his/her family were analysed using direct sequencing or endonuclease restriction digestion. The relatives (affected and non-affected) were all informed about the results, and genetic counselling was provided.
Splice site scores
To evaluate if mutations found in the intron-exon boundary region affected the splice site, we performed four different analysis using software for splice site scores calculation present on the web:
-
SCORE 1: Splice site analyzer tool: http://ast.bioinfo.tau.ac.il/SpliceSiteFrame.htm
-
SCORE 2: Alex Dong Li’s splice site score calculator (v 0.1): http://www.genet.sickkids.on.ca/∼ali/splicesitescore.html
-
SCORE 3: Splice site score calculation: http://rulai.cshl.edu/new_alt_exon_db2/HTML/score.html
-
SCORE 4: MaxEntScan::score5ss for human 5′ splice sites: http://genes.mit.edu/burgelab/maxent/Xmaxentscan_scoreseq.html
-
MaxEntScan::score3ss for human 3′ splice sites: http://genes.mit.edu/burgelab/maxent/Xmaxentscan_scoreseq_acc.html
All these tools are based on the paper by Shapiro and Senapathy (1987), Carmel et al. (2004) and Yeo and Burge (2004).
Access date is May 25th, 2007 for all these software programs.
Results
Mutations and polymorphisms identified in our study are summarized in Tables 1 and 2 respectively. Nomenclature follows suggestions reported by den Dunnen and Antonarakis (2000). GenBank RefSeq-file accession number are NM_000118.1 for ENG and NM_000020.1 for ACVRL1. Nucleotide numbering uses the A of the ATG translation initiation start site as nucleotide +1.
In ENG we identified 26 different mutations in 29 subjects; 13 are, to the best of our knowledge, unpublished.
We found 12 nucleotide substitutions, 12 deletions and two insertions. As to their effects, 16/26 (61.5%) were truncating mutations, 5/26 (19.3%) were splice-site mutations, 4/26 (15.4%) were missense mutation and 1/26 (3.8%) was an in-frame deletion of 7 amino acids. Results of splice-site scores for these last mutations are reported in Table 3.
Mutations 6 and 8 were shared by three and two families respectively (see Table 1); only for the former could a common geographical origin be recorded.
We also identified nine different polymorphisms; two of them were, to the best of our knowledge, unpublished.
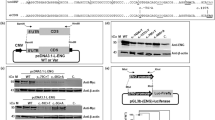
As to P131L, we identified a family in which two subjects were homozygous for this DNA change. The index case, a 9-year-old child, presented with epistaxes, which were present although limited to childhood also in the father and in two paternal uncles. Some cutaneous atypical telangiectases were also present. This phenotype did not segregate with either homozygosity or heterozygosity for the P131L variant. No other ENG or ACVRL1 mutation was found in this family, and it was therefore excluded from any calculation.
In ACVRL1, we identified 50 different mutations in 72 subjects; 11 are, to the best of our knowledge, unpublished.
The mutations can be classified as nucleotide substitution (36), deletions (8) and insertions/duplications (6).
As to their effect, 26/50 (52%) were amino-acid replacement, 20/50 (40%) were truncating mutations, 2/50 (4%) were deletions in frame of 2 amino acids and 2/50 (4%) affected a splice site (see Table 1). All the missense mutations involved conserved amino acids, when the sequence of orthologous genes were compared (Rattus, Bos, Xenopus, Mouse, and Gallus, data not shown). The results of splice site scores for the change-affecting splice sites are reported in Table 3.
Ten mutations (nos. 31,34,35,37,41,44,55,62,68,70 in Table 1) were found in more than one index case; all families carrying the same mutation were unrelated going back for at least four generations, but shared a common geographic origin, with the exception of mutation no. 44 (Table 1): ID1 is a sporadic case and ID287, ID701 and ID705 have a different geographical origin.
We identified six intronic polymorphisms (outside splicing regions); four of them are unpublished and were found in a small proportion of patients (<5%); we also determined the prevalence of g.IVS3 + 11 c > t polymorphisms (Olivieri et al. 2002) (see Table 2) which was in Hardy-Weinberg equilibrium.
In one of our patients, a history of colon cancer at age 34 suggested MADH4 involvement; mutation analysis performed by C. Gallione identified a previously unreported MADH4 mutation (R361L). The son of the index case carries the same mutation, shows a typical HHT clinical picture, but has no evidence of polyposis at age 19 (as assessed by colonoscopy). The healthy mother of the index case, as expected, does not carry the mutation; the father died because of a pancreatic cancer at age 56, and no biological samples were available for analysis.
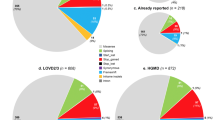
We then compared our results to the screenings recently published by other groups on different European populations [Germany (Kuehl et al. 2005; Schulte et al. 2005; Wehner et al. 2006), Spain (Fernandez-L et al. 2006), Denmark (Brusgaard et al. 2004), France (Lesca et al. 2004, 2006), Netherlands (Letteboer et al. 2005)] and to the work of Lenato et al. 2006 on a smaller group of Italian patients (see Figs. 1,2).
The distribution of the mutations in the two genes [number of ENG mutations/total number of mutations] varied from 37% in Spain to 54% in the Netherlands. When considering the Italian population, ENG mutations were found in 29 % in our cohort and in 40% in the study by Lenato et al. 2006.
ENG mutations are almost evenly distributed in the various exons coding for the extracellular domain, while the ACVRL1 gene shows clustering of mutations in exons 3, 7, 8, and this uneven distribution is confirmed in all the European samples studied. Conversely, exon 5 is consistently less involved in all published reports.
Discussion
The analysis of a group of 137 index patients affected with HHT allowed us to identify 76 different mutations in 101 subjects, 24 unpublished; 162 out of the 356 family members subsequently tested were proven to be mutation carriers.
All but two of the cases (one Indian, one Swedish) were of Italian ancestry (see Table 1). On the basis of these data and of other recent reports, the number of different known mutations arises to over 200 for each gene; a further increase in the number of new mutations in both ENG or ACVRL1 is to be expected.
A “founder effect” for specific mutations observed with high frequency was suggested in the Netherlands Antilles (ENG: c.1238G > T; g.IVS1 + 1 g > a; Gallione et al. 2000), France (ACVRL1: c.1112_1113 dupG; Lesca et al. 2004) and Denmark (ENG: c.360C > A; Brusgaard et al. 2004).
We observed a small cluster in the North Italian county of Bergamo (ACVRL1: c.289 del CACAAC, p. H97-N98 del in frame; Table 1, mutation 41) with the same mutation in ten apparently unrelated families with 40 affected subjects, from a population for the county of 1,021,700 in 2004. Other HHT families with different mutations are known in the same area.
This cluster may partially explain the higher percentage of ACVRL1 mutation in our cohort as compared to the results of Lenato et al. 2006.
We also identified several polymorphisms in both genes by DHPLC analysis, a method that does not allow homozygote identifications; as most of them are intronic variations, outside splicing sites, and thus are unlikely to have any effect on the protein, we did not check if these polymorphisms were in Hardy-Weinberg equilibrium.
The Endoglin P131L substitution was reported as a HHT-causing mutation (Kjeldsen et al. 2005; Fernandez-L et al. 2006; Cymerman et al. 2003) and was subsequently listed as a variant (Abdalla and Letarte 2006). Homozygous Endoglin null mice have an embryonic lethal phenotype (Arthur et al. 2000), and a previous report showed no live children homozygous for ENG mutation in a consanguineous family where both parents were affected (Karabegovic et al. 2004). Our results (no segregation with phenotype, presence of two homozygotes) provide conclusive evidence that P131L is probably a polymorphism whose biological effect, if any, is still to be assessed.
The ENG protein contains a large extracellular domain (encoded by exons 1–12), a transmembrane and an intracellular domain (exons 13 and 14). The mutations are distributed almost evenly in all exons with the exception of a clear lower involvement of exons 1 and 12 to 14 (HHT mutation database; Wehner et al. 2006); our data support this distribution (Fig 1).
Most mutations result in a truncated protein (61.5% in our sample), which is in agreement with the model of haploinsufficiency proposed for the pathogenesis of the disease (Abdalla and Letarte 2006). This model is supported by studies in KO mouse (Arthur et al. 2000; Bordeau et al. 2000) and by the demonstration of a 50% level of protein in HUVEC HHT1 cells (Pece et al. 1997; Cymerman et al. 2000).
A number of DNA changes resulting in amino acid variation but interpreted as polymorphisms have also been reported as [MTHFR, A222V]. A high population frequency of the observed change “per se” cannot be held as a demonstration that the variation has no effect on the protein function and in turn on the general phenotype of the subject carrying it (Paquet et al. 2001). For instance, the [MTHFR, A222V] will result in a protein with definitely modified behaviour, whose presence is associated with vascular disorders and other diseases.
Endoglin polymorphisms resulting in amino acid changes and not causing HHT (as a second obviously pathogenetic mutation in the same gene or in ACVRL1 gene is concurrently present) should thus be reported and studied to demonstrate if they have any effect on protein function. This situation is observed in cases ID170, ID131 and ID300, in which a ACVRL1 mutation was associated to the ENG G191D polymorphism. Until this datum is obtained, we cannot formally exclude the possibility that (minor) changes in protein activity may exert an addictive effect on the HHT phenotype or may be a risk factor for different phenotypes.
The ALK1 protein consists of a small extracellular domain (encoded by exons 2,3, partially 4), a short transmembrane domain (exons 4,5 partially), and a large intracellular domain including a glycine-serine rich (GS) domain (exon 5) and the serine/threonine kinase (S/T-K) domain (exons 5 to 10).
The mutations are unevenly distributed, with exons 3,7,8 carrying a larger number of mutations (61.46% of the total number of ACVRL1 mutations, as calculated from the HHT mutation database). This trend is observed in all European countries, and also in our cohort of cases, with the only peculiarity of a higher number of mutations in exon 3.
In fact in our group of HHT2 patients, we found 28/72 subjects (39%) to be carriers of exon 3 mutations; if only the different ACVRL1 mutations are considered, exon 3 still contains 14/50 mutations (28%), versus 22% in Europe and in the database.
In exon 5, only 5 mutations are known; among them, the only mutation identified in the GS domain (Harrison et al. 2003) was found in a patient with PAH but, apparently, without clinical signs or family history of HHT. Other mutations in exon 5 generate a truncated protein or are in the S/T-K domain, as our mutation 49 in Table 1.
For two mutations in the S/T-K domain (S333I and C344Y), Gu et al. (2006) suggested a dominant negative effect, on the basis of the evaluation of ALK1 expression and activity in monkey and human cell lines. We found a C344F mutation (Table 1, n.58) in which cysteine is replaced by phenylalanine instead of tyrosine as for the study by Gu et al.; the two substituted amino acids are structurally similar, and the hypothesis of a dominant negative may thus be considered for our mutation too. Moreover, the same authors demonstrated that the R411Q mutation results in a protein which retains its activity, although at a reduced level.
Very interestingly, Fernandez-L et al. (2005) demonstrated on BOECs (Blood Outgrowth Endothelial Cells) from HHT2 patients carrying the R374W missense mutation an ALK1 amount similar to that of controls by western blot analysis; this result suggests that some missense mutations may be expressed. Both the R374W and the R411Q are present in our cohort (Table 1, mutation 62 and 70).
The use of the SIFT Tool allowed Prigoda et al. (2006) to show that the vast majority of, but not all, ALK1 missense mutations lead to altered structures, which might imply loss of function.
Altogether, these data indicate that the pathogenesis of HHT2 may be more complex than previously thought; haploinsufficiency may not be the only disease-causing mechanism, and different mutations may lead to (slightly) different phenotypes.
In vivo studies and the development of functional biological tests will settle this point.
In the ACVRL1 gene we did not find any polymorphism resulting in amino acid change, in agreement with previously published data.
We found seven different DNA changes (two in ACVRL1) occurring at a splice site; as all of our work was performed on DNA samples, and data on the expression were not available, we studied these changes in silico using four different tools (Table 3). They were considered as disease-causing mutations when segregating with the disease in the family, and when the power of the splicing was reduced to an extent similar to the result reported by Roca et al. 2003.
A mutation in either ENG or ACVRL1 was found in 75.6% of the Italian index cases and in 2/6 of the non-Italian patients; in addition, 1 MADH4 mutation was found in a family selected because of the phenotype of the index case. The reasons for failure in identification of a mutation in a proportion of the remainder of cases have been extensively and clearly discussed by Lesca et al. (2004) and include technical limits and genetic heterogeneity.
The comparison of our data with those of other European countries indicates that Italy, Spain and France homogeneously show an higher percentage of ACVRL1 mutations with a narrow range: 60–62%. In other countries such as Germany, Denmark and The Netherlands, the percentage of HHT2 patients is lower, and the picture seems less homogeneous, as the percentage of ACVRL1 mutations ranges from 45 to 57%. Knowledge of the relative distribution of the mutations in the two genes is helpful in developing strategies for testing HHT patients.
References
Abdalla SA, Letarte M (2006) Hereditary haemorrhagic telangiectasia: current views on genetics and mechanisms of disease. J Med Genet 43:97–110
Abdalla SA, Pece-Barbara N, Vera S, Tapia E, Paez E, Bernabeu C, Letarte M (2000) Analysis of ALK-1 and endoglin in newborns from families with hereditary hemorrhagic telangiectasia type 2. Hum Mol Genet 9:1227–1237
Abdalla SA, Cymerman U, Johnson RM, Deber CM, Letarte M (2003) Disease-associated mutations in conserved residues of ALK-1 kinase domain. Eur J Hum Genet 11:279–287
Abdalla SA, Gallione CJ, Barst RJ, Horn EM, Knowles JA, Marchuk DA, Letarte M, Morse JH (2004) Primary pulmonary hypertension in families with hereditary haemorrhagic telangiectasia. Eur Respir J 23:373–377
Abdalla SA, Cymerman U, Rushlow D, Chen N, Stoeber GP, Lemire EG, Letarte M (2005) Novel mutations and polymorphisms in genes causing hereditary hemorrhagic telangiectasia. Hum Mutat 25:320–321
Arthur HM, Ure J, Smith AJ, Renforth G, Wilson DI, Torsney E, Charlton R, Parums DV, Jowett T, Marchuk DA, Burn J, Diamond AG (2000) Endoglin, an ancillary TGFbeta receptor, is required for extraembryonic angiogenesis and plays a key role in heart development. Dev Biol 217:42–53
Bayrak-Toydemir P, McDonald J, Akarsu N, Toydemir RM, Calderon F, Tuncali T, Tang W, Miller F, Mao R (2006a) A fourth locus for hereditary hemorrhagic telangiectasia maps to chromosome 7. Am J Med Genet A 140:2155–2162
Bayrak-Toydemir P, McDonald J, Markewitz B, Lewin S, Miller F, Chou LS, Gedge F, Tang W, Coon H, Mao R (2006b) Genotype-phenotype correlation in hereditary hemorrhagic telangiectasia: Mutations and manifestations. Am J Med Genet 140A:463–470
Berg JN, Gallione CJ, Stenzel TT, Johnson DW, Allen WP, Schwartz CE, Jackson CE, Porteous ME, Marchuk DA (1997) The activin receptor-like kinase 1 gene: genomic structure and mutations in hereditary hemorrhagic telangiectasia type 2. Am J Hum Genet 61:60–67
Bossler AD, Richards J, George C, Godmilow L, Ganguly A (2006) Novel mutations in ENG and ACVRL1 identified in a series of 200 individuals undergoing clinical genetic testing for hereditary hemorrhagic telangiectasia (HHT): correlation of genotype with phenotype. Hum Mutat 27:667–675
Bourdeau A, Faughnan ME, Letarte M (2000) Endoglin-deficient mice, a unique model to study hereditary hemorrhagic telangiectasia. Trends Cardiovasc Med 10:279–285
Brusgaard K, Kjeldsen AD, Poulsen L, Moss H, Vase P, Rasmussen K, Kruse TA, Horder M (2004) Mutations in endoglin and in activin receptor-like kinase 1 among Danish patients with hereditary haemorrhagic telangiectasia. Clin Genet 66:556–561
Carmel I, Tal S, Vig I, Ast G (2004) Comparative analysis detects dependencies among the 5′ splice-site positions. RNA 10: 828–840
Cole SG, Begbie ME, Wallace GM, Shovlin CL (2005) A new locus for hereditary haemorrhagic telangiectasia (HHT3) maps to chromosome 5. J Med Genet 42:577–582
Cymerman U, Vera S, Pece-Barbara N, Bourdeau A, White RI Jr, Dunn J, Letarte M (2000) Identification of hereditary hemorrhagic telangiectasia type 1 in newborns by protein expression and mutation analysis of endoglin. Pediatr Res 47:24–35
Cymerman U, Vera S, Karabegovic A, Abdalla S, Letarte M (2003) Characterization of 17 novel endoglin mutations associated with hereditary hemorrhagic telangiectasia. Hum Mutat 21:482–492
David L, Mallet C, Mazerbourg S, Feige JJ, Bailly S (2007) Identification of BMP9 and BMP10 as functional activators of the orphan activin receptor-like kinase 1 (ALK1) in endothelial cells. Blood 109:1953–1961
den Dunnen JT, Antonarakis SE (2000) Mutation nomenclature extensions and suggestions to describe complex mutations: a discussion. Hum Mutat 15:7–12
Fernandez-L A, Sanz-Rodriguez F, Zarrabeitia R, Perez-Molino A, Hebbel RP, Nguyen J, Bernabeu C, Botella LM (2005) Blood outgrowth endothelial cells from Hereditary Haemorrhagic Telangiectasia patients reveal abnormalities compatible with vascular lesions. Cardiovasc Res 68:235–248
Fernandez-L A, Sanz-Rodriguez F, Zarrabeitia R, Perez-Molino A, Morales C, Restrepo CM, Ramirez JR, Coto E, Lenato GM, Bernabeu C, Botella LM (2006) Mutation study of Spanish patients with hereditary hemorrhagic telangiectasia and expression analysis of Endoglin and ALK1. Hum Mutat 27:295
Gallione CJ, Klaus DJ, Yeh EY, Stenzel TT, Xue Y, Anthony KB, McAllister KA, Baldwin MA, Berg JN, Lux A, Smith JD, Vary CP, Craigen WJ, Westermann CJ, Warner ML, Miller YE, Jackson CE, Guttmacher AE, Marchuk DA (1998) Mutation and expression analysis of the endoglin gene in hereditary hemorrhagic telangiectasia reveals null alleles. Hum Mutat 11:286–294
Gallione CJ, Scheessele EA, Reinhardt D, Duits AJ, Berg JN, Westermann CJ, Marchuk DA (2000) Two common endoglin mutations in families with hereditary hemorrhagic telangiectasia in the Netherlands Antilles: evidence for a founder effect. Hum Genet 107:40–44
Gallione CJ, Repetto GM, Legius E, Rustgi AK, Schelley SL, Tejpar S, Mitchell G, Drouin E, Westermann CJ, Marchuk DA (2004) A combined syndrome of juvenile polyposis and hereditary haemorrhagic telangiectasia associated with mutations in MADH4 (SMAD4). Lancet 363:852–859
Gallione CJ, Richards JA, Letteboer TG, Rushlow D, Prigoda NL, Leedom TP, Ganguly A, Castells A, Ploos van Amstel JK, Westermann CJ, Pyeritz RE, Marchuk DA (2006) SMAD4 mutations found in unselected HHT patients. J Med Genet 43:793–797
Gu Y, Jin P, Zhang L, Zhao X, Gao X, Ning Y, Meng A, Chen YG (2006) Functional analysis of mutations in the kinase domain of the TGF-beta receptor ALK1 reveals different mechanisms for induction of hereditary hemorrhagic telangiectasia. Blood 107:1951–1954
Harrison RE, Flanagan JA, Sankelo M, Abdalla SA, Rowell J, Machado RD, Elliott CG, Robbins IM, Olschewski H, McLaughlin V, Gruenig E, Kermeen F, Halme M, Raisanen-Sokolowski A, Laitinen T, Morrell NW, Trembath RC (2003) Molecular and functional analysis identifies ALK-1 as the predominant cause of pulmonary hypertension related to hereditary haemorrhagic telangiectasia. J Med Genet 40:865–871
Johnson DW, Berg JN, Baldwin MA, Gallione CJ, Marondel I, Yoon SJ, Stenzel TT, Speer M, Pericak-Vance MA, Diamond A, Guttmacher AE, Jackson CE, Attisano L, Kucherlapati R, Porteous ME, Marchuk DA (1996) Mutations in the activin receptor-like kinase 1 gene in hereditary haemorrhagic telangiectasia type 2. Nat Genet 13:189–195
Karabegovic A, Shinawi M, Cymerman U, Letarte M (2004) No live individual homozygous for a novel endoglin mutation was found in a consanguineous Arab family with hereditary haemorrhagic telangiectasia. J Med Genet 41:e119
Kjeldsen AD, Moller TR, Brusgaard K, Vase P, Andersen PE (2005) Clinical symptoms according to genotype amongst patients with hereditary haemorrhagic telangiectasia. J Intern Med 258:349–355
Klaus DJ, Gallione CJ, Anthony K, Yeh EY, Yu J, Lux A, Johnson DW, Marchuk DA (1998) Novel missense and frameshift mutations in the activin receptor-like kinase-1 gene in hereditary hemorrhagic telangiectasia. Hum Mutat 12:137, MIB n. 164
Kuehl HK, Caselitz M, Hasenkamp S, Wagner S, El-Harith el-HA, Manns MP, Stuhrmann M (2005) Hepatic manifestation is associated with ALK1 in hereditary hemorrhagic telangiectasia: identification of five novel ALK1 and one novel ENG mutations. Hum Mutat 25(3):320
Lastella P, Sabba C, Lenato GM, Resta N, Lattanzi W, Gallitelli M, Cirulli A, Guanti G (2003) Endoglin gene mutations and polymorphisms in Italian patients with hereditary haemorrhagic telangiectasia. Clin Genet 63:536–540
Lenato GM, Lastella P, Di Giacomo MC, Resta N, Suppressa P, Pasculli G, Sabba C, Guanti G (2006) DHPLC-based mutation analysis of ENG and ALK-1 genes in HHT Italian population. Hum Mutat 27:213–214
Lesca G, Plauchu H, Coulet F, Lefebvre S, Plessis G, Odent S, Riviere S, Leheup B, Goizet C, Carette MF, Cordier JF, Pinson S, Soubrier F, Calender A, Giraud S; French Rendu-Osler Network (2004) Molecular screening of ALK1/ACVRL1 and ENG genes in hereditary hemorrhagic telangiectasia in France. Hum Mutat 23:289–299
Lesca G, Burnichon N, Raux G, Tosi M, Pinson S, Marion MJ, Babin E, Gilbert-Dussardier B, Riviere S, Goizet C, Faivre L, Plauchu H, Frebourg T, Calender A, Giraud S; French Rendu-Osler Network (2006) Distribution of ENG and ACVRL1 (ALK1) mutations in French HHT patients. Hum Mutat 27:598
Lesca G, Olivieri C, Burnichon N, Pagella F, Carette MF, Gilbert-Dussardier B, Goizet C, Roume J, Rabilloud M, Saurin JC, Cottin V, Honnorat J, Coulet F, Giraud S, Calender A, Danesino C, Buscarini E, Plauchu H; French-Italian Rendu-Osler Network (2007) Genotype-phenotype correlations in hereditary hemorrhagic telangiectasia: data from the French-Italian HHT network. Genet Med 9:14–22
Letteboer TG, Zewald RA, Kamping EJ, de Haas G, Mager JJ, Snijder RJ, Lindhout D, Hennekam FA, Westermann CJ, Ploos van Amstel JK (2005) Hereditary hemorrhagic telangiectasia: ENG and ALK-1 mutations in Dutch patients. Hum Genet 116:8–16
Letteboer TG, Mager HJ, Snijder RJ, Koeleman BP, Lindhout D, Ploos van Amstel HK, Westermann KJ (2006) Genotype–phenotype relationship in Hereditary Hemorrhagic Telangiectasia. J Med Genet 43:371–377
McAllister KA, Baldwin MA, Thukkani AK, Gallione CJ, Berg JN, Porteous ME, Guttmacher AE, Marchuk DA (1995) Six novel mutations in the endoglin gene in hereditary hemorrhagic telangiectasia type 1 suggest a dominant-negative effect of receptor function. Hum Mol Genet 4:1983–1985
Olivieri C, Mira E, Delu G, Pagella F, Zambelli A, Malvezzi L, Buscarini E, Danesino C (2002) Identification of 13 new mutations in the ACVRL1 gene in a group of 52 unselected Italian patients affected by hereditary haemorrhagic telangiectasia. J Med Genet 39:E39
Olivieri C, Lanzarini L, Pagella F, Semino L, Corno S, Valacca C, Plauchu H, Lesca G, Barthelet M, Buscarini E, Danesino C (2006) Echocardiographic screening discloses increased values of pulmonary artery systolic pressure in 9 of 68 unselected patients affected with hereditary hemorrhagic telangiectasia. Genet Med 8:183–190
Paquet ME, Pece-Barbara N, Vera S, Cymerman U, Karabegovic A, Shovlin C, Letarte M (2001) Analysis of several endoglin mutants reveals no endogenous mature or secreted protein capable of interfering with normal endoglin function. Hum Mol Genet 10:1347–1357
Pece N, Vera S, Cymerman U, White RI Jr, Wrana JL, Letarte M (1997) Mutant endoglin in hereditary hemorrhagic telangiectasia type 1 is transiently expressed intracellularly and is not a dominant negative. J Clin Invest 100:2568–2579
Prigoda NL, Savas S, Abdalla SA, Piovesan B, Rushlow D, Vandezande K, Zhang E, Ozcelik H, Gallie BL, Letarte M (2006) Hereditary haemorrhagic telangiectasia: mutation detection, test sensitivity and novel mutations. J Med Genet 43:722–728
Ravnic-Glavač M, Glavač D, Dean M (1994) Sensitivity of single strand conformation polymorphism and heteroduplex method for mutation detection in the cystic fibrosis gene. Hum Mol Genet 3: 801–807
Roca X, Sachidanandam R, Krainer AR (2003) Intrinsic differences between authentic and cryptic 5′ splice sites. Nucleic Acids Res 31:6321–6333
Schulte C, Geisthoff U, Lux A, Kupka S, Zenner HP, Blin N, Pfister M (2005) High frequency of ENG and ALK1/ACVRL1 mutations in German HHT patients. Hum Mutat 25:595
Shapiro MB, Senapathy P (1987) RNA splice junctions of different classes of eukaryotes: sequence statistics and functional implications in gene expression. Nucleic Acids Res 15:7155–7174
Shovlin CL, Hughes JM, Scott J, Seidman CE, Seidman JG (1997) Characterization of endoglin and identification of novel mutations in hereditary hemorrhagic telangiectasia. Am J Hum Genet 61:68–79
Shovlin CL, Guttmacher AE, Buscarini E, Faughnan ME, Hyland RH, Westermann CJ, Kjeldsen AD, Plauchu H (2000) Diagnostic criteria for hereditary hemorrhagic telangiectasia (Rendu-Osler-Weber syndrome). Am J Med Genet 91:66–67
Trembath RC, Thomson JR, Machado RD, Morgan NV, Atkinson C, Winship I, Simonneau G, Galie N, Loyd JE, Humbert M, Nichols WC, Morrell NW, Berg J, Manes A, McGaughran J, Pauciulo M, Wheeler L (2001) Clinical and molecular genetic features of pulmonary hypertension in patients with hereditary hemorrhagic telangiectasia. N Engl J Med 345:325–334
Wehner LE, Folz BJ, Argyriou L, Twelkemeyer S, Teske U, Geisthoff UW, Werner JA, Engel W, Nayernia K (2006) Mutation analysis in hereditary haemorrhagic telangiectasia in Germany reveals 11 novel ENG and 12 novel ACVRL1/ALK1 mutations. Clin Genet 69:239–245
Yeo G, Burge CB (2004) Maximum entropy modeling of short sequence motifs with applications to RNA splicing signals. J Comput Biol 11:377–394
Acknowledgments
The authors are indebted to Dr. M. Letarte for helpful discussions on mutation analysis and on the manuscript. We also wish to thank Dr. C. Gallione and Prof. D. A. Marchuk for their contributions in MADH4 analysis, and Prof. E. Arbustini for DHPLC analysis. This work was partially supported by grant no. 80170 - IRCCS Policlinico S.Matteo of Pavia, by Fondazione Cariplo, Milano and by Fondazione Banca del Monte di Lombardia, Pavia, Italy. We wish to thank for continuous support the Fondazione Italiana “O.Carini” per la Teleangectasia Emorragica Ereditaria, all the patients and their families.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Olivieri, C., Pagella, F., Semino, L. et al. Analysis of ENG and ACVRL1 genes in 137 HHT Italian families identifies 76 different mutations (24 novel). Comparison with other European studies. J Hum Genet 52, 820–829 (2007). https://doi.org/10.1007/s10038-007-0187-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-007-0187-5
Keywords
This article is cited by
-
Stereotactic radiosurgery for brain arteriovenous malformations in patients with hereditary hemorrhagic telangiectasia
Acta Neurochirurgica (2024)
-
uAUG creating variants in the 5’UTR of ENG causing Hereditary Hemorrhagic Telangiectasia
npj Genomic Medicine (2023)
-
Impact of SARS-CoV-2 infection in patients with hereditary hemorrhagic telangiectasia: epidemiological and clinical data from the comprehensive Italian retrospective multicenter study
Internal and Emergency Medicine (2023)
-
Current HHT genetic overview in Spain and its phenotypic correlation: data from RiHHTa registry
Orphanet Journal of Rare Diseases (2020)
-
Characterization of a family mutation in the 5’ untranslated region of the endoglin gene causative of hereditary hemorrhagic telangiectasia
Journal of Human Genetics (2019)