Abstract
Mutations in the CLCN5 gene have been detected in Dent’s disease and its phenotypic variants (X-linked recessive nephrolithiasis, X-linked recessive hypophosphatemic rickets, and idiopathic low-molecular-weight proteinuria of Japanese children). Dent’s disease is a tubular disorder characterized by low-molecular-weight proteinuria, and nephrolithiasis associated with nephrocalcinosis and hypercalciuria. ClC-5 is the first chloride channel for which a definitive role in the trafficking and acidification-dependent recycling of apical membrane proteins has been established. In the course of CLCN5 SSCP analysis in patients with hypercalciuric nephrolithiasis, we detected a novel mutation at intron 2 of the CLCN5 gene, a T-to-G substitution, located 17 bp upstream of the AG acceptor site. To determine the effect of IVS2–17 T>G mutation on the correct splicing of intron 2, we studied ClC-5 transcripts in a patient’s peripheral blood leukocytes by means of quantitative comparative RT/PCR, and found a new ClC-5 5’ UTR isoform characterized by the untranslated exon 1b and by retention of intron 1b. This new isoform—isoform B1—was not correlated with mutation since it was detected also in control leukocytes and in renal tissues of kidney donors, thus confirming its physiological role. By RACE analysis we determined the putative transcriptional start site which is located at intron 1a, 251 nt upstream of the first nucleotide of the untranslated exon 1b. ORF analysis revealed that intron 1b retention in isoform B1 stabilizes the initiation of translation to the AGT at position 297 of the ClC-5 cDNA coding region.
Similar content being viewed by others
Introduction
The chloride channel ClC-5 belongs to the ClC family of voltage-gated chloride channels (ClC 1–7, ClC-Ka and ClC-Kb) that regulates cell volume, membrane excitability, and transepithelial transports (Jentsch et al. 2002). Some ClC proteins are membrane channels, while others are thought to reside predominantly in intracellular membranes. In the kidney, ClC-5 is expressed in the S3 segment of proximal tubules, in the ascending limb of Henle’s loop, and in the intercalated cells of the collecting duct (Luyckx et al. 1998). CLCN5 mutations inactivating ClC-5 have been detected in Dent’s disease and in three other disorders referred as X-linked recessive nephrolithiasis (XRN), X-linked recessive hypophosphatemic rickets (XLRH), and idiopathic low-molecular-weight proteinuria of Japanese children (JILMWP), which are now considered phenotypic variants of a unique disease (Thakker 2000). Dent’s disease is a tubular disorder characterized by low-molecular-weight proteinuria, and nephrolithiasis associated with nephrocalcinosis and hypercalciuria. This disease is clinical relevant because of its major risk of evolution into uremia between the 3rd and 5th decades of life. ClC-5 is the first chloride channel for which a definitive role in the trafficking and acidification-dependent recycling of apical membrane proteins has been established (Gunther et al. 1998; Devuyst et al. 1999; Sakamoto et al. 1999; Sayer and Simmons 2002). Animal models of knockout ClC-5 mice demonstrated that impairment of the endocytotic traffic is the major cause not only of tubular proteinuria but also of hypercalciuria and consequently kidney stones (Piwon et al. 2000; Wang et al. 2000; Gunther et al. 2003). However, given the high interfamilial and intrafamilial phenotypic variability of Dent’s disease and its variants, it is conceivable that modifying genes could determine the phenotypic manifestation of the disease.
Very recently, it has been hypothesized that the splicing machinery could be one of such modifying systems since changes in the level of normal transcripts or in the relative pattern of different mRNA isoforms could affect disease expression leading to phenotypic variability (Nissim-Rafinia and Kerem 2002). Two different 5’ UTR ends of ClC-5 cDNA have been described in the kidney. One of them (type B) was first described by Fisher et al. (1994, 1995) and contains the untranslated exon 1b. The other (type A) was recently identified by Hayama et al. (2000) and consists in a newly untranslated exon 1a that was found 2.1 kb upstream of exon 2 containing the translation start site. By RNase protection assay and primer extension analysis, only for the type A isoform was the transcription start determined. These different 5’ UTR ends constitute the two mRNA isoforms, type A and type B, whose expression in the kidney or in other tissues have not yet been determined. In the course of CLCN5 mutational analysis in patients with hypercalciuric nephrolithiasis, we detected a novel mutation at intron 2 of the CLCN5 gene, a T-to-G substitution, 17 bp upstream of the AG acceptor site. To determine the effect of IVS2–17 T>G mutation on the correct splicing of intron 2, we studied CLCN5 transcripts in a patient’s peripheral blood leukocytes and found a new ClC-5 5’ UTR isoform characterized by the untranslated exon 1b and by retention of intron 1. This isoform was not correlated with mutation since it was detected also in control leukocytes and in renal tissues of kidney donors, thus confirming its physiological role.
Materials and methods
Patient
The patient was a 26-year-old man with hypercalciuria, bilateral nephrolithiasis by the age of 18 years, and a family history of nephrolithiasis. Proteinuria was absent as well as other typical signs of Dent’s disease tubulopathy.
Mutational analysis of the CLCN5 gene
The patient’s and parents’ leukocyte DNA was extracted by NucleoSpin Blood Quick Pure minicolumns (Macherey-Nagel GmbH & Co. KG, Duren, Germany) and used with specific primers for PCR amplification of the 12 exons and exon-intron boundaries of CLCN5 gene. Primers and PCR conditions were the same used by Lloyd et al. (1997). Single strand conformation polymorphism (SSCP) analysis was applied to detect DNA mutations as previously described (Anglani et al. 1993). PCR products demonstrating mobility shift under specific electrophoretic conditions were directly sequenced.
RNA extraction from leukocytes and kidney biopsies
Total RNA was extracted from the patient’s frozen whole blood by RNAzol B method (Biotex). As control samples, leukocytes of normal individuals, basal biopsies of transplanted kidneys—performed before reperfusion and collected after informed consent—and leukocytes of renal transplant recipients were used. RNAzol B method and QIAmp RNA Blood Mini kit (Qiagen) were used for purifying RNA from biopsies and leukocytes respectively. RNA extracted from the primary culture of human proximal tubular cells was also utilized.
RT/PCR analysis of ClC-5 mRNA
To analyze the CLCN5 gene expression, three sets of primers were designed (Fig. 1). One set is specific for the common region of the two isoforms spanning the translated exon 2 and exon 6 (primers Ex 2–3F- Ex 6R). The other two are constituted by the same reverse (Ex 4R) while the forward one was 5’-end specific, i.e., located in ex 1a (ex 1aF) or in exon 1b (Ex 1bF). The nucleotide composition of primers and the length of amplified products are reported in Table 1.
One hundred nanograms of total RNA were reverse-transcribed in a total volume of 20 µl containing 5 mM MgCl2, 1 mM dNTPs (Boehringer), 2.5 µM random hexamers (Applied Biosystem), 1 U RNase inhibitor (Applied Biosystem) and 2.5 U MuLV reverse transcriptase (Applied Biosystem) in a buffer of 50 mM KCl, 10 mM Tris HCl pH 8.3. The reaction was carried out at 42°C for 30 min, and 5 min at 99°C in a Thermal cycler (MJ Research). An aliquot of 2 µl of RT reaction was used to amplify the three different regions of ClC-5 mRNA in a final volume of 25 µl containing 1.5 mM MgCl2, 0.2 mM dNTPs, 0.4 µM specific primers, 2 U JumpStart Taq (Sigma) in 50 mM KCl, 10 mM Tris HCl pH 8.3. Amplification profile was the same for each primer set and consisted of 45 s at 94°C (denaturation), 45 s at 60°C (annealing), and 2 min at 72°C (extension). PCR amplification of glyceraldheyde-3-phosphate dehydrogenase (GAPDH) as internal standard was performed under the same conditions.
To obtain quantitative data, the kinetic strategy was applied as described in order to determine the opportune cycles in which PCR products exponentially increased (Ceol et al. 2001). The specific cycle numbers are reported in Table 1.
RT/PCR products were analyzed by polyacrylamide gel electrophoresis (PAGE) followed by silver staining as previously described (Del Prete et al. 1998). Gel bands were quantified directly in the gel by densitometric analysis using Gel-Pro Analyzer software (Media Cybernetics). The relative levels of each mRNA were reported calculating the ratio of the target gene and GAPDH transcripts.
The 5’-rapid amplification of cDNA ends (RACE) of human ClC-5 cDNA
A 5’ RACE system for rapid amplification of cDNA ends (Clontech) was used to clone more of the 5’ end portion of the human ClC-5 cDNA. Matathon-Ready human kidney cDNA (Clontech) was utilized as a template for RACE reactions. Three gene-specific antisense primers (GSP) were designed to isolate the 5’ ends of isoforms A and B: rRACE1a (5’ -cagcccgcatttggtcattcaggttg- 3’), rRACE1b (5’ -tggctaaaggcagtgtgaatggaaagca- 3’), and rRACE Ex2 (5’ -acaccagggattggctcctccaagaa- 3’) located respectively in exon 1a, exon 1b, and exon 2 (Fig. 1). Before RACE, a PCR with internal known primers was performed to verify that the cDNA library contained ClC-5 cDNA. RACE amplifications were made with the forward adaptor primer AP1 and each of the three GSPs using Advantage 2 polymerase (Clontech). For the isolation of more specific 5’ end fragments, a second nested PCR, on the first RACE performed with the GSP rRACE Ex2 primer, was performed utilizing the AP2 forward primer and the rRACE1b or rRACE 1a reverse primer. Touchdown thermal cycling conditions for the first amplification were 94°C for 1 min, 5 cycles (94°C 30 s, 72°C 3 min), 5 cycles (94°C 30 s, 70°C 3 min) and 25 cycles (94°C 30 s, 68°C 3 min). The nested PCR conditions were 94°C 1 min followed by 25 cycles each of 94°C 30 s, 68°C 3 min. RACE products were analyzed by PAGE on 7% polyacrylamide gel, isolated for sequencing on 2% low-melting agarose gel, and then purified using Sephaglas BandPrep kit (Amersham Biosciences).
Sequencing
Direct sequencing of PCR, RT/PCR, and 5’ RACE products was performed using ABI Prism GENESCAN 373 A DNA sequencer and the BigDye Terminator Cycle Sequencing Ready Reaction Kit (Applied Biosystem).
Sequence comparison
To compare ClC-5 5’ UTR cDNAs and coding region with its genomic sequences, the NCBI Blast 2 sequence alignment program was used. The coding region of ClC-5 cDNA from human kidney (GeneBank accession number MN000084) and the human DNA sequence from clone RP13–377G1 on chromosome Xp11.22–11.3 (GenBank accession number version GI: 18121562) and from the CLCN5 promoter region (GeneBank accession number AB0205b7) were used for comparison. Amino acid sequence analysis was subjected to computational analysis with the ORF Finder program.
Results
CLCN5 gene mutational analysis
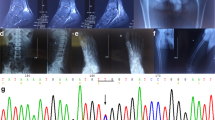
We screened all the coding sequences (exons 2 to 12) and the exon-intron boundaries of the CLCN5 gene by SSCP analysis. As shown in Fig. 2A, the electrophoretic pattern of patient PCR products obtained with primers spanning intron 2 - exon 3 - intron 3 boundaries was clearly different from that of controls’ and parent’s samples. Sequencing of purified PCR products revealed a T-to-G substitution at position –17 bp of the invariant consensus acceptor site sequence of intron 2 (IVS2 –17 T>G) (Fig. 2B and C). The nucleotide substitution was a de novo mutation since it was not present in the mother. The mutation did not affect directly the AG acceptor site but was located in a region that could be critical for the corrected intron 2 splicing. The effect of this acceptor splice-site mutation on ClC-5 transcript was studied analyzing total RNA extracted from leukocytes. RT/PCR and SSCP analysis of RT/PCR products revealed, in both patient and control samples, an apparently normal mRNA except for the presence of a longer isoform B (Fig. 3B). In fact, the expected size of RT/PCR amplicons, generated by primers Ex 1bF-Ex 4R, was 514 bp while the observed one was greater (~ 650 bp). RT/PCR analysis, carried out in renal biopsies and leukocytes from kidney recipients, confirmed the presence of the 650 bp product in almost all samples (Fig. 3B). Sequencing and Blast 2 analysis disclosed that the longer B isoform resulted from retention of intron 1 in the mature ClC-5 mRNA (Fig. 4). ORF analysis revealed that retention of intron 1 stabilizes the initiation of translation to the AGT at position 297 of the ClC-5 cDNA coding region (Fig. 5).
A SSCP analysis of CLCN5 exon 3 and intron-exon boundaries in the genomic DNA of patient (lane 4), parents (lanes 2 and 3) and control (lane 1). MW molecular-weight marker, ΦX Hae III. The profile of DNA fluorescence autosequencing, using forward primer, of B PCR product with altered mobility shift from patient DNA and C PCR product from mother’s DNA
RT/PCR analysis of ClC-5 mRNA in kidney biopsies and leukocytes. MW molecular-weight marker ΦX Hae III, P patient’s leukocytes. RT/PCR products were generated using primer pairs: A Ex 2–3F-Ex 6R (expected band size 573 bp) corresponding to the common region of the two isoforms; B Ex 1bF-Ex 4R (expected band size 514 bp, observed one ~ 650 bp) corresponding to isoform B1; C Ex 1a F-Ex 4R (expected band size 487 bp) corresponding to isoform A; D GAPDH F-GAPDH R housekeeping gene
Expression study of ClC-5 5’ UTR ends
Quantification of 5’ UTR isoforms of ClC-5 mRNA conducted by kinetic comparative RT/PCR in both leukocytes of kidney recipients and in renal biopsies (Table 2, Fig. 3) showed that the isoform A (Fig. 3C) was indeed the most abundant in renal biopsies, whereas the isoform B, described by Fisher et al. (1995), was not present in renal tissue as not in leukocytes. The new 5’ UTR isoform (B1) (Fig. 3B) was expressed at a lower level compared with isoform A in renal tissue, while both isoform A and B1 had low but comparable levels of expression in leukocytes. These levels did not correspond to that detected with the coding region from exon 3 to 6 (Fig. 3A). We hypothesize that unrecognized 5’ UTR ends could be present in leukocytes.
The already described isoform B was indeed detected as a very faint band on total RNA extracted from leukocytes of normal individuals. Using primers Ex 1bF-Ex 4R, the expected 514 bp amplicon was coamplified, along with the 648 bp amplicon in all samples (Fig. 6).
Analysis of ClC-5 cDNA 5’ ends by RACE
Human kidney cDNA containing the ClC-5 transcripts was used to perform RACE PCR. Using primers designed to study ClC-5 expression in renal tissue samples and leukocytes, we confirmed the presence in the library of both the new isoform B1 and isoform A described by Hayama et al. (2000), but we did not succeed in detecting the classical isoform B described by Fisher et al. (1994, 1995) (data not shown). A RACE fragment of 410 bp was obtained using AP2 and rRACE1b primers and was sequenced. We succeeded in extending the 5’ end of the new isoform B by 251 nucleotides beyond exon 1b (Fig. 7). This new putative transcriptional start site is located at intron 1a (at position 2049 of the ClC-5 promoter region DDBJ/EMBL/GeneBank accession number AB020597). Analysis of the 5’ flanking region was performed using www Signal and Promoter Scan IMD Search Service, which revealed the presence of various Sp1 sites, two AP2 elements, a JCV repeated sequence, and an EARLY-SEQ1. Similarly to the promoter of isoform A described by Hayama et al. (2000), the alternative promoter region we found at intron 1a lacked a TATA box, a CAAT box and a CpG island; however, the region contained Sp1 binding sites near the putative transcription start site, a typical feature of a promoter lacking a TATA box.
Discussion
We found a novel mutation affecting the region of intron 2 splice-site consensus sequences of the CLCN5 gene in a patient with hypercalciuric nephrolithiasis. However, since the T-to-G substitution is located 17 bp upstream of the AG acceptor site, its consequence on the mRNA splicing process is unpredictable. RT/PCR analysis of the patient’s leukocyte ClC-5 transcripts revealed the presence of two mRNA of the same size as control samples. SSCP analysis of RT/PCR products showed no mobility shift, thus confirming that the nucleotide composition of both patient and control transcripts was identical. The presence of normal ClC-5 mRNAs in the patient’s leukocytes may suggest that the nucleotide substitution is indeed a polymorphism. However, against this hypothesis, three major points/objections can be raised: (1) nucleotide substitution is a de novo mutation since it is absent in maternal X chromosome, (2) the patient exhibits some clinical signs characteristic of Dent’s disease, and (3) since the patient’s kidney biopsy was not available, we could not exclude that an abnormal splicing of ClC-5 mRNA occurred in renal tissue. In fact, while an increasing body of evidence indicates that RNA sequence elements important for regulation of pre-mRNA splicing can be located outside the traditional splice sites, either internally within the exon or in flanking intron sequences, it is becoming more and more evident that the recognition of splicing regulatory elements by nuclear splicing factors is tissue specific (Brudno et al. 2001), and that mutations in splicing motifs can result in different splicing patterns in different tissues (Càceres and Komblihtt 2002). Furthermore, the IVS2 –17 T>G mutation, belonging to the subclass II splicing mutations, which include mutations in the less-conserved consensus motifs adjacent to both donor or acceptor sites, should lead either to an aberrantly or correctly spliced mRNA (Nissim-Rafinia and Kerem 2002). Accordingly, the expected phenotype should be milder owing to the partial formation of a normal transcript in the affected tissue. Indeed, the patient carrying the mutation showed no other signs of Dent’s tubulopathy except for hypercalciuria and nephrolithiasis.
Studying the effect of the mutation on pre-mRNA splicing, we detected a novel 5’ UTR isoform of ClC-5 transcript present in both renal tissue and in mononuclear peripheral blood cells of normal individuals. This novel isoform, that we called isoform B1, is similar to that described by Fisher et al. (1995) but retains the 134-bp-long intron 1 in the untranslated region of mature ClC-5 mRNA. The isoform was also detected in two different human kidney cDNA libraries purchased by Clontech. In these libraries, by RACE we identified the putative transcriptional start site of the newly identified 5’ UTR isoform. It is localized at intron 1a, 251 nucleotides upstream of the beginning of exon 1b. Analysis of the 5’ flanking region conducted by www Signal Scan IMD Search Service revealed the presence of a putative promoter region. Similarly to the promoter region of isoform A described by Hayama et al. (2000), it lacks a TATA box, a CAAT box, and a CpG island, but it contains Sp1 binding sites near the putative transcription start site, a typical feature of promoter lacking a TATA box.
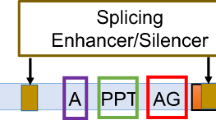
In human pre-mRNAs, the exons are usually short (typically 100–200 bp) and the introns are much longer, averaging about 3 kb. Information for splicing of short introns may be contained entirely within the intron. In particular, accurate short intron recognition in human and plant transcripts is not sufficiently directed by the 5’ and 3’ splice sites, but intronic enhancer motif can dramatically contribute to the accuracy of splicing machinery (Lim and Burge 2001). According to the criterion defined by Lim and Burge (2001), intron 1 of the 5’ UTR ClC-5 mRNA should be considered as a short intron as it is 134 bp long. The invariant consensus splice sites are of classical type GT-AG, so we cannot ascribe the phenomenon of intron retention to the presence of a weak variant GC donor site (Thanaraj and Clark 2001). More likely, the presence of a weak enhancer intronic motif could be the reason for intron retention. Although an accurate estimation of base composition of such intronic motif is still lacking, an intronic putative enhancer motif constituted by the pentamer AAGGG is indeed present in the retained intron 1 (Lim and Burge 2001). Thus, intron 1 retention cannot be ascribed to an inefficient splicing process.
While alternative splicing in the coding region of a gene leads to protein and function diversification, the one occurring in the 5’ UTR region may be relevant to the proper initiation of translation (van der Velden and Thomas 1999). The majority of eukaryotic mRNAs are thought to be translated by a ribosome-scanning mechanism in which the 40S ribosomal subunit binds to the cap structure (cap-dependent translation). However, an alternative cap-independent mechanism of regulation of translation initiation mediated by ribosome binding to internal ribosomal entry site (IRES) has been proposed (Kozak 2001). IRES was first discovered in picornavirus (Jackson and Kaminski 1995) and was also found in a number of growth factors whose translation initiation is specifically regulated during development, growth, and stress (Holcik et al. 2000; van der Velden and Thomas 1999; Shalev et al. 2002; Morrish and Rumsby 2002). We hypothesize that the retention of intron 1 in the ClC-5 B transcript might be explained by the necessity of tubular cells to arrange, under stress conditions, a more efficient site of translation initiation. The data we obtained from the analysis of the expression pattern of ClC-5 5’ UTR isoforms seem to support this hypothesis. Indeed, the originally described isoform B was exclusively found in leukocytes of normal subjects, whereas solely the new isoform with retained intron 1 was observed in renal biopsies from ischemic kidneys or in leukocytes from kidney transplant recipients. It is conceivable that ischemia and/or inflammatory conditions could have conditioned the transcription of this 5’ UTR isoform.
In conclusion, we detected a novel mutation affecting the less conserved consensus motif adjacent to the acceptor site of CLCN5 intron 2. It is a de novo mutation detected in a patient with hypercalciuric nephrolithiasis but without the classical Dent phenotype. Furthermore, we identified a novel 5’ UTR end of ClC-5 mRNA whose putative transcriptional start site is located at intron 1a, 251 nt upstream of the first nucleotide of exon 1b first described by Fisher et al. This newly identified isoform is expressed in leukocytes and renal tissue at a lower level compared with isoform A and is characterized by intron 1 retention. ORF analysis revealed that intron 1 retention in isoform B stabilizes the initiation of translation to the AGT at position 297 of the ClC-5 cDNA coding region. Furthermore, we suggest that intron 1 could contain IRES elements.
References
Anglani F, Murgia A, Bresin S, Bernardi P, Clementi M, Tenconi R (1993) A new disease-causing mutation in the GAP-related domain of the NF1 gene. Hum Mol Genet 2(7):1057–1059
Brudno M, Gelfand MS, Spengler S, Zorn M, Dubchak I, Conboy JG (2001) Computational analysis of candidate intron regulatory elements for tissue-specific alternative pre-mRNA splicing. Nucleic Acids Res 29(11):2338–2348
Càceres JF, Komblihtt AR (2002) Alternative splicing: multiple control mechanisms and involvement in human disease. Trends Genet 18(4):186–193
Ceol M, Forino M, Gambaro G, Sauer U, Schleicher ED, D’Angelo A, Anglani F (2001) Quantitation of TGF-beta 1 mRNA in porcine mesangial cells by comparative kinetic RT/PCR: comparison with nuclease protection assay and in situ hybridization. J Clin Lab Anal 15(4):215–222
Del Prete D, Forino M, Gambaro G, D’Angelo A, Anglani F (1998) Comparative kinetic RT/PCR strategy for the quantitation of mRNAs in microdissected human renal biopsy specimens. Exp Nephrol 6(6):563-567
Devuyst O, Christie PT, Courtoy PJ, Beauwens R, Takker RV (1999) Intra-renal and subcellular distribution of the human chloride channel, ClC-5, reveals a pathophysiological basis for Dent’s disease. Hum Mol Genet 8(2):247–257
Fisher SE, Black GCM, Lloyd SE, Hatchwell E, Wrong O, Thakker RV, Craig IW (1994) Isolation and partial characterization of a chloride channel gene which is expressed in kidney and is a candidate for Dent’s disease (an X-linked hereditary nephrolithiasis). Hum Mol Genet 3(11):2053-2059
Fisher S, Van Bakel I, Lloyd SE, Pearce SHS, Thakker RV, Craig IW (1995) Cloning and characterization of CLCN5, the human kidney chloride channel gene implicated in Dent disease (an X-linked hereditary nephrolithiasis). Genomics 29:598–606
Gunther W, Luchow A, Cluzeaud F, Vandewalle A, Jentsch TJ (1998) ClC-5, the chloride channel mutated in Dent’s disease, colocalizes with the proton pump in endocytotically active kidney cells. Proc Natl Acad Sci USA 95:8075–8080
Gunther W, Piwon N, Jentsch TJ (2003) The ClC-5 chloride channel knock-out mouse an animal model for Dent’s disease. Eur J Physiol 445:456–462
Hayama A, Uchida S, Sasaki S, Marumo F (2000) Isolation and characterization of the human CLC-5 chloride channel gene promoter. Gene 261:355–364
Holcik M, Sonenberg N, Korneluk RG (2000) Internal ribosome initiation of translation and the control of cell death. Trends Genet 16(10):469–473
Jackson RJ, Kaminski A (1995) Internal initiation of translation in eukaryotes: the picornavirus paradigm and beyond. RNA1(10):985–1000
Jentsch TJ, Stein V, Weinreich F, Zdebik AA (2002) Molecular structure and physiological function of chloride channels. Physiol Rev 82(2):503–568
Kozak M (2001) New Ways of Initiating translation in eukariotes?. Mol Cell Biol 21(6):1899–1907
Lim LP, Burge CB (2001) A computational analysis of sequence features involved in recognition of short introns. Proc Natl Acad Sci USA 98(20):11193–11198
Lloyd SE, Gunther W, Pearce SHS, Thomson A, Bianchi ML, Bosio M, Craig IW, Fisher SE, Scheinman SJ, Wrong O, Jentsch TJ, Thakker RV (1997) Characterization of renal chloride channel, CLCN5, mutations in hypercalciuric nephrolithiasis (kidney stones) disorders. Hum Mol Genet 6(8):1233–1239
Luyckx V, Goda FO, Mount DB, Nishio T, Hall A, Hebert SC, Hammod TG, Yu ASL (1998) Intrarenal and subcellular localization of rat CLC5. Am J Physiol 275(5Pt2):F761–F769
Morrish BC, Rumsby MG (2002) The 5’ untranslated region of protein kinase C delta directs translation by an internal ribosome entry segment that is most active in densely growing cells and during apoptosis. Mol Cell Biol 22(17):6089–6099
Nissim-Rafinia M, Kerem B (2002) Splicing regulation as a potential genetic modifier. Trends Genet 18(3):123–127
Piwon N, Gunther W, Schwake M, Bosl MR, Jentsch TJ (2000) ClC-5 Cl- channel disruption impairs endocytosis in a mouse model for Dent’s disease. Nature 408:369–373
Sakamoto H, Sado Y, Naito I, Kwon T, Inoue S, Endo K, Kawasaki M, Uchida S, Nielsen S, Sasaki S, Marumo F (1999) Cellular and subcellular immunolocalization of ClC-5 channel in mouse kidney: colocalization with H+-ATPase. Am J Physiol 277(6):F957-F965
Sayer JA, Simmons NL (2002) Urinary stone formation: Dent’s disease moves understanding forward. Exp Nephrol 10:176–181
Shalev A, Blair PJ, Hoffmann SC, Hirshberg B, Peculis BA, Harlan DM (2002) A proinsulin gene spice variant with increased translation efficiency is expressed in human pancreatic islet. Endocrinology 143(7):2541–2547
Thakker RV (2000) Molecular pathology of renal chloride channels in Dent’s disease and Bartter’s syndrome. Exp Nephrol 8:351–360
Thanaraj TA, Clark F (2001) Human GC-AG alternative intron isoforms with weak donor sites show enhanced consensus at acceptor exon positions. Nucleic Acids Res 29(12):2581–2593
Van der Velden AW, Thomas AAM (1999) The role of 5’ untranslated region of an mRNA in translation regulation during development. Int J Biochem Cell Biol 31:87–106
Wang SS, Devuyst O, Courtoy PJ, Wang X, Wang H, Wang Y, Takker RV, Guggino S, Guggino WB (2000) Mice laking renal chloride channel, CLC-5, are a model for Dent’s disease, a nephrolithiasis disorder associated with defective receptor-mediated endocytosis. Hum Mol Genet 9(20):2937–2945
Acknowledgements
This work was supported by grants from Ricerca Regionale Sanitaria Finalizzata 1999–2001 (no. 798/03/98) and from Ministero dell’Istruzione, dell’Università e della Ricerca 2002–2004 (no. 2002062925–003).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Forino, M., Graziotto, R., Tosetto, E. et al. Identification of a novel splice site mutation of CLCN5 gene and characterization of a new alternative 5’ UTR end of ClC-5 mRNA in human renal tissue and leukocytes. J Hum Genet 49, 53–60 (2004). https://doi.org/10.1007/s10038-003-0108-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-003-0108-1
Keywords
This article is cited by
-
Genetics and phenotypic heterogeneity of Dent disease: the dark side of the moon
Human Genetics (2021)
-
Functional analysis of suspected splicing variants in CLCN5 gene in Dent disease 1
Clinical and Experimental Nephrology (2020)
-
Complexity of the 5′UTR region of the CLCN5gene: eleven 5′UTR ends are differentially expressed in the human kidney
BMC Medical Genomics (2014)
-
Association between OPG, RANK and RANKL gene polymorphisms and susceptibility to acute coronary syndrome in Korean population
Journal of Genetics (2012)
-
A missense mutation in the chloride/proton ClC-5 antiporter gene results in increased expression of an alternative mRNA form that lacks exons 10 and 11. Identification of seven new CLCN5 mutations in patients with Dent’s disease
Journal of Human Genetics (2007)