Abstract
Down’s syndrome (DS), a chromosomal disorder due to trisomy 21, results mostly from nondisjunction in maternal meiosis. The present case-control study examined the association of genetic polymorphisms with predisposition to nondisjunction. Two common polymorphisms (SNPs), C677T and A1298C, in the 5,10-methylenetetrahydrofolate reductase (MTHFR) gene involved in folate metabolism, are known to lower the activity of this enzyme. Three hundred and fourteen mothers (with DS children and controls), mostly from the eastern states of India, were genotyped for the two above-mentioned SNPs. Significant association with both of these SNPs were detected, more specifically, in the mothers of DS children homozygous for the polymorphic alleles 677 T and 1298 C. The relative risk of T (C677T) and C (A1298C) homozygosity in mothers for DS-affected pregnancy was 7 (OR 7.67, 95% CI 1.67–35.08, P=0.003) and 4 (OR 4.40, 95% CI 1.45–13.26, P=0.008), respectively. Moreover, all 677TT mothers studied were less than 31 years of age, whereas no correlation with maternal age was observed for A1298C genotypes. Interestingly, all of the young 677TT mothers had either a first- or secondborn child with DS. Thus, this study reports that young Indian mothers with TT genotypes are genetically predisposed to nondisjunction due to abnormal folate metabolism.
Similar content being viewed by others
Introduction
The majority of DS cases occur due to trisomy of chromosome 21. Around 90% of these are caused by nondisjunction in maternal meiosis. Advanced maternal age (post 35 years) is a known risk factor for DS (Hassold and Hunt 2001). Nevertheless, a fairly high number of DS children are also born to young mothers (1/1,250 births). One of the major challenges in DS etiology is to identify possible risk factors responsible for predisposing young mothers to DS-affected pregnancy. In a pilot case-control study on mothers with DS children (case mothers) and control mothers from a population in USA, James et al. (1999) reported the association of a SNP, C677T, in the 5,10-methylenetetrahydrofolate reductase (MTHFR) gene (OMIM*607093; chromosome 1p36.3) with case mothers. A more extensive study from the same group (Hobbs et al. 2000) on different populations in the USA further substantiated their earlier observations. They also showed that a SNP in another gene associated with methionine–homocysteine metabolism, methionine synthase reductase (MTRR, OMIM*602568; chromosome 5p15.2-.3), also has an association with case mothers (with DS children). However, subsequent studies from different populations from Europe, the USA and Turkey have produced a mixed picture, finding no association with MTHFR C677T and showing either no or weak association with MTRR and another MTHFR SNP, A1298C (Chadefaux-Vekemans et al. 2002; O’Leary et al. 2002; Boduroglu et al. 2004).
Both MTHFR and MTRR code for enzymes involved in homocysteine–methionine metabolism. Methionine is converted to S-adenosylmethionine, the lone donor of a methyl group for epigenetic modification of DNA (cytosine methylation) and post-translational methylation of histones (Fuks et al. 2003). The C to T transition (C677T) in exon 4 of the MTHFR gene results in the substitution of alanine with valine (A222V) in the N-terminal catalytic domain that renders the enzyme thermolabile and less active (Frosst et al. 1995). Lowering of the enzyme activity would purportedly lead to hypomethylation of DNA and histones that might globally affect gene expression. In exon 7 of the same gene, an A to C transversion at A1298C leads to a glutamate to alanine substitution (E429A) in the C-terminal regulatory domain of the enzyme, without overtly affecting enzyme activity (Van der Put and Blom 2000).
In the present study, case mothers (with DS children) and control mothers (with no DS children) have been genotyped for MTHFR SNPs (both C677T and A1298C) in an Indian population mostly from eastern India. Significant associations with both SNPs were detected in homozygous 677TT and 1298CC case mothers. The transmission pattern of the parental 677T allele to the DS child was also studied.
Materials and methods
A total of 149 case mothers (mean maternal age 27.4±6.7, range 18–47 years; 101 mothers aged <31 years and 48 mothers >31 years) with a DS child and 165 control mothers (mean age 26.8±8.0 years, range 19–53 years) with normal children that have no reported abnormalities were recruited for the study. The vast majority of samples were taken from the Paediatrics and Gynaecology divisions of the University Hospital in Varanasi (situated in the eastern part of India). In order to check the possibility of the results being unique to the population in this part of the country, a small group of samples (26 case and 20 control) was obtained from Ahmedabad (western India ~1,300 km west of Varanasi). The Ahmedabad samples were used only for C677T analysis, and since the genotype and allele frequencies obtained were comparable to those from Varanasi they were pooled for further analyses. Pedigree was recorded for each family. Blood was drawn in heparinized syringes from DS child, case and control mothers by venipuncture. DNA was isolated from the blood. Using PCR-RFLP (Hobbs et al. 2000; Van der Put et al. 1998), individuals were genotyped for C677T SNP. For A1298C polymorphism, only 89 case and 70 control mothers were genotyped. Karyotyping of direct blood culture was performed on the children to confirm trisomy 21.
Statistical comparisons were made using the SPSS (version 0.6, Chicago, IL, USA) software package. Allele frequency was calculated for each genotype and the difference in allele frequency between controls and cases was determined using the χ2 test for independence with Pearson statistics. Expected genotype frequencies were calculated from the allele frequencies under the assumption of Hardy–Weinberg equilibrium. Odds ratios (OR) and 95% confidence intervals (CI) were calculated using the calculator developed by Dr. Hutchon, Consultant Gynecologist, Memorial Hospital, Darlington, England. Odds ratio was used as an estimate of relative risk. Statistical significance was interpreted as two tailed P values of <0.05 (using GraphPad software).
Results
C677T
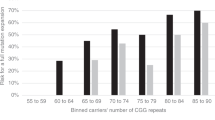
Table 1 shows that the frequencies of the wild-type C allele and CC genotype are high in both the case and control mothers. However, the incidences of the T allele and CT/TT genotypes were higher in case mothers and the differences from the control mothers were statistically significant. The frequency of the homozygous TT genotype in the case mothers was 7.6 times higher than in the controls. The distribution of C677T genotypes according to age groups revealed that all 12 homozygous TT mothers were aged less than 31 years (total mothers <31 year = 101) (Fig. 1). The genotypic distribution in case mothers who were more than 31 years of age was comparable to the controls. The pedigrees of 149 families could be divided into type-I (T-I) and type-II (T-II) depending upon the number of siblings and the position of the DS child in the pedigree. Those with T-I pedigree had up to four children, with either first or the second child affected with DS, while those with T-II had four to eight children, and the DS child was usually the last born in the family. Most mothers of DS children older than 35 years of age belonged to the T-II pedigree, a classical old age case of DS (48 out of 149 families; 32%) (Fig. 2).
In order to understand the pattern of transmission of the 677 C allele and the 677 T allele from parents to DS children, 84 family triads comprising mother, father and DS child were genotyped. Amongst these, five triads were uninformative because the parents and the DS offspring were heterozygous (CT). The frequencies of the T allele and the homozygous TT genotype in DS children and their mothers were comparable. In fathers, the frequency of T was considerably lower than that of the mothers and DS children (Table 2). Both 677 C and T alleles were transmitted equally from the mothers to DS children, as per the Mendelian ratio. Out of 20 informative transmissions from the mother to DS child, 11 involved the C allele and 9 involved the T allele, whereas the T allele was preferentially transmitted to DS children from the father (of 13 informative transmissions from father to DS child, the T allele was involved 12 times and the C allele just once); see Table 3.
A1298C
The polymorphism of A1298C, studied in 89 case and 70 control mothers, revealed that the frequencies of the C allele and the homozygous CC genotype were significantly higher in the case mothers (Table 4). The odds ratio for CC in case mothers was 4.4 (95%CI: 1.45–13.26) versus the controls. Genotypes were distributed uniformly in the case mothers across different age groups (data not shown).
Since the homozygosity for each of the polymorphic alleles 677T and 1298C carried high odds, the possible effects of the combined genotypes were assessed in 89 case and 70 control mothers (Table 5). Homozygous 677CC/1298CC genotypes were observed in the case and control mothers with an odds ratio of >4, whereas the rare 677CT/1298CC genotype occurred only in the case mothers, suggesting that the 677T allele resulted in added severity to the homozygous 1298CC genotype. The 677TT/1298AA, 677TT/1298AC and 677TT/1298CC combined genotypes were present only in case mothers, not in controls, indicating that the homozygous 677TT is an independent risk irrespective of the A1298C genotypes.
Discussion
5,10-Methylenetetrahydrofolate reductase, a key enzyme in methionine–homocysteine metabolism, maintains the folate pool between the DNA synthesis and methylation pathways. Expression of a large number of genes (especially tissue-specific) is also regulated by DNA and/or histone methylation, though nearly 10% of the mammalian genome carries imprinted genes. Hence, hypomethylation of DNA/histones could affect various cellular functions, such as DNA repair, replication, gene expression and chromatin conformation, leading to disease conditions.
While C677T SNP, particularly in the homozygous state, is a risk factor for multifactorial disorders (such as cardiovascular, spina bifida (NTD) and also infertility, Singh et al. 2005), the other SNP, A1298C, does not seem to affect the enzyme, though its risk association with NTD has been recorded (Van der Put et al. 1998).
James et al. (1999) were the first to demonstrate that in certain American populations young mothers with 677TT genotypes run the risk of delivering DS children. The present study demonstrates the association of both C677T and A1298C SNPs with mothers of DS children and a correlation with the mother’s age in an Indian population. However, various other studies did not find a similar association with DS mothers (Petersen et al. 2000; O’Leary et al. 2002; Chadefaux-Vekemans et al. 2002; Boduroglu et al. 2004; Chango et al. 2005; Da Silva et al. 2005). Da Silva et al. (2005) have shown a higher frequency of the 677T allele and elevated homocysteine levels in mothers with DS children, but consider it a minor predisposing risk factor for DS. O’Leary et al. 2002, did not find an association of MTHFR with DS, but observed that an SNP in MTRR, another gene in the methionine–homocysteine metabolic pathway, is associated with DS.
Chadefaux-Vekemans et al. (2002) did stratify case mothers according to their ages (average age 35 years) into two categories: less than and more than 40 years, and found no difference in the allele and genotype frequencies between the two categories for C677T. In contrast to these reports, the present study revealed that all TT homozygous mothers were less than 31 years of age, although no such age relation was observed with A1298C genotypes, possibly due to its weak association with mothers of DS children. One of the reasons for the disparity in the results obtained from young and old mothers could be due to region-specific differences in the frequency of the T allele. Since the frequency of DS-affected pregnancy increases sharply beyond 35 years of maternal age, a greater proportion of case mothers examined by other researchers happened to be of advanced age (>35 years). Therefore, it is also possible that either lack of age classification or fewer number of young mothers in their studies could have diluted the age-based genotypic distinction among mothers.
Pedigree analysis of all the 149 families revealed that the DS children of all TT homozygous mothers were either the first or the second in the brood, suggesting the possibility of them being genetically predisposed to abnormal disjunction.
Dietary folate deficiency of an individual along with MTHFR polymorphisms leads to DNA hypomethylation (Ulrey et al. 2005; Castro et al. 2004; Friso et al. 2002; Roizen and Patterson 2003). It is also known that hypomethylation of pericentromeric heterochromatic DNA may lead to nondisjunction during cell division (Hassold and Hunt 2001; James et al. 1999). Additionally, in populations where no correlation has been observed between the SNPs and mothers with DS children, the frequency of the T allele in the normal population is much higher (20–40%) than in the population presently examined by us or in other Asian and African populations (~10% with very low T homozygosity, Schneider et al. 1998; Botto and Yang 2000; Ogino and Wilson 2003). In general, there is also good evidence of nutritional folate fortification in Western populations. Girelli et al. (1998) have clearly demonstrated that 677TT subjects with low folates have very high homocysteine (Hcy) levels, whereas those with adequate folates have normal Hcy levels, regardless of the C677T genotype. However, the fact that nutritional fortification of folates substantially obviates the adverse effect of the SNP explains why a large section of the population with CT or TT genotypes of C677T is completely unaffected. Moreover, in certain American and European “normal” populations, the frequency of the T allele is as high as 40% (Schneider et al. 1998).
In India, a few studies on the folate and Hcy levels in general populations have revealed that the basal level of homocysteine is higher whereas that of folate is lower than the recommended WHO standards (Lakshmi et al. 2001; Refsum et al. 2001; Kumar et al. 2005). In a large-scale study on MTHFR SNPs from southern India, Radha Rama Devi et al. (2004) showed that the frequency of the T allele is lower than those seen in European and American populations. Thus the present study suggests that in certain populations with low folate and high homocysteine, the C allele of C677T has a distinct selective advantage against T. It further indicates that, while the effects of T allele polymorphisms may be alleviated in Western populations due to nutritional folate fortification, it will be a risk factor in any population with a dietary deficiency of folates. This nutritional constraint may also lead one to assume that a weak SNP, A1298C, could also be a risk factor for DS mothers in this population.
The present paper also reports the transmission pattern of C677T to the DS child. Inheritance of T from the CT-heterozygous parents showed that both C and T maternal alleles followed Mendelian segregation, whereas there was preferential transmission of the T allele from father to DS child. Hobbs et al. (2002), using a larger sample, also observed that the paternal T allele was preferentially transmitted to the DS child. The significance of this segregation distortion in the paternal meiosis is not immediately apparent. Hobbs et al. (2002) have speculated that the presence of the 677T allele might confer a survival advantage to a DS child, but why the transmission occurs preferentially from the father is not clear.
Thus the present study re-emphasizes that folate metabolism can be modulated by genetic factors (like SNPs in MTHFR, MTRR genes) and dietary folate supplements.
References
Boduroglu K, Alanay Y, Koldan B, Tuncbilek E (2004) Methylenetetrahydrofolate reductase enzyme polymorphism as maternal risk for Down syndrome among Turkish women. Am J Med Genet 127A:5–10
Botto LD, Yang Q (2000) 5,10-Methylenetetrahydrofolate reductase gene variants and congenital anomalies: a HuGE review. Am J Epidemiol 151:862–877
Castro R, Rivera I, Ravasco P, Camilo ME, Jakobs C, Blom HJ, de Almeida IT (2004) 5,10-Methylenetetrahydrofolate reductase (MTHFR) 677C->T and 1298A->C mutations are associated with DNA hypomethylation. J Med Genet 41:454–458
Chadefaux-Vekemans B, Coude M, Muller F, Oury JF, Chabli A, Jais J, Kamoun P (2002) Methylenetetrahydrofolate reductase polymorphism in the etiology of Down syndrome. Pediatr Res 51:766–767
Chango A, Fillon-Emery N, Mircher C, Blehaut H, Lambert D, Herbeth B, James SJ, Rethore MO, Nicolas JP (2005) No association between common polymorphisms in genes of folate and homocysteine metabolism and the risk of Down’s syndrome among French mothers. Br J Nutr 94:166–169
Da Silva LR, Vergani N, Galdieri Lde C, Ribeiro Porto MP, Longhitano SB, Brunoni D, D’Almeida V, Alvarez Perez AB (2005) Relationship between polymorphisms in genes involved in homocysteine metabolism and maternal risk for Down syndrome in Brazil. Am J Med Genet 135:263–267
Friso S, Choi SW, Girelli D, Marson JB, Dolnikowski GG, Bagley PJ, Olivieri O, Jacques PF, Rosenberg IH, Corrocher R, Selhub J (2002) A common mutation in the 5,10-methylenetetrahydrofolate reductase gene affects genomic DNA methylation through an interaction with folate status. Proc Natl Acad Sci USA 99:5606–5611
Frosst P, Blom HJ, Kluijtmans LAJ, van den Heuvel LP, Rozen R (1995) A candidate genetic risk factor for common mutation in methylenetetrahydrofalate reductase. Nat Genet 10:111–113
Fuks F, Hurd PJ, Wolf D, Nan X, Bird AP, Kouzarides T (2003) The methyl-CpG-binding protein MeCP2 links DNA methylation to histone methylation. J Biol Chem 278:4035–4040
Girelli D, Frso S, Trabetti E, Olivieri O, Russo C, Pessotto R, Faccini G, Pignatti PF, Mazzucco A, Corrocher R (1998) Methylenetetrahydrofolate reductase C677T mutation, plasma homocysteine, and folate in subjects from northern Italy with or without angiographically documented severe coronary atherosclerotic disease: evidence for an important genetic-environmental interaction. Blood 91:4158–4163
Hassold T, Hunt P (2001) To err (meiotically) is human: the genesis of human aneuploidy. Nat Rev Genet 2:280–291
Hobbs CA, Sherman SL, Yi P, Hopkins SE, Torfs CP, Hine RJ, Pogribna M, Rozen R, James SJ (2000) Polymorphisms in genes involved in folate metabolism as maternal risk factors for Down syndrome. Am J Hum Genet 67:623–630
Hobbs CA, Cleves MA, Lauer RM, Burns TL, James SJ (2002) Preferential transmission of the MTHFR 677 T allele to infants with Down syndrome: implication for survival advantage. Am J Med Genet 113:9–14
James SJ, Pogribna James SJ, Pogribna M, Pogribny IP, Melnyk S, Hine RJ, Gibson JB, Yi P, Tafoya DL, Swenson DH, Wilson VL, Gaylor DW (1999) Abnormal folate metabolism and mutation in the methylenetetrahydrofolate reductase gene may be maternal risk factors for Down syndrome. Am J Clin Nutr 70:495–501
Kumar J, Das SK, Sharma P, Karthikeyan G, Ramakrishnan L, Sengupta S (2005) Homocysteine levels are associated with MTHFR A1298C polymorphism in Indian population. J Hum Genet 50:655–663
Lakshmi AV, Maniprebha C, Krishana TP (2001) Plasma homocysteine level in relation to folate and vitamin B6 status in apparently normal men. Asia Pac J Clin Nutr 10:194–96
O’Leary VB, Parle-McDermott A, Molly AM, Kirke PN, Johnson Z, Conley M, Scott JM, Mills JL (2002) MTRR and MTHFR polymorphism: link to Down syndrome? Am J Med Genet 107:151–155
Ogino S, Wilson RB (2003) Genotype and haplotype distribution of MTHFR677C>T and 1298A>C single nucleotide polymorphism: a meta-analysis. J Hum Genet 48:1–7
Petersen M, Grigoriadou M, Mikkelsen M (2000) A common mutation in the methylenetetrahydrofolate reductase gene is not a risk factor for Down syndrome. Am J Hum Genet 67(Suppl 2):141
Radha Rama Devi A, Govindaiah V, Ramakrishna G, Naushad SM (2004) Prevalence of methylene tetrahydrofolate reductase polymorphism in South Indian population. Curr Sci 86:440–443
Refsum H, Yajnik CS, Gadkari M, Schneede J, Vollset SE, Orning L, Guttormsen AB, Joglekar A, Sayyad MG, Ulvik A, Ueland PM (2001) Hyperhomocysteinemia and elevated methylmalonic acid indicate a high prevalence of cobalamin deficiency in Asian Indians. Am J Clin Nutr 74:233–241
Roizen NJ, Patterson D (2003) Down’s syndrome. Lancet 361:1281–1289
Schneider JA, Rees DC, Liu YT, Clegg JB (1998) World distribution of a common methylenetetrahydrofalate reductase mutation. Am J Hum Genet 62:1258–1260
Singh K, Singh SK, Sah R, Singh I, Raman R (2005) Mutation C677T in the methylenetetrahydrofolate reductase gene is associated with male infertility in an Indian population. Int J Androl 28:115–119
Ulrey CL, Liu L, Andrews LG, Tollefsbol TO (2005) The impact of metabolism on DNA methylation. Hum Mol Genet 14:R139–R147
Van der Put NM, Blom HJ (2000) Reply to Donnelly. Am J Hum Genet 66:744–775
Van der Put NM, Gabreels F, Stevens EM, Smeitinik JA, Trijbels FJ, Eskes TK, van den Heuvel LP, Blom HJ (1998) A second common mutation in the methylenetetrahydrofalate reductase gene: an additional risk factor for neural-tube defects? Am J Hum Genet 62:1044–1051
Acknowledgements
We thank Prof. Mercy J. Raman of the Department of Zoology for helpful discussions, Prof. Sulekha Pandey (Head) and Dr. Sukhbir Kaur Sidhu of the Department of Obstetrics and Gynecology, Institute of Medical Sciences for their help when procuring blood from the controls. Competing interest: None declared. Ethical approval: Written consents were obtained from all of the participants; approved by the “Research Ethical Committee” of the Institute of Medical Sciences, Banaras Hindu University, India. Contributors: RR and AKR designed the study, AK referred the DS cases, LKP provided control mothers from the University hospital, AKR, SS and SM carried out the DNA analysis, and AKR, SS did the statistical analysis. All authors have approved the final manuscript. RR is the guarantor. Funding: Department of Biotechnology, New Delhi, India to RR.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rai, A.K., Singh, S., Mehta, S. et al. MTHFR C677T and A1298C polymorphisms are risk factors for Down’s syndrome in Indian mothers. J Hum Genet 51, 278–283 (2006). https://doi.org/10.1007/s10038-005-0356-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-005-0356-3
Keywords
This article is cited by
-
RETRACTED ARTICLE: Analyzing gene polymorphism and metal folic acid interactions in neural tube defects using optimized deep recurrent neural networks
Personal and Ubiquitous Computing (2023)
-
Choline metabolic pathway gene polymorphisms and risk for Down syndrome: An association study in a population with folate-homocysteine metabolic impairment
European Journal of Clinical Nutrition (2017)
-
Whole genome sequencing identifies ANXA3 and MTHFR mutations in a large family with an unknown equinus deformity associated genetic disorder
Molecular Biology Reports (2016)
-
Synthetic combinations of missense polymorphic genetic changes underlying Down syndrome susceptibility
Cellular and Molecular Life Sciences (2016)
-
Maternal MTHFR polymorphism (677 C–T) and risk of Down’s syndrome child: meta-analysis
Journal of Genetics (2016)