Abstract
In single nucleotide polymorphism (SNP) data analysis, the allelic odds ratio and its confidence interval (CI) are usually used to evaluate the association between disease and alleles at each SNP. The usual formula for calculating the CI of the allelic odds ratio based on the Hardy–Weinberg equilibrium (HWE) may, however, lead to errors beyond the control assured by the nominal confidence level if HWE is not true. We therefore present a generalized formula for CI that does not assume HWE. CIs calculated by this generalized formula are likely to be wider than those by the usual method if the Hardy–Weinberg disequilibrium (HWD) is toward a relative deficiency of the heterozygotes (fixation index greater than 0), whereas they are likely to be narrower if HWD is toward a relative excess of the heterozygotes (fixation index less than 0). A simulation experiment to examine the influence of the generalization was performed for the case where 2% of SNPs had a fixation index greater than 0. The result revealed that the generalized method slightly decreased the mean number of falsely detected SNPs.
Similar content being viewed by others
Introduction
Recently, we have witnessed the completion of the project for identifying the human genome sequence (International Human Genome Sequencing Consortium 2004), the accumulation of enormous SNP-related data into public databases (Sachidanadam et al. 2001; Haga et al. 2002), and the development of high throughput SNP typing technologies. This progress has provided modern molecular biology with an ability to identify a genotype (combination of alleles) at any particular genetic locus for a large number of individuals (Hirschhorn et al. 2005).
In genetic association studies, the phenotype of interest is typically associated with an allele or genotype for biallelic markers, such as SNPs, and consequently many researchers are interested in calculating the allelic odds ratio and its confidence interval (CI) for identifying SNPs that may have a close association, e.g., to a certain disease. The usual method, which calculates the CI using Eq. 1 based on the logarithm \(\left(\log \hat{\psi} \right)\) of the estimated allelic odds ratio, the upper α/2 quantile (z α/2) of the standard normal distribution, and observed frequencies n ij’s in Table 1 (Balding et al. 2001), assumes the Hardy–Weinberg equilibrium (HWE) in study populations.
Hardy–Weinberg disequilibrium (HWD) is often encountered when experimental errors occur in the SNP typing. However, even after the careful quality control of the genotyping, the genotype distribution may depart from HWE for a variety of other reasons, such as stratification, selection, inbreeding, assortative or disassortative mating (Wright 1951, 1965; Nei 1987). Under such a Hardy–Weinberg disequilibrium (HWD), the standard error of the estimated allelic odds ratio given in the last term of Eq. (1) will either be overestimated or underestimated. In order to solve this problem, Schaid and Jacobsen (1999) provided a correction method based on determining the correct variance for the observed allele frequency difference \(\left(\hat{P}_{11} - \hat{P}_{12} \right)\) between cases and controls, and quantified the effect on the type I error rate of Pearson’s chi-square test induced by HWD. Additionally, the standard error of relative risk under HWD was shown by Zaykin et al. (2004). In this article, we present a generalized formula for calculating the CI of the allelic odds ratio based on the estimated standard error, which is valid under both HWE and HWD, and then examine the effect of this generalization in a genome-wide association study.
Materials and methods
Derivation of the generalized method of CI calculation
In case-control studies, allelic frequencies are compared between cases and controls. Assuming that two alleles X and x exist at a certain SNP locus, the genotype data are given in a 3 × 2 contingency table as shown in Table 1, the observed frequencies (n 1j, n 2j, n 3j) being distributed as a trinomial distribution Tn (n .j; π1j, π2j, π3j) for j=1 (case) and j=2 (control), where (π1j, π2j, π3j) are the population proportions of genotype (XX, Xx, xx), respectively, and n .j (j=1, 2) is the sample size for each population. Of course, π1j+π2j+π3j=1 and n 1j+n 2j+n 3j=n .j (j=1, 2).
Let the population proportions of allele X in cases and controls be P 11 and P 12. Then P 11=π11+π21/2 and P 12=π12+π22/2, and they are estimated as \(\hat{P}_{1j} = {(2n_{{1j}} + n_{{2j}})} \mathord{\left/ {\vphantom {{(2n_{{1j}} + n_{{2j}})} {(2n_{{.j}})}}} \right. \kern-\nulldelimiterspace} {(2n_{{.j}})} (j = 1, 2)\) (Li and Horvitz 1953; Sasieni 1997) in Table 2. The estimator of allelic odds ratio \(\psi = \frac{{P_{11} {\left({1 - P_{12} } \right)}}}{{(1 - P_{11})P_{12} }}\) is given by Eq. 2. (See Appendix.)
When n .1 and n .2 are large, \(\log \hat{\psi}\) is asymptotically distributed as normal with mean and variance given by Eqs. 3 and 4, respectively. (See Appendix.)
where F 1 and F 2 are fixation indices of case and control populations, respectively.
Based on the estimated standard error \({\rm SE}\left(\log \hat{\psi} \right)\) that is given by Eqs. 5 and 6, an approximate 100(1 − α)% CI for ψ is given by Eq. 7. (See Appendix.)
When HWE is true without doubt, Eq. 5 should be changed to \(\hat{F}_{1} = \hat{F}_{2} = 0\) and then Eq. 7 reduces to Eq. 1, which implies that calculating CI by Eq. 7 is a generalization of the usual method. The essential derivation idea of the generalized method is to introduce the fixation index (F j) into the population probabilities of genotypes (π1j, π2j and π3j). In actuality, as F j approaches 0, one automatically arrives at the usual Eq. 1.
Numerical evaluation of the difference of the two formulas
It is obvious from Eq. 5 that the calculated CI is wider in the generalized method than the one in the usual method if \(\hat{F}_{1} > 0\;\hbox{and}\;\hat{F}_{2} > 0,\) while it is narrower if they are less than 0. However, the difference of the two methods should be evaluated numerically, because it is influenced by sampling errors of F 1 and F 2. We evaluated the difference by a numerical calculation of expected upper and lower confidence limits for various values of the fixation indices and sample sizes in the case of P 11=0.10 and P 12=0.15. In the calculation, we used a normal approximation to the trinomial distribution and the software SAS for computing.
Simulation experiment to examine the influence of generalization
In SNP data analysis, we simultaneously investigate the association between thousands of SNPs and a disease. Some SNPs among them may be under HWD with a distribution of fixation index, while others may be under HWE (F=0). We have to examine the performance of the generalized method for CI calculation, assuming that the fixation indices have a distribution among thousands of SNPs. Consequently, we conducted a Monte Carlo simulation experiment to statistically identify disease-associated SNPs using the decision rule that an association was judged as positive if the calculated CI did not include 1.0.
As the framework of simulation, we set the following conditions referring to the genome-wide association study (Sato et al. 2004):
Condition 1 The total number of SNPs to be examined was set as N=10,000 and the number of disease-associated SNPs (positive SNPs) was set as N .p=50, referring to the literature (Sing et al. 1996; Wright et al. 1999; Pharoah et al. 2002; Ponder 2001).
Condition 2 Allelic odds ratio for positive N .p SNPs was ψ=1.5 or 2.0, but ψ=1.0 for the remaining N−N .p SNPs.
Condition 3 The sample size was varied as n=n .1=n .2=188, 376 or 752.
Condition 4 The proportion P 12 of allele X in the control population was a random variable uniformly distributed in unit interval (0.05, 0.95), and P 11 in the case population was automatically determined by P 12 through Eq. 20 in Appendix. This condition was set with reference to Fig. 1, to which a uniform distribution is plausible, for the distribution of alleles in the database of Japanese Single Nucleotide Polymorphisms (Haga et al. 2002; Hirakawa et al. 2002). In our genome-scan, we did not include these SNPs with low allele frequency (P 11>0.95 or P 11<0.05). Note that (π1j, π2j, π3j, j=1, 2) were fixed through Eq. 12 in Appendix when (P 11, P 12, F) or, equivalently, (P 11, ψ, F) was determined.
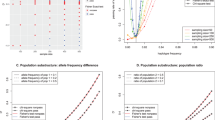
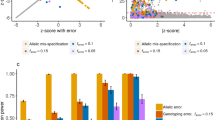
Condition 5 In a case-group, the fixation index F was specified by a mixed distribution of a constant 0 with probability 1−w and a normal distribution N(μ, 0.102) with probability w, where w=0.02, 0.06 or 0.10, and μ was set as 0.0 (in the null case), 0.2, or 0.4. On the other hand, F was set to 0 for a control group. Note that this condition was set referring to Figs. 2 and 3 taken from a database, Genome Medicine Database of Japan. In order to determine whether normally distributed or not, we showed a quantile–quantile plot in Fig. 2. It showed that the core data reasonably fit a normal distribution, but the tail data do not. Therefore, the distribution of observed F does not have a normal distribution with mean 0. Moreover, around 2% of the larger tail area in Fig. 3 was laid outside the distribution of observed F under the null hypothesis that the fixation index was equal to 0 and the mean of the outlying values was around 0.2 or more.
Condition 6 The criteria to evaluate the performance of the decision rule were two indicators, positive predictive value R p and sensitivity R s, defined by Eqs. 8 and 9 with notations in Table 3.
Condition 7 The Monte-Carlo simulation to observe R p and R s was repeated 1,000 times, and the mean values, together with the mean number of N TP and N FP, were used for comparison of the two methods.
An example of the minor allele frequency distribution of SNP. The data are from the JSNP database (http://www.snp.ims.u-tokyo.ac.jp/)
Quantile–quantile plot for fixation index F in a case-group obtained from Genome Medicine Database of Japan, http://www.gemdbj.nibio.go.jp/dgdb/)
Note that N .p was a constant fixed by Condition 1, whereas N p. was a random variable realized as the sum of N TP and N FP in the simulation experiment. Note further that these N TP and N FP have a trade-off relationship depending on the nominal confidence level, but that we fix the nominal confidence level as 1 − α=0.999, taking the multiplicity of SNPs into consideration.
The procedure to conduct the simulation experiment was as follows:
-
Step 1.
Assign a set of values to N, N .p, ψ, and n according to the above-described conditions.
-
Step 2.
Assign the value ψ=1.5 or 2.0 to the first N .p SNPs and ψ=1.0 to the remaining N−N .p SNPs.
-
Step 3.
Generate 10,000 random numbers of F according to Condition 5 and assign them to 10,000 SNPs.
-
Step 4.
Generate random numbers (n 11, n 21, n 31) and (n 12, n 22, n 32) distributed as Tn(n, π11, π21, π31) and Tn(n, π12, π22, π32), respectively, for each 10,000 SNPs.
-
Step 5.
Calculate CIs using Eq. 1 (usual method) and Eq. 8 (Generalized method) with α=0.001 and calculate N TP, N FP, R p, and R s for each 10,000 SNPs.
-
Step 6.
Repeat Steps 1–5 1,000 times and calculate the mean of the realized values.
-
Step 7.
Repeat Steps 1–6, changing parameters ψ in Condition 2, n in Condition 3, and w and μ in Condition 4.
Results
A summarized result of numerical evaluation of the expected confidence limits in a typical case is shown in Table 4 for various values of the fixation index F=F 1=F 2 when the sample size was set at n .1=n .2=188 or 752. Table 4 suggests that the difference of the two methods is not ignorable, on average, judging by statistical significance when F≥0.4, because the CI by the generalized method included 1.0, whereas CI by the usual method did not.
The essential feature of the influence of the generalized method on the judgment of association can be seen in Table 5, which is the mean of R p, R s, N TP, and N FP obtained from the 1,000 simulation repetitions. When ψ=1.5 or 2.0, w=0.02, 0.06 or 0.10 and μ=0.2 or 0.4, the false positive number of SNPs in the generalized method was, on average, slightly less than that in the usual method.
Discussion
The essential improvement achieved by the generalized method is summarized in Table 5. In this table, for example, the average number of falsely detected SNPs by the usual method was 22.0 (n=188) or 22.0 (n=752), whereas it was 20.4 (n=188) or 19.7 (n=752) by the generalized method when ψ=2.0, w=0.10 and μ=0.4. The amount of the improvement was not great, but it may be appreciated in certain research circumstances, because a difference of even a few SNPs would be highly significant in the advanced stages of gene hunting following an association study, such as large-scale, multiethnic replication studies or lengthy functional analyses on model animals. It should be noted that a substantial investment in the post-association study is often necessary, especially in a hypothesis-free genome scan, in which a prior probability of the gene is minimal.
Deviation from HWE is not a rare, exceptional case in association studies. Figure 2 shows an example of the distribution of the fixation index in a large-scale SNP typing project, in which 84,542 SNP typing data on autosomal chromosomes were obtained for 940 individuals in the Millennium Genome Project of Japan (Haga et al. 2002; Yoshida and Yoshimura 2003). In this dataset, the operating protocol of our SNP typing laboratory includes routine quality check steps to filter simple experimental errors. However, even after the careful check for the genotyping errors, a sizable fraction of about 2% of the 84,542 SNPs showed a fixation index outside the normal range of variation under the hypothesis that the population fixation index was 0.
As for other data, Wittke-Thompson et al. (2005) did a survey of HWD in several recent reviews of association studies (Xu et al. 2002; Gyorffy et al. 2004; Kocsis et al. 2004a, b; Osawa et al. 2004) and identified 41 studies with 60 polymorphisms showing a departure from HWE: 35 polymorphisms that depart from HWE in cases only, 21 that departed in controls only, 2 that departed in the same direction in cases and controls, and 1 that departed in the opposite direction in cases and controls. Wittke-Thompson et al. (2005) emphasized the importance not only of correctly assessing HWE for genotype data but also of understanding whether an observed HWD was consistent with a genetic model of disease susceptibility.
In a previous study, Schaid and Jacobsen (1999), Zaykin et al. (2004) and Salanti et al. (2005) each recommended the correction of the variance of the observed statistics which is allele frequency difference, relative risk or odds ratio under HWD, respectively, because the type I error for gene-disease associations tested on the level of alleles was inflated when the estimated inbreeding coefficient was positive, while the error deflates for the negative coefficient. However, under circumstances where the assumptions of HWE in controls and codominance between the alleles do not hold well, Sasieni (1997) recommended simply to abandon the allelic odds ratio for an association study, because the allelic odds ratio and chi-square statistics are not robust under such circumstances. These previous studies were targeted at a candidate–gene association study or meta-analysis and did not examine a genome-wide association study. Here, we scrutinized the situation in a genome-wide association study and showed that around 2% of the large tail area was laid outside the distribution of F, suggesting the importance of the correction under HWD. Because the cardinal feature of the genome-wide association study is a screening, we believe that Sasieni’s recommendation may be too conservative to be accepted, and the generalized method should be applied as a sensitivity analysis in a genome-wide association study to improve both false positive rate (for F>0) and false negative rate (for F<0).
References
Agresti A (2001) Categorical data analysis, 2nd edn. Wiley, New York
Balding D, Bishop M, Cannings C (2001) Handbook of statistical genetics. Wiley, New York
Bishop Y, Fienberg S, Holland P (1975) Discrete multivariate analysis: theory and practice. MIT Press, Cambridge
Gyorffy B, Kocsis I, Vasahelyi B (2004) Biallelic genotype distributions in papers published in Gut between 1998 and 2003: altered conclusions after recalculating the Hardy–Weinberg equilibrium. Gut 53:614–616
Haga H, Yamada R, Ohnishi Y, Nakamura Y, Tanaka T (2002) Gene-based SNP discovery as part of the Japanese Millennium Genome Project: identification of 190,562 genetic variations in the human genome. Single-nucleotide polymorphism. J Hum Genet 47:605–610
Hirakawa M, Tanaka T, Hashimoto Y, Kuroda M, Takagi T, Nakamura Y (2002) JSNP: a database of common gene variations in the Japanese population. Nucleic Acids Res 30:158–162
Hirschhorn JN, Daly MJ (2005) Genome-wide association studies for common diseases and complex traits. Nat Rev Genet 6:95–108
International Human Genome Sequencing Consortium (2004) Finishing the euchromatic sequence of the human genome. Nature 431:931–945
Kocsis I, Gyorffy B, Nemeth E, Vasarhelyi B (2004a) Examination of Hardy–Weinberg equilibrium in papers of Kidney International: an underused tool. Kidney Int 65:1956–1958
Kocsis I, Vasarhelyi B, Gyorffy A, Gyorffy B (2004b) Reanalysis of genotype distributions published in Neurology between 1999 and 2002. Neurology 63:357–358
Li CC, Horvitz DG (1953) Some methods of estimating the inbreeding coefficient. Am J Hum Genet 5:107–117
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Osawa H, Yamada K, Onuma H, Murakami A, Ochi M, Kawata H, Nishimiya T, Niiya T, Shimizu I, Nishida W, Hashiramoto M, Kanatsuka A, Fujii Y, Ohashi J, Makino H (2004) The G/G genotype of a resistin single-nucleotide polymorphism at −420 increases type 2 diabetes mellitus susceptibility by inducing promoter activity through specific binding of Sp1/3. Am J Hum Genet 75:678–686
Pharoah PD, Antoniou A, Bobrow M, Zimmern RL, Easton DF, Ponder BA (2002) Polygenic susceptibility to breast cancer and implications for prevention. Nat Genet 31:33–36
Ponder BA (2001) Cancer genetics. Nature 17:336–341
Sachidanandam R, Weissman D, Schmidt SC, Kakol JM, Stein LD, Marth G, Sherry S et al (2001) A map of human genome sequence variation containing 1.42 million single nucleotide polymorphisms. Nature 409:928–933
Salanti G, Amountza G, Ntzani EE, Ioannidis JP (2005) Hardy–Weinberg equilibrium in genetic association studies: an empirical evaluation of reporting, deviations, and power. Eur J Hum Genet 13:840–848
Sasieni PD (1997) From genotypes to genes: doubling the sample size. Biometrics 53:1253–1261
Sato Y, Suganami H, Hamada C, Yoshimura I, Yoshida T, Yoshimura K (2004) Designing a multistage, SNP-based, genome screen for common diseases. J Hum Genet 49:669–676
Schaid DJ, Jacobsen SJ (1999) Biased tests of association: comparisons of allele frequencies when departing from Hardy–Weinberg proportions. Am J Epidemiol 149:706–711
Sing F, Haviland B, Reilly L (1996) Genetic architecture of common multifactorial diseases. In: Variation in the Human Genome (Ciba Foundation Symposium 1997). Wiley, Chichester, pp 211–229
Wittke-Thompson JK, Pluzhnikov A, Cox NJ (2005) Rational Inferences about Departures from Hardy–Weinberg Equilibrium. Am J Hum Genet 76:967–986
Wright S (1951) The genetical structure of populations. Ann Eugen 15:23–354
Wright S (1965) The interpretation of population structure by F-statistics with special regard to systems of mating. Evolution 19:395–420
Wright AF, Carothers AD, Pirastu M (1999) Population choice in mapping genes for complex diseases. Nat Genet 23:397–404
Xu J, Turner A, Little J, Bleecker ER, Meyers DA (2002) Positive results in association studies are associated with departure from Hardy–Weinberg equilibrium: hint for genotyping error? Hum Genet 111:573–574
Yoshida T, Yoshimura K (2003) Outline of disease gene hunting approaches in the Millennium Genome Project of Japan. Proc Jpn Acad Ser B 79:34–50
Zaykin DV, Meng Z, Ghosh SK (2004) Interval estimation of genetic susceptibility for retrospective case-control studies. BMC Genet 5:9
Acknowledgments
We thank Professor Toshiya Sato at Kyoto University and Dr. Takashi Sozu at Tokyo University of Science for their valuable advice in improving this paper. We are grateful to the anonymous reviewers for their useful comments, which greatly improved this paper. This study was supported by the Program for Promotion of Fundamental Studies in Health Sciences of the National Institute of Biomedical Innovation of Japan.
Author information
Authors and Affiliations
Corresponding author
Appendix: Mathematical details
Appendix: Mathematical details
Let the population probabilities of genotypes “XX”, “Xx”, and “xx” be π1, π2 and π3 (π1 + π2 + π3=1), respectively, then those of alleles “X” and “x” for a SNP are given by Eq. 10.
When we use the fixation index (Li et al. 1953) defined by
(πi; i=1, 2, 3) are expressed as Eq. 12.
Therefore, (π1, π2, π3) is equivalent to (P 1, P 2, F).
When the Hardy–Weinberg equilibrium (HWE) holds, F=0 and Eq. 12 reduces to Eq. 13; that is, the second term in the right side of Eq. 12 represents the degree of disequilibrium.
For a random sample of size n, the observed frequency (n 1, n 2, n 3); (n 1 + n 2 + n 3=n) of genotypes (XX, Xx, xx) is distributed as trinomial distribution Tn(n; π1, π2, π3) and, therefore, the maximum likelihood estimator of πi (i=1, 2, 3) is p i=n i/n (i=1, 2, 3). Likewise, the maximum likelihood estimators of P 1 and allele odds \({P_{1} } \mathord{\left/ {\vphantom {{P_{1} } {{\left({1 - P_{1} } \right)}}}} \right. \kern-\nulldelimiterspace} {{\left({1 - P_{1} } \right)}}\;\hbox{are}\; \hat{P}_{1} = p_{1} + {p_{2}} \mathord{\left/ {\vphantom {{p_{2} } 2}} \right. \kern-\nulldelimiterspace} 2 = {{\left({2n_{1} + n_{2} } \right)}} \mathord{\left/ {\vphantom {{{\left({2n_{1} + n_{2} } \right)}} {{\left({2n} \right)}}}} \right. \kern-\nulldelimiterspace} {{\left({2n} \right)}},\;\hbox{and}\;{\hat{P}_{1} } \mathord{\left/ {\vphantom {{\hat{P}_{1} } {{\left({1 - \hat{P}_{1} } \right)}}}} \right. \kern-\nulldelimiterspace} {{\left({1 - \hat{P}_{1} } \right)}},\) respectively.
Since the means, variances, and covariance of p 1 and p 2 are given (Bishop et al. 1975; Agresti 2001) by
the mean and variance of \(\hat{P}_{1}\) is, after a simple but tedious algebra, derived as Eqs. 15 and 16.
When F=0, the last term is the well-known formula for binomial proportion for the size 2n and probability P 1. It reflects that the distribution of the frequency of allele X under HWE is the same as that of allele X randomly chosen from 2n alleles with P 1 as the proportion of X.
Since \(\hat{P}_{1}\) tends to P 1 in probability when n tends to infinity, the logarithm of estimated allelic odds, \(\log {\left({{\hat{P}_{1}} \mathord{\left/ {\vphantom {{\hat{P}_{1} } {{\left({1 - \hat{P}_{1} } \right)}}}} \right. \kern-\nulldelimiterspace} {{\left({1 - \hat{P}_{1} } \right)}}} \right)},\) can be approximated by the first order Taylor expansion as Eq. 17.
Consequently, the mean and variance of \(\log {\left({{\hat{P}_{1} } \mathord{\left/ {\vphantom {{\hat{P}_{1} } {{\left({1 - \hat{P}_{1} } \right)}}}} \right. \kern-\nulldelimiterspace} {{\left({1 - \hat{P}_{1} } \right)}}} \right)}\) are asymptotically approximated by Eqs. 18 and 19:
When we consider the populations of cases and controls of a disease, the association between allele and disease is conventionally represented by the allele odds ratio ψ defined by Eq. 20, where the case and the control are differentiated with the second subscript 1 (case) and 2 (control).
Consider we have random samples of size n .1 and n .2 from cases and controls, respectively. Then the maximum likelihood estimator \(\hat{\psi}\) of ψ is given by Eq. 21, where \(\hat{P}_{11}\;\hbox{and}\;\hat{P}_{12}\) are the maximum likelihood estimators based on samples of case and control, respectively.
Since the sample of case and that of control can be assumed independent, we obtain Eqs. 22 and 23.
where F 1 and F 2 are fixation indices of case and control, respectively.
When we construct an asymptotic confidence interval of log (ψ) with confidence level 1−α, we should replace \(V{\left\{ {\log {\left({\hat{\psi }} \right)}} \right\}}\) with its estimator given by Eq. 24.
where \(\hat{F}_{1}, \hat{F}_{2}\) are as follows:
Rights and permissions
About this article
Cite this article
Sato, Y., Suganami, H., Hamada, C. et al. The confidence interval of allelic odds ratios under the Hardy–Weinberg disequilibrium. J Hum Genet 51, 772–780 (2006). https://doi.org/10.1007/s10038-006-0020-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-006-0020-6
Keywords
This article is cited by
-
Susceptibility to leishmaniasis is affected by host SLC11A1 gene polymorphisms: a systematic review and meta-analysis
Parasitology Research (2019)
-
Impact of Hardy–Weinberg equilibrium deviation on allele-based risk effect of genetic association studies and meta-analysis
European Journal of Epidemiology (2010)
-
Variance estimation of allele-based odds ratio in the absence of Hardy–Weinberg equilibrium
European Journal of Epidemiology (2008)