Abstract
Susceptibility to lung cancer has been shown to be modulated by inheritance of polymorphic genes encoding cytochrome P450 1A1 (CYP1A1) and glutathione S transferases (GSTM1 and GSTT1), which are involved in the bioactivation and detoxification of environmental toxins. As the incidence of lung cancer is known to differ according to ethnicity, we have conducted a case-control study of 146 South Indian lung cancer patients along with 146 healthy controls, to assess any association between CYP1A1, GSTM1 and GSTT1 polymorphisms, either separately or in combination, with the likelihood of development of lung cancer in our population. The current weight of evidence from our study indicated that the frequency of CYP1A1 MspI homozygous variant alleles was significantly higher in cases (OR=3.178). We observed a considerable difference in the GSTT1 null deletion frequency in this population when compared with other populations (OR=2.472, 95% CI: 1.191–5.094, P=0.014). There was no relative risk in GSTM1 null genotype when analysed singly (P=0.453). Considering genotype combinations, risk of lung cancer increased remarkably significantly in individuals having one variant allele of CYP1A1, GSTM1, or GSTT1, suggesting gene–gene interactions. Rare genotypic combinations (such as CYP1A1 wild GSTM1 or GSTT1 either null; CYP1A1 variant both GSTM1 and GSTT1 present; CYP1A1 variant GSTM1 or GSTT1 either null), were at higher risk compared to the reference group. Moreover, patients who had smoked <20 pack years and harboured the CYP1A1 variant allele or the GSTT1 null genotype also had a significant risk of lung cancer. Hence our study—the first to analyse a South Indian population—suggests the importance of combined CYP1A1, GSTM1 and GSTT1 polymorphisms in the development of smoking-induced lung cancer.
Similar content being viewed by others
Introduction
Lung cancer is the leading cause of cancer-related deaths in both men and women worldwide. Molecular epidemiology of lung cancer has received widespread attention because the primary etiology, namely tobacco smoking, is well established, but it is also known that only some smokers develop lung cancer while others do not (Hecht 2002). The influence of environment–gene interactions on lung carcinogenesis has been well demonstrated by phase I and II enzymes that are involved in the metabolic activation and detoxification of carcinogens primarily present in tobacco smoke. Functional polymorphisms in genes encoding xenobiotic metabolizing enzymes such as phase I cytochrome P-450s, cytochrome P450 1A1 (CYP1A1) and glutathione S-transferase Mu (GSTM1), as well as theta (GSTT1) phase II detoxifying enzymes, which are involved in the formation and elimination of carcinogens, have been extensively studied as possible modulators of risk for lung cancer that could explain varying susceptibilities to the disease (Taningher et al. 1999).
Human cytochrome P450 enzymes, which represent a large multigene family with differing substrate specificities, are important in phase I detoxification reactions (Nebert 1991). The CYP1A1 (MIM#108330) gene is involved in the activation step in the metabolism of polycyclic aromatic hydrocarbons (PAHs), such as those found in tobacco smoke, converting them to carcinogens (Gonzalez 1990). Several restriction fragment length polymorphism (RFLP) patterns in the CYP1A1 gene have been identified, including an MspI RFLP produced by a 3801C→T mutation in the 3′-non-coding region (3′-UTR), 250 bp downstream of the polyadenylation signal (Kawajiri et al. 1990). The CYP1A1 m1 polymorphic allele has been associated with alterations in regulation and transcript half-life, which result in elevated induction of the enzyme, and thus increased levels of activated intermediates (Landi et al. 1994). The variant genotype has a high prevalence among Asians (Garte et al. 2001) compared to Caucasians (Tefre et al. 1991) and African Americans (Taioli et al. 1998).
Glutathione transferases (GSTs) comprise a multigene family encoding enzymes that catalyse the conjugation of glutathione to a wide variety of compounds with an electrophilic centre (Hayes and Pulford 1995). GSTs are divided into four classes: alpha, mu, pi and theta, based on amino acid sequence similarity and antibody cross-reactivity (Pemble et al. 1994). GSTM1 (MIM#138350) is involved in the detoxification of tobacco-related carcinogens, such as epoxides and hydroxylated metabolites of benzo(a)-pyrene (Ketterer et al. 1992), whereas GSTT1 (MIM#600436) is involved in the biotransformation of several low molecular weight toxins such as ethylene oxides, butadiene, etc. (Guengerich et al. 1995), which are constituents of tobacco smoke. GSTT1 and GSTM1 are the most extensively studied genes in the GST gene superfamily. Deletion polymorphisms of GSTM1 and GSTT1 that result in no functional enzymatic activity for each locus have been characterised and assigned. Approximately 50% of Caucasian and Asian populations (Seidegard et al. 1988) show GSTM1 null, and 16–50% of different ethnic and racial populations have both copies of the GSTT1 gene deleted (Pemble et al. 1994). We were interested to test the hypothesis that polymorphisms in genes coding for xenobiotic-metabolizing enzymes like CYP1A1 MspI, GSTM1 and GSTT1 may be causally associated with lung carcinogenesis. It is likely that several genetic polymorphisms cooperate in increasing individual risk. There may be specific genotypes or genotype combinations that greatly increase the risk of developing lung cancer. Therefore, the study of gene–gene interactions might be important to identify high susceptibility subgroups. In view of the prevalence of tobacco smoking, and the increased incidence of lung cancer in India, we investigated the distribution of polymorphisms in the CYP1A1, GSTM1 and GSTT1 genes singly, as well as in combination, in lung cancer patients and healthy controls in a South Indian population to determine whether any of the polymorphisms confer an increased risk of developing lung cancer.
Materials and methods
The present hospital-based case-control study of incident cases of lung cancer was conducted at the Regional Cancer Centre, Thiruvananthapuram, Kerala, India. Eligible cases included all patients with newly diagnosed lung cancer presented between January 2003 and December 2004. All cases were newly diagnosed and previously untreated patients. Histological type and degree of differentiation of all cancer cases were assessed by histological examination of surgery or bronchoscopy specimens. During the study period, we included 146 lung cancer cases (133 males and 13 females) as well as 146 healthy controls (128 males and 18 females). Ethical approval was obtained from the Ethical Committee of the Regional Cancer Centre, Thiruvananthapuram, Kerala, India. Controls of the same geographic origin were randomly selected from individuals who attended the outpatient department of the nearby medical college.
Data and sample collection
A thorough, structured questionnaire was completed by both patient and control groups to provide relevant information regarding the risk factors for lung cancer. The information collected included socio-demographic characteristics such as gender, age, lifetime occupational history (including exposure to known carcinogens), area of origin, family history of cancer among first degree relatives, smoking status, which included smoking duration and pack years smoked, medication history and pre-existence of respiratory or lung diseases. In order for the age and gender distributions of controls to match those of lung cancer patients, most of the controls were age matched and the majority were males. Controls were also interviewed and asked about histories of cancer, occupation and smoking habits. Smoking information included past and/or present smoking status, amount smoked and duration of smoking. Smoking status of the subjects was calculated as the average tobacco consumption expressed in pack years. Pack years were computed as the number of cigarettes smoked per day multiplied by the duration of smoking in years. Blood samples were collected from study subjects after obtaining their written informed consent. Peripheral blood (2 ml) collected from patients and all controls and was stored at −80°C until use.
Statistical methods
The principal measure of interest was the relative risk of lung cancer for individuals with the CYP1A1 MspI, GSTM1 and GSTT1 polymorphisms, compared to individuals without the respective variants. Variables selected from the data set are age, gender, smoking status (never a smoker and long-term smokers), pack years of smoking, and polymorphisms in the CYP1A1, GSTM1 and GSTT1 genes. We estimated the study-specific odds ratios (OR) of lung cancer for each polymorphism using unconditional logistic regression modeling with 95% confidence intervals (CIs), and the difference in genotype prevalence and association between case and control group were assessed and adjusted for age, gender and smoking status. To determine whether the genotype frequencies were significantly different between the patient and control population, a probability of P<0.05 was considered. Age, gender, smoking status and pack years were included as covariates as well as all the possible genotypes studied. GSTM1 and GSTT1 polymorphism was dichotomized into null genotype and wild type, while CYP1A1 MspI polymorphism was categorized into homozygous wild type- and variant allele-containing genotypes.
Besides the main effect of CYP1A1, GSTM1 and GSTT1 polymorphism on lung cancer, we were also interested in the possible combined effect of CYP1A1 variants with GSTM1 and GSTT1 null genotypes. Wild type of CYP1A1 and non-null genotypes of GSTM1 and GSTT1 were used as reference groups to assess the combined effects of the two genes. To evaluate the possible interaction between genetic polymorphisms and smoking, a group of subjects with non-null genotype and no current smoking habits was used as a reference group.
DNA extraction
Genomic DNA was extracted from whole blood using proteinase K followed by phenol–chloroform extraction and ethanol precipitation according to standard procedures (Sambrook et al. 1989) and was stored at −20°C for genotype analysis.
Genotyping
Genotyping of the CYP1A1 MspI polymorphism was performed for 146 lung cancer patients and 146 healthy controls using a polymerase chain reaction-RFLP (PCR-RFLP) method as previously described (Hayashi et al. 1991) with slight modifications. Briefly, PCR amplification of a 340 bp DNA fragment containing an MspI restriction site was performed using the primers 5′-CAGTGAAGAGGTGTAGCCGCT-3′ and 5′-TAGGAGTCTTGTCTCATGCCT-3′. A total of 50–100 ng DNA was amplified in a total volume of 25 μl containing 1× buffer, 0.5 U Taq polymerase, 10 pmol of each primer and 200 μM deoxynucleotide triphosphates. PCR was performed at 95°C for 10 min for the initial denaturation, followed by 35 cycles of denaturation at 95°C for 30 s, annealing at 55°C for 1 min and final extension at 72°C for 1 min. A 10 μl aliquot of PCR product was digested with 10 U MspI restriction enzyme (New England Biolabs, Beverly, MA) at 37°C overnight, and the digested PCR product was then resolved by electrophoresis on a 2.5% agarose gel. Three different genotypes were defined for the individual polymorphism: the homozygous wild type (wt/wt) characterized by an approximately 340 bp fragment, a heterozygous variant (wt/m1) with fragments of 340, 200 and 140 bp, and a homozygous variant (m1/m1) with 200 and 140 bp fragments. Genotypes for the GSTM1 and GSTT1 deletions were determined by multiplex PCR using slightly modified published methods (Bell et al. 1993; Chen et al. 1996). Briefly, GSTM1- and GSTT1-specific primer pairs were used together with a third primer for β-globin as an internal control, in a multiplex PCR analysis. The absence of the GSTM1- (210 bp) and/or GSTT1 (473 bp)-specific PCR product indicated the corresponding null genotype, whereas a β-globin-specific fragment (260 bp) confirmed proper functioning of the reaction. These assays do not distinguish between heterozygous and homozygous GSTM1- and GSTT1-positive genotypes. To ensure laboratory quality control, any sample with ambiguous results was re-tested, and a random selection of 10% of all samples was repeated. No discrepancies were encountered upon replicate testing.
Results
In the present study, a total of 292 study subjects comprising 146 lung cancer patients and 146 healthy controls were recruited. Relevant characteristics of the patient group are given in Table 1. The median age in the patient and control groups was 60.00 (mean age 58.17±10.95) and 57.50 (56.06±10.67) years, respectively. The age range of the patients was 20–80 years old while in controls it was 22–75. The affected and control populations were not significantly different in terms of age and gender by analysis of means and SD. The majority of the lung cancer cases were males (91.09%; females 8.9%). A positive family history of cancer among first degree relatives was observed in 24 patients (16.44%) and 17 (11.64%) normal controls, with a relative risk of OR=1.493, 95% CI: 0.765–2.914, P=0.240. With regard to smoking status, 102 (69.9%) patients were smokers and 44 (30.1%) were non-smokers. The control group included 62 (45.2%) smokers and 84 (57.5%) non-smokers. A significant increase in lung cancer risk was found in subjects having a smoking habit (OR=3.141, 95% CI: 1.939–5.087, P=0.001). When smoking data was stratified into pack years, 61 patients (41.8%) had less than 20 pack years and 41 (28.1%) had more than 20 pack years of smoking. Among the control group, 40 (27.4%) were light smokers and 22 (15.1%) were heavy smokers. Those subjects with <20 pack years had a relative risk of 2.911, 95% CI: 1.696–4.998, P=0.001, while for those with >20 pack years, the risk for lung cancer was much higher (OR 3.557, 95% CI: 1.888–6.703, P=0.001). The PCR-RFLP analysis for the CYP1A1 MspI polymorphism is presented in the Fig. 1. The multiplex PCR analysis for the GSTM1 and GSTT1 polymorphisms, with β-globin as a positive internal control are shown in Fig. 2.
Genotype distribution of CYP1A1, GSTM1 and GSTT1 and lung cancer incidence
Table 2 shows the distribution of individual genotypes for CYP1A1 MspI, GSTM1 and GSTT1 in the study population. The proportions of homozygous wild (wt/wt), heterozygous variant (wt/m1) and homozygous variant (m1/m1) genotypes in CYP1A1 were 71 (48.6%), 53 (36.3%) and 22 (15.1%) in the patient group. On the other hand, the control group showed 93 (63.7%), 45 (30.8%) and 8 (5.5%), respectively. Evaluation of the CYP1A1 MspI polymorphism analysis revealed that, compared to the homozygous wild type, the homozygous variant genotype had a significant higher risk, with an adjusted OR of 3.178 (95% CI: 1.294–7.803, P=0.012). Further analysis was carried out by combining CYP1A1 heterozygous and homozygous genotypes, and the OR estimate revealed a significant 1.76-fold higher risk of developing lung cancer compared with the wild type genotype (1.072–2.902, P=0.026).
About 107 controls (73.3%) and 100 patients (68.5%) had an intact GSTM1 gene. GSTM1 null genotype was observed in 39 (26.7%) of the healthy controls, but 46 patients (31.5%) had the GSTM1 gene deletion (OR 1.232; 95% CI: 0.714–2.126). However, the OR estimate revealed that these differences did not attain formal statistical significance (P=0.453). Regarding the GSTT1 genotype, the GSTT1 gene was present in 114 patients (78.1%) and 133 controls (91.1%). The frequencies of the GSTT1 null genotype increased in cases, with 32 patients (21.9%) having the GSTT1 deletion as compared to 13 (8.9%) controls. The OR was 2.472 (95% CI: 1.191–5.094, P=0.014), which was statistically significant. Thus, individuals harboring the null deletion of GSTT1 gene had a 2.4-fold increase in the risk of developing lung cancer.
Any analysis of lung cancer without separation of smokers and non-smokers would be strongly influenced by the results among smokers, who account for at least 90% of cases. We also examined the association between polymorphisms and lung cancer risk by cumulative smoking status and pack years. Detailed results are given in Table 3. Even after adjustment for smoking, age and gender, there was not much effect modification in the genotypes. However, when stratified on the basis of non-smokers vs smokers, the age- and gender-adjusted OR in the group of individuals who had smoked seemed to indicate a significant 2.2-fold risk associated with the GSTT1 null genotype (OR=2.242, 95% CI: 1.020–4.929, P=0.045), the CYP1A1 homozygous mutant (OR=2.947, 95% CI: 1.090–7.968, P=0.033) and the CYP1A1 heterozygous and homozygous mutant genotypes combined (OR=1.832, 95% CI: 1.024–3.277, P=0.041). On stratification based on pack years, those individuals who had <20 pack years and GSTT1 null genotype (OR=3.952, 95% CI: 1.065–14.668, P=0.040), CYP1A1 heterozygous (OR=3.427, 95% CI: 1.126–10.427, P=0.030) and CYP1A1 heterozygous and homozygous mutant combined (OR=3.997, 95% CI: 1.711–13.088, P=0.003) had a 4-fold higher significant relative risk of lung cancer.
Combined genotyping of the CYP1A1, GSTM1 and GSTT1 genes
To further elucidate the genetic factors associated with susceptibility to the development of lung cancer, and to assess the existence of any interactions between the different genotypes studied, the role of combined genetic polymorphisms in the CYP1A1, GSTM1 and GSTT1 genes was investigated. CYP1A1 MspI heterozygote (wt/m1) and homozygote (m1/m1) variants were combined for the analysis. CYP1A1 wild (wt/wt) genotype and individuals with GSTM1 and GSTT1 genes served as the reference group. Analyses were performed considering the two loci, GSTM1 and GSTT1, simultaneously. Fifty-two (35.6%) cases compared to 43 (29.5%) controls had at least one of the alleles deleted. The relative risk was 1.367 (95% CI: 0.812–2.302, P=0.240). When analysis was carried out selecting both null genotypes together, 13 (8.9%) cases compared to 5 (3.4%) controls carried a null deletion at both loci. We found that carrying a deletion at both the GSTM1 and the GSTT1 loci increased the OR to 2.98, but this value was not significant (95% CI: 0.984–9.024, P=0.053).
The genotype combinations between CYP1A1, GSTM1 and GSTT1 examined are given in Table 4. Although combinations between CYP1A1 and GSTM1 indicated a higher OR compared to reference groups (both involving CYP1A1, which had at least one variant allele), formal statistical significance was not obtained. Individuals who possessed a CYP1A1 wild GSTM1− genotype had a 1.43-fold increased risk of cancer (95% CI: 0.706–2.927, P=0.317), and those with CYP1A1 variant GSTM1+ genotype had a 1.7-fold higher risk (OR=1.726, 95% CI: 0.968–3.076, P=0.064). Similarly, those with both variant alleles, CYP1A1 variant GSTM1−, also had a significantly higher OR (OR=2.124, 95% CI: 0.965–4.671, P=0.061). Gene combinations of CYP1A1 wild GSTT1− (OR= 4.469, 95% CI: 1.632–12.239, P=0.004), CYP1A1 variant GSTT1+ (2.106, 95% CI: 1.235–3.592, P=0.006) and CYP1A1 variant GSTT1− (OR=2.472, 95% CI: 0.895–6.826, P=0.081) also gave substantially higher risk compared to the reference group.
In addition, we also evaluated whether lung cancer risk is modified by rare genotype combinations of CYP1A1 MspI, GSTM1 and GSTT1. The ORs for genotype combinations of these three genes in the risk of lung cancer among the study population are shown in Table 5. Six different genotypes from different combinations of these genes were analysed. Although the sample size was small, the genotype combinations had ORs >2. It was noteworthy that three combinations, i.e. (1) CYP1A1 wild GSTM1 or GSTT1 either null (OR=2.230, 95% CI: 1.105–4.501, P=0.025), (2) CYP1A1 variant both GSTM1 and GSTT1 present (OR=2.331, 95% CI: 1.246–4.361, P=0.008), and (3) CYP1A1 variant GSTM1 or GSTT1 either null (OR=2.323, 95% CI: 1.057–5.102, P=0.036) were significantly associated with lung cancer. Even though the genotype combinations GSTM1 null, GSTT1 null, CYP1A1 wild (OR=4.401, 95% CI: 0.819–23.662, P=0.084) and GSTM1 null, GSTT1 null, CYP1A1 variant (OR=3.747, 95% CI: 0.084–17.038, P=0.087) appeared to confer a higher risk, there was no statistical significance.
Discussion
A plethora of studies have examined the risk for lung cancer among individuals possessing variant forms of CYP1A1, GSTM1 and GSTT1. Polymorphic genes involving metabolic polymorphisms almost universally exhibit ethnic and racial variation (Garte et al. 2001), and thus may have large population impacts due to a high prevalence of the relevant polymorphism. Among several candidates for high-risk alleles for lung cancer, CYP1A1, GSTM1 and GSTT1 have been investigated most extensively because of their potential involvement in carcinogenesis. The levels of expression and catalytic activities of cytochrome P450 and GSTM1 and GSTT1 enzymes in lungs, and their metabolic balance, may be an important determinant host factor underlying lung cancer.
In the present analysis, we evaluated the effect of genetic polymorphisms in a series of South Indian lung cancer patients and controls. Worldwide, numerous studies investigating the association between CYP1A1 MspI, GSTM1 and GSTT1 polymorphisms have led to conflicting reports, although several studies have pointed out that individuals polymorphic for the carcinogen-activating or -detoxifying genes are at increased risk for the development of lung cancer. To the best of our knowledge, this is the first report on the combined effect of CYP1A1, GSTM1 and GSTT1 genes in South Indian lung cancer patients.
The CYP1A1 gene is important for the activation of pre carcinogens (Ingelman-Sundberg et al. 2001). In the present study, there was a high frequency of CYP1A1 MspI gene polymorphisms (wt/m1 and m1/m1). A higher prevalence of the CYP1A1 homozygous variant genotype was recorded among lung cancer patients compared to controls with an OR of 3.178 (95% CI: .294–7.803, P=0.012). The MspI polymorphism located in the 3′ flanking region of the CYP1A1 gene was originally found to be associated with lung cancer in Asians (Kawajiri et al. 1990). This was in accordance with the results of a clear association between the m1/m1 genotype and lung cancer risk in Caucasians (Taioli et al. 2003). For further analysis, those having at least one mutant allele of the CYP1A1 MspI gene were combined, and this combination gave a significant OR of 1.763 (95% CI: 1.072–2.902, P=0.026), thus indicating a possible risk genotype. The results of our study are comparable with another study from North India where the relative risk for the carriers of variant CYP1A1 genotypes was high (Sobti et al. 2003). Strong correlations between lung cancer risk and homozygosity for the CYP1A1 variant alleles have been reported in several studies (Xu et al. 1996). A similar relative risk for lung cancer associated with a single mutated allele in CYP1A1 was obtained in a Chilean population (Quinones et al. 1999). We therefore suggest that this mutated genotype plays an important role in the development of lung cancer.
The GSTM1 and GSTT1 members of the glutathione S-transferase multigene family are candidate cancer susceptibility genes because of their ability to regulate the conjugation of carcinogenic compounds to excretable hydrophilic metabolites. Individuals who are carriers of homozygous deletions in the GSTM1 and GSTT1 genes may have an impaired ability to eliminate carcinogenic compounds metabolically and may therefore be at an increased cancer risk. The frequencies of homozygous GSTM1 and GSTT1 deletion carriers are surprisingly high in most human populations, and noticeable differences between ethnic groups exist. Nelson et al. (1995) reported that the GSTM1 null or deletion genotype was present in about 50% of Caucasians, 33% of African Americans and 45% of Japanese, which is in accordance with the studies of Roy et al. (2001). Similar results were obtained in a few other studies (Persson et al. 1999; Chan et al. 2004). Previous studies showed an increased lung cancer risk for GSTM1 null patients independent of ethnic background (Gao and Zhang 1999; Chen et al. 2001). In the case of GSTT1, Nelson et al. (1995) reported that the GSTT1 null genotype is present in 64% of Chinese, 60% of Koreans, 28% of Caucasians and 22% of African Americans. On the basis of our study results, 31.5% of lung cancer cases are carriers of the GSTM1 null genotype, and 21.9% of diseased individuals are homozygous GSTT1 deletion carriers in our geographic region compared to 26.7 and 8.9% in controls. In our study, the GSTM1 null genotype was not associated with an increased risk of lung cancer, although a higher proportion of the patients exhibited the GSTM1 null genotype compared to controls. The OR obtained for the GSTM1 null genotype was 1.262, although this was not statistically significant. For the first time in this population, we have also demonstrated that the GSTT1 null genotype is associated with a significantly increased risk of lung cancer compared with the GSTT1 gene, with an OR of 2.870. This is consistent with some other similar findings (Taioli et al. 2003; Sorensen et al. 2004), but conflicts with other reports (Lan et al. 2000). Regarding the frequency of the homozygous null deletion at the GSTM1 and GSTT1 loci, the putative worst genotype combination of GSTM1 null and GSTT1 null was observed in 13 (8.9%) cases and 5 (3.4%) controls (OR=3.143, 95% CI: 1.075–9.184, P=0.036) with significant risk. Although intra ethnic as well as inter ethnic differences exist in the Indian population, the prevalence of null genotypes in GSTM1 and/or both GSTT1 genes in our population is comparable with that found in other Indian studies (Naveen et al. 2004; Mishra et al. 2004). Thus, our results signify an impact of ethnicity in the distribution of these genotypes. However, in most of studies the ORs have been found to be higher than 1, suggesting a moderate effect of the GSTT1 null genotype (Lewis et al. 2002; Sunaga et al. 2002). As a result of deletions at one or both of these loci and, consequently, less detoxification of xenobiotic toxic substances, an individual may become susceptible to diseases produced by toxic substances present in the environment; hence, finding a positive correlation raises the possibility that the two enzymes are working in tandem rather than in a complementary way.
Overall, the distribution of patients showed that variant genotypes were predominant in the smoking group. In the analysis combining polymorphisms and interactions with smoking, those patients who were smokers and having a GSTT1 null genotype had an increased lung cancer risk (OR=2.242, 95% CI: 1.020–4.929, P=0.045) after adjustment for age and gender. Patients who smoked and had the CYP1A1 mutant allele also had a higher risk (OR=2.947, 95% CI: 1.090–7.968, P=0.033) when compared with their non-smoking counterparts who had wild-type genotypes. In the analysis including heterozygous and homozygous variants together, patients who were smokers and with variant genotypes also had increased ORs when compared to the reference group. Lung cancer risk rose significantly with higher cumulative cigarette consumption, confirming the association with smoking-related lung cancer risk. Stratified analysis between tobacco smoking and variant genotypes revealed, for light smokers with <20 pack years, increasing risks with the presence of at least one copy of the CYP1A1 mutant allele (OR=3.427, 95% CI: 1.126–10.427, P=0.030) or GSTT1 null genotype (OR=3.952, 95% CI: 1.065–14.668, P=0.040) suggesting a gene–environment interaction (Alexandrie et al. 2004). When CYP1A1 heterozygous and homozygous individuals were combined, for light smokers (<20 pack years), cancer risk was increased up to ∼4-fold (OR=3.997, 95% CI: 1.418–11.266, P=0.009). In a study conducted by Ishibe et al. (1997), an approximately 2-fold increase in lung cancer risk among individuals with one or more of the variant alleles was observed among light smokers (defined as having smoked ≤30 pack years). No such increase in risk was found among heavy smokers (>30 pack years). Thus, patients with a combination of high inducibility variants at CYP1A1 together with the GSTT1 null genotype, particularly at low smoking levels, confers a highly significant increase in risk.
Several studies have correlated lung cancer risk with combinations of CYP1A1, GSTM1 and GSTT1 genotypes, which has been of particular interest since the combined action of phases I and II enzymes with increased activation and decreased metabolism of PAHs in tobacco smoke have been hypothesized to lead to increased lung cancer risk. Individuals having a defective genotype for more than one of these genes can thus be expected to be at greater risk for lung cancer than those having a defective genotype of only one gene. The importance of GSTM1, GSTT1 and CYP1A1 genotypes is further emphasized by the increased significance of their interactions. Even though genotype combinations between GSTM1 and CYP1A1 did not achieve formal statistical significance for the combination studied, we obtained an OR of above 1.5 for all combinations. Regarding GSTT1 and CYP1A1 combinations, the risk was found to be significantly high for the genotype combination with the following variant alleles: CYP1A1 wild GSTT1− (OR=4.469, 95% CI: 1.632–12.239, P=0.004) and CYP1A1 variant GSTT1+ (2.106, 95% CI: 1.235–3.592, P=0.006). Co-inheritance of two variants, such as CYP1A1 polymorphism with GSTM1/GSTT1 null genotype was associated with increased cancer risk in several studies (Hayashi et al. 1991). This latter study reported a relative risk of 5.8 for lung cancer in individuals with CYP1A1 and GSTM1 null combination genotype. The variant CYP1A1 genotype had a higher risk when present in individuals with the GSTM1 null genotype (Alexandrie et al. 2004). In a study from North India, Sobti et al. (2004) reported a 2-fold elevated risk for lung cancer in individuals with a single copy of the variant CYP1A1 and null GSTM1.
Combinations of the three genes, e.g. (1) CYP1A1 wild (wt/wt), GSTM1, or GSTT1 either null (OR=2.230, 95% CI: 1.105–4.501); (2) CYP1A1 variant (wt/m1 or m1/m1), both GSTM1 and GSTT1 present (OR=2.331, 95% CI: 1.246–4.361); or (3) CYP1A1 variant (wt/m1 or m1/m1) GSTM1 or GSTT1 either null (OR=2.323, 95% CI: 1.057–5.102) were significantly associated with lung cancer with high OR. There was a trend of 4.4-fold increased relative risk for the GSTM1 null, GSTT1 null, CYP1A1 wild genotype combination (OR=4.401, 95% CI: 0.819–23.662) and a 3.5-fold increased risk, although not statistically significant (OR=3.747, 95% CI: 0.824–17.038), in individuals possessing the GSTM1 null, GSTT1 null, CYP1A1 variant genotype combination. A similar combination association of similar combination genotypes with lung cancer has been reported previously (Dialyna et al. 2003). Due to gene–environment interactions, the effects of inheritance at independent loci can be additive, and each locus contributes incrementally to cancer development or epistasis in which inheritance at multiple loci is required to develop lung cancer. Thus, inheritance of variant genotypes in patients can be additive and may contribute to the development of lung cancer with different susceptibility in the presence of environmental exposures like smoking compared to individuals with a single common variant of the gene (Vineis 2002).
Certain limitations that may give rise to some uncertainty regarding our results should be mentioned. In the present study, the small sample size might have impaired our ability to recognize association with smaller effects. Although combined genotype comparison of a relatively small sample size might show bias and uncertainty, we observed a significant trend towards association between these gene polymorphisms and the development of lung cancer. In conclusion, the results from this study indicate that the CYP1A1 homozygous variant genotype and GSTT1 null genotypes are more strongly associated with the risk of lung cancer. This study suggest a plausible combined role for these enzymes in lung cancer susceptibility. The presence of variant CYP1A1 protein may result in increased formation of carcinogenic metabolites due to the hyperactive phase I enzyme, with detoxification of these reactive metabolites being restrained by the absence of functional phase II enzymes like GSTM1 and GSTT1, which might have a role in the initiation or progression of lung cancer. It seems that combinations of rare metabolic genotypes should be considered more appropriate for cancer risk assessment rather than individual genotypes. The associations discussed in this paper demonstrate the importance of genetic variation in phase I and II genes in lung cancer susceptibility and it is conceivable that these variants will interact with environmental carcinogens and that certain combinations of genotypes will better define at-risk groups. Thus, the high public health significance of these low prevalence genes will hopefully allow us to obtain more mechanistic insights into human lung carcinogenesis, as well as targeted-preventive approaches to individuals at risk.
References
Alexandrie AK, Nyberg F, Warholm M, Rannug A (2004) Influence of CYP1A1, GSTM1, GSTT1, and NQO1 genotypes and cumulative smoking dose on lung cancer risk in a Swedish population. Cancer Epidemiol Biomarkers Prev 13:908–914
Bell DA, Taylor JA, Paulson DF, Robertson CN, Mohler JL, Lucier GW (1993) Genetic risk and carcinogen exposure: a common inherited defect of the carcinogen-metabolism gene glutathione S-transferase M1 (GSTM1) that increases susceptibility to bladder cancer. J Natl Cancer Inst 85:1159–1164
Chan YM, Tan UKC, Ip MS, Tsang KW, Ho SP, Ho JC, Chan H, Lam WK (2004) Lung cancer susceptibility and polymorphisms of glutathione-S-transferase genes in Hong Kong. Lung Cancer 45:155–160
Chen H, Sandler DP, Taylor JA, Shore DL, Liu E, Bloomfield CD, Bell DA (1996) Increased risk for myelodysplastic syndromes in individuals with glutathione transferase theta 1 (GSTT1) gene defect. Lancet 347:295–297
Chen S, Xue K, Xu L, Ma G, Wu J (2001) Polymorphisms of the CYP1A1 and GSTM1 genes in relation to individual susceptibility to lung carcinoma in Chinese population. Mutat Res 458:41–47
Dialyna IA, Miyakis S, Georgatou N, Spandidos DA (2003) Genetic polymorphisms of CYP1A1, GSTM1 and GSTT1 genes and lung cancer risk. Oncol Rep 10:1829–1835
Gao Y, Zhang Q (1999) Polymorphisms of the GSTM1 and CYP2D6 genes associated with susceptibility to lung cancer in Chinese. Mutat Res 444:441–449
Garte S, Boffetta P, Caporaso N, Vineis P (2001) Metabolic gene allele nomenclature. Cancer Epidemiol Biomarkers Prev 10:1305–1306
Gonzalez FJ (1990) Molecular genetics of the P-450 superfamily. Pharmacol Ther 45:1–38
Guengerich FP, Thier R, Persmark M, Taylor JB, Pemble SE, Ketterer B (1995) Conjugation of carcinogens by theta class glutathione S-transferases: mechanisms and relevance to variations in human risk. Pharmacogenetics 5:S103–S107
Hayashi S, Watanabe J, Nakachi K, Kawajiri K (1991) Genetic linkage of lung cancer-associated MspI polymorphisms with amino acid replacement in the heme binding region of the human cytochrome P450IA1 gene. J Biochem 110:407–411
Hayes JD, Pulford DJ (1995) The glutathione S-transferase supergene family: regulation of GST and the contribution of the isoenzymes to cancer chemoprotection and drug resistance. Crit Rev Biochem Mol Biol 30:445–600
Hecht SS (2002) Cigarette smoking and lung cancer: chemical mechanisms and approaches to prevention. Lancet Oncol 3:461–469
Ingelman-Sundberg M, Oscarson M, Daly AK, Garte S, Nebert DW (2001) Human cytochrome P-450 (CYP) genes: a web page for the nomenclature of alleles. Cancer Epidemiol Biomarkers Prev 10:1307–1308
Ishibe N, Wiencke JK, Zuo ZF, McMillan A, Spitz M, KT Kelsey (1997) Susceptibility to lung cancer in light smokers associated with CYP1A1 polymorphisms in Mexican- and African-Americans. Cancer Epidemiol Biomarkers Prev 6:1075–1080
Kawajiri K, Nakachi K, Imai K, Yoshii A, Shinoda N, Watanabe J (1990) Identification of genetically high risk individuals to lung cancer by DNA polymorphisms of the cytochrome P450IA1 gene. FEBS Lett 263:131–133
Ketterer B, Harris JM, Talaska G, Meyer DJ, Pemble SE, Taylor JB, Lang NP, Kadlubar FF (1992) The human glutathione S-transferase supergene family, its polymorphism, and its effects on susceptibility to lung cancer. Environ Health Perspect 98:87–94
Lan Q, He X, Costa DJ, Tian L, Rothman N, Hu G, Mumford JL (2000) Indoor coal combustion emissions, GSTM1 and GSTT1 genotypes, and lung cancer risk: a case-control study in Xuan Wei, China. Cancer Epidemiol Biomarkers Prev 9:605–608
Landi MT, Bertazzi PA, Shields PG, Clark G, Lucier GW, Garte SJ, Cosma G, Caporaso NE (1994) Association between CYP1A1 genotype, mRNA expression and enzymatic activity in humans. Pharmacogenetics 4:242–246
Lewis SJ, Cherry NM, Niven RM, Barber PV, Povey AC (2002) GSTM1, GSTT1 and GSTP1 polymorphisms and lung cancer risk. Cancer Lett 180:165–171
Mishra DK, Kumar A, Srivastava DS, Mittal RD (2004) Allelic variation of GSTT1, GSTM1 and GSTP1 genes in North Indian population. Asian Pac J Cancer Prev 5:362–365
Naveen AT, Adithan C, Padmaja N, Shashindran CH, Abraham BK, Satyanarayanamoorthy K, Anitha P, Gerard N, Krishnamoorthy R (2004) Glutathione S-transferase M1 and T1 null genotype distribution in South Indians. Eur J Clin Pharmacol 60:403–406
Nebert DW (1991) Role of genetics and drug metabolism in human cancer risk. Mutat Res 247:267–281
Nelson HH, Wiencke JK, Christiani DC, Cheng TJ, Zuo ZF, Schwartz BS, Lee BK, Spitz MR, Wang M, Xu X (1995) Ethnic differences in the prevalence of the homozygous deleted genotype of glutathione S-transferase theta. Carcinogenesis 16:1243–1245
Pemble S, Schroeder KR, Spencer SR, Meyer DJ, Hallier E, Bolt HM, Ketterer B, Taylor JB (1994) Human glutathione S-transferase theta (GSTT1): cDNA cloning and the characterization of a genetic polymorphism. Biochem J 300:271–276
Persson I, Johansson I, Lou YC, Yue QY, Duan LS, Bertilsson L, Ingelman-Sundberg M (1999) Genetic polymorphism of xenobiotic metabolizing enzymes among Chinese lung cancer patients. Int J Cancer 81:325–329
Quinones L, Berthou F, Varela N, Simon B, Gil L, Lucas D (1999) Ethnic susceptibility to lung cancer: differences in CYP2E1, CYP1A1 and GSTM1 genetic polymorphisms between French Caucasian and Chilean populations. Cancer Lett 141:167–171
Roy B, Majumder PP, Dey B, Chakraborty M, Banerjee S, Roy M, Mukherjee N, Sil SK (2001) Ethnic differences in distributions of GSTM1 and GSTT1 homozygous “null” genotypes in India. Hum Biol 73:443–450
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Seidegard J, Vorachek WR, Pero RW, Pearson WR (1988) Hereditary differences in the expression of the human glutathione transferase active on trans-stilbene oxide are due to a gene deletion. Proc Natl Acad Sci USA 85:7293–7297
Sobti RC, Sharma S, Joshi A, Jindal SK, Janmeja A (2003) CYP1A1 and CYP2D6 polymorphism and risk of lung cancer in a North Indian population. Biomarkers 8:415–428
Sobti RC, Sharma S, Joshi A, Jindal SK, Janmeja A (2004) Genetic polymorphism of the CYP1A1, CYP2E1, GSTM1 and GSTT1 genes and lung cancer susceptibility in a north indian population. Mol Cell Biochem 266:1–9
Sorensen M, Autrup H, Tjonneland A, Overvad K, Raaschou-Nielsen O (2004) Glutathione S-transferase T1 null-genotype is associated with an increased risk of lung cancer. Int J Cancer 110:219–224
Sunaga N, Kohno T, Yanagitani N, Sugimura H, Kunitoh H, Tamura T, Takei Y, Tsuchiya S, Saito R, Yokota J (2002) Contribution of the NQO1 and GSTT1 polymorphisms to lung adenocarcinoma susceptibility. Cancer Epidemiol Biomarkers Prev 11:730–738
Taioli E, Ford J, Trachman J, Li Y, Demopoulos R, Garte S (1998) Lung cancer risk and CYP1A1 genotype in African Americans. Carcinogenesis 19:813–817
Taioli E, Gaspari L, Benhamou S, Boffetta P, Brockmoller J, Butkiewicz D, Cascorbi I, Clapper ML, Dolzan V, Haugen A, Hirvonen A, Husgafvel-Pursiainen K, Kalina I, Kremers P, Le Marchand L, London S, Rannug A, Romkes M, Schoket B, Seidegard J, Strange RC, Stucker I, To-Figueras J, Garte S (2003) Polymorphisms in CYP1A1, GSTM1, GSTT1 and lung cancer below the age of 45 years. Int J Epidemiol 32:60–63
Taningher M, Malacarne D, Izzotti A, Ugolini D, Parodi S (1999) Drug metabolism polymorphisms as modulators of cancer susceptibility. Mutat Res 436:227–261
Tefre T, Ryberg D, Haugen A, Nebert DW, Skaug V, Brogger A, Borresen AL (1991) Human CYP1A1 (cytochrome P(1)450) gene: lack of association between the MspI restriction fragment length polymorphism and incidence of lung cancer in a Norwegian population. Pharmacogenetics 1:20–25
Vineis P (2002) The relationship between polymorphisms of xenobiotic metabolizing enzymes and susceptibility to cancer. Toxicology 181–182:457–462
Xu X, Kelsey KT, Wiencke JK, Wain JC, Christiani DC (1996) Cytochrome P450 CYP1A1 MspI polymorphism and lung cancer susceptibility. Cancer Epidemiol Biomarkers Prev 5:687–692
Acknowledgements
L.S. is a recipient of an Indian Council for Medical Research (ICMR) Senior Research Fellowship. This study was supported by grants from the Department of Science and Technology, Government of India.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sreeja, L., Syamala, V., Hariharan, S. et al. Possible risk modification by CYP1A1, GSTM1 and GSTT1 gene polymorphisms in lung cancer susceptibility in a South Indian population. J Hum Genet 50, 618–627 (2005). https://doi.org/10.1007/s10038-005-0303-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-005-0303-3
Keywords
This article is cited by
-
Health effects associated with smoking: a Burden of Proof study
Nature Medicine (2022)
-
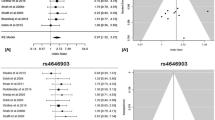
A comprehensive meta-analysis and a case–control study give insights into genetic susceptibility of lung cancer and subgroups
Scientific Reports (2021)
-
Genetic polymorphism of lysyl oxidase, glutathione S-transferase M1, glutathione-S-transferase T1, and glutathione S-transferase P1 genes as risk factors for lung cancer in Egyptian patients
Molecular Biology Reports (2021)
-
Acetylcholinesterase, Butyrylcholinesterase and Glutathione S-Transferase Enzyme Activities and Their Correlation with Genotypic Variations Based on GST M1 and GST T1 Loci in Long Term-Pesticide-Exposed Tea Garden Workers of Sub-Himalayan West Bengal
Toxicology and Environmental Health Sciences (2019)
-
Glutathione S-transferase M1 and T1 null genotype frequency distribution among four tribal populations of western India
Journal of Genetics (2018)