Abstract
We have identified a novel gene, designated KRAP (Ki-ras-induced actin-interacting protein), encoding a protein of 1,259 amino acids with coiled-coil regions and transmembrane regions, from the cDNA library of human colon cancer HCT116 cells, as one of the genes upregulated by activated Ki-ras. While KRAP was rarely expressed in normal colon epithelium, deregulated constitutive KRAP expression was observed in some other colon cancer cells. In normal tissues, KRAP was strongly expressed in pancreas and testis. Anti-KRAP polyclonal antibodies detected endogenous KRAP as the molecular size of Mr 180,000, and immunofluorescence microscopy and cytochalasin E treatment revealed that KRAP was clearly associated with the actin filaments. Furthermore, KRAP was localized as a membrane-bound form with extracellular regions. These results together suggested KRAP might be involved in the regulation of filamentous actin and signals from the outside of the cells.
Similar content being viewed by others
Introduction
Ras has been implicated in controlling cell proliferation, differentiation, and apoptosis (Barbacid 1987; Bos 1989; Almoguera et al.1998; Bos et al. 1987; Forrester et al. 1987). We previously established HCT116-derived cells in which activated Ki-ras was disrupted through gene targeting (Shirasawa et al. 1993) and had reported the several critical molecular mechanisms of the activated Ki-ras-mediated signals in tumorigenesis utilizing these cells (Shirasawa et al. 1993; Allgayer et al. 1999; Okada et al. 1998; Rak et al. 1995; Baba et al. 2000; Tsunoda et al. 2002; Okumura et al. 1999). The cleavage signal-1 protein, CS-1, a protein of 249 amino acids reported to be detected as the molecular size of Mr 33,000 in Western blot using anti-CS-1 polyclonal antibodies, was originally identified as a sperm surface antigen involved in regulation of early cleavage of fertilized eggs (Javed and Naz 1992; Naz 1992). Here we have identified a novel gene, designated KRAP (Ki-ras-induced actin-interacting protein), including CS-1 sequences in the COOH terminal region. KRAP was overexpressed through the activated Ki-ras-mediated signals in HCT116 cells, and anti-KRAP polyclonal antibodies detected endogenous KRAP as a single band of the molecular size of Mr 180,000. Of great interest is that KRAP was colocalized with filamentous actin and was also localized at the cell surface.
Materials and methods
Isolation of differentially expressed genes
Isolation of differentially expressed genes between HCT116 and HKe3 cells was as described (Baba et al. 2000). KRAP cDNA was isolated by conventional plaque hybridization from the HCT116 cDNA library, and the full length of KRAP cDNA was determined by 5’ rapid amplification of cDNA ends. Mouse KRAP cDNA was isolated from a mouse heart cDNA library using a human KRAP cDNA as a probe, and the full length of cDNA was determined as described above. DNA sequences were determined by ABI Prism Big Dye Terminator Cycle Sequencing (Applied Biosystems, Foster, CA. USA) with ABI 3100 sequencer.
Northern blot
Total RNA of cell lines was isolated by ISOGEN (Nippon Gene, Tokyo, Japan) following the manufacturer’s protocol. Total RNA (20 μg) was separated by electrophoresis in a 1% agarose-formaldehyde gel, transferred to nylon membrane (Pall, East Hills, NY, USA), as described (Baba et al. 2000; Tsunoda et al. 2002). Human endocrine multiple tissue Northern (MTN) blot filter and human MTN blot II filter were purchased from Clontech (Palo Alto, CA, USA) on which 2 μg of polyA RNA from each tissue was loaded. These filters were hybridized with probes for human KRAP cDNA (full length of the coding region), and β-actin cDNAs, as described (Okumura et al. 1999).
Anti-KRAP polyclonal antibodies
Anti-KRAP polyclonal antibodies (FT017–2) were raised in rabbits using the polypeptide (amino acids 1,178–1,192 of human KRAP: VDAAEGAPEVVGPKS). Polyclonal antibodies were purified from serum by using protein-G Sepharose column.
Transfections and stable transfectants
COOH terminal hemagglutinin (HA)-tagged (Baba et al. 2000) KRAP cDNA was subcloned into pSI vector (Promega, Madison, WI, USA). Cells were transiently transfected using Polyfect (QIAGEN, Hilden, Germany) according to the manufacturer’s protocol. Total cell lysates were extracted 24 h after transfection then subjected to immunoblot analysis using Western blots (Okumura et al. 1999). For stable transfectants, COOH terminal HA-tagged KRAP cDNA was subcloned into pMG vector (Invivogen, San Diego, CA, USA), and the linearized pMG-KRAP-HA was transfected into NIH3T3 cells using Polyfect. Cells maintained in Dulbecco’s modified eagle’s medium (DMEM) with 10% fetal calf serum (FCS) were split after 2 days, and cells were selected with 100 μg/ml of hygromycin B (Invitrogen, Carlbad, CA, USA). Colonies surviving in selection media in 14 days were picked up under the microscope and expanded and characterized for the expression of KRAP-HA by Western blot using anti-HA antibody (3F10) (Roche, Mannheim, Germany).
Immunofluorescence staining
Cells were grown on collagen-coated glass coverslips in growth medium. Twenty-four hours after transfection, cells were fixed with 3.7% paraformaldehyde in phosphate buffered saline (PBS) for 10 min at room temperature. After blocking with 1.4% milk in PBS containing 0.1% saponin, cells were incubated with anti-HA antibody (3F10) in PBS containing 0.1% bovine serum albumin (BSA) and 0.1% saponin for 1 h at room temperature, followed by incubation with Alexa Fluor 488 anti-rat IgG and/or Alexa Fluor 594 phalloidin (Molecular Probes, Eugene, OR, USA) at a dilution 1:2,000 for 1 h at room temperature. For counterstaining the nucleic acids, cells were incubated with 100 μg/mL of DNase-free RNase in 2×SSC (0.3 M NaCl, 0.03 M sodium citrate, pH7.0) followed by incubation with 500 nM solution of propidium iodide (PI) (Molecular Probes, Eugene, OR, USA) for 5 min at 37°C. Cells were covered with a drop of GEL/MOUNT (Biomeda, Foster City, CA, USA), viewed by confocal laser scanning microscopy (FLUOVIEW FV500, OLYMPUS). Pretreatment of cytochalasin E (Sigma, Deisenhofen, Germany) (100 ng/ml or 1.0 μg/ml in DMEM with 10% FCS) was done 3 h before fixing.
Surface biotinylation and immunoprecipitation
Biotinylation of cell-surface proteins was done as described (Baba et al. 2000). The surface-biotinylated lysates of the transfected HCT116 cells (4×107) were immunoprecipitated with 2 μg/ml of anti-HA antibody for overnight at 4°C, then protein G-Sepharose was added for 2 h. Protein G-Sepharose complexes were boiled for 10 min, and the supernatants were run in a 6% SDS-PAGE. Western blots were done using streptavidin-horseradish peroxidase (SA-HRP) (Amersham Biosciences, Piscataway, NJ, USA) and anti-HA antibody (3F10).
Results
To identify the genes located downstream of the activated Ki-ras-mediating signals, a PCR-based cDNA subtraction library was constructed between HCT116 cells and HCT116-derived activated Ki-ras-disrupted clone cells (HKe3) through homologous recombination (Shirasawa et al. 1993). The KRAP cDNA fragment was detected from this subtracted library, and the full length of KRAP cDNA was isolated from the HCT116 cDNA library (Baba et al. 2000). To confirm the differential expression of KRAP between HCT116 cells and HKe3 cells, Northern blot analysis was done under exponential condition. Northern blot detected the 5.2 kb-major KRAP transcript and the other minor 6.0 kb-KRAP transcript (Fig. 1). KRAP was strongly expressed in HCT116, whereas it was rarely observed in HKe3 (Fig. 1A). Furthermore, HKe3-stable transfectants expressing the activated Ki-ras (e3-MKRas#9, e3-MKRas#14) (Baba et al. 2000) evidently expressed KRAP (Fig. 1A). Northern blots revealed that KRAP expression was rarely evident in the normal colon epithelium, whereas it was strongly expressed in several kinds of human colon cancer cells (Fig. 1B). On the other hand, KRAP expression in human normal tissues was strongly observed in pancreas and testis (Fig. 1C).
Deregulated Ki-ras-induced actin-interacting protein (KRAP) mRNA expression in human cancer cell lines and normal tissues. A Twenty micrograms of total RNA of HCT116, HKe3, and HKe3-derived transfectants expressing activated Ki-ras (e3-MKRas#9 and e3-MKRas#14) were run in a 1% agarose-formaldehyde gel, then Northern blots were done using the coding region of KRAP (upper panel) and β-actin (lower panel) as probes. B Twenty micrograms of total RNA of each cancer cell line were run in a 1% agarose-formaldehyde gel, and Northern blots were done using KRAP (upper panel) and β-actin (lower panel) as probes. C Human endocrine system multiple tissue Northern (MTN) blot filter (Clontech) (left panel) and human MTN blot II (Clontech) (right panel), on which 2 μg of polyA RNA from each tissue was loaded, hybridized with KRAP as probes
Human KRAP cDNA was isolated from the HCT116 cDNA library and the full length of KRAP cDNA was determined by 5’ rapid amplification of cDNA ends (AB116937). KRAP encodes a hypothetical protein of 1,259 amino acids (a.a.) with the transmembrane (a.a.870–890) and coiled-coil regions (a.a 992–1,038) (Fig. 2). To identify a mouse homologue of human KRAP, mouse cDNA library was screened using human KRAP cDNA fragment as a probe, and mouse KRAP cDNA encoding a protein of 1,252 a.a. was cloned (AB120565). Mouse KRAP cDNA had 81% identity with human KRAP at the nucleotides level and 86% similarity at the amino acid level (Fig. 2).
Anti-KRAP polyclonal antibodies (FT017–2) raised in rabbits using the peptides of the COOH terminal region of KRAP (a.a.1,178–1,192), recognized a single band of the molecular size of about Mr 180,000 in HCT116 and e3-MKRas#9, but barely in HKe3 (Fig. 3). The 293T cells were transiently transfected with KRAP-HA, HA-tagged in the COOH terminal region of KRAP, and KRAP-HA was evidently detected by FT017–2 at almost the same molecular size of KRAP detected in HCT116 cells, but was not detected in the nontransfected 293T cells. Furthermore, any processed KRAP was not detected in other human cancer cell lines examined by FT017–2 (data not shown).
Activated Ki-ras-induced actin-interacting protein (KRAP) expression. Anti-KRAP rabbit polyclonal antibodies detected the endogenous KRAP at the molecular size of Mr 180,000 (arrow). HCT116, HKe3, and HKe3-MKRas#9 cells were grown at exponential growth conditions; 293T cells were transfected with pSI-KRAP-hemagglutinin (HA) or empty vector, and then whole cell lysates were run in a 6% SDS-PAGE
To elucidate the localization of KRAP in the cells, NIH3T3-derived stable transfectants expressing KRAP-HA (NIH3T3-KRAP-HA#1) were stained with anti-HA antibody. KRAP-HA seemed to be colocalized with phalloidin-specific dye for F-actin in NIH3T3-KRAP-HA#1 (Fig. 4A), indicating that KRAP is colocalized with actin filaments. Cytochalasin E, an epoxide-containing metabolite of Aspergillus clavatus, is a compound with diverse activities on cellular function, including inhibition of actin polymerization and glucose transport (Udagawa et al. 2000). To elucidate the interaction of KRAP with the actin filaments, treatment with cytochalasin E was done. Phalloidin staining indicated that disruption of actin filaments by cytochalasin E occurred in a dose-dependent manner, and the interaction of KRAP with phalloidin gradually disappeared (Fig. 4B, C). These results together suggested that KRAP is associated with actin filaments in NIH3T3 cells.
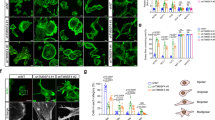
Colocalization of Ki-ras-induced actin-interacting protein (KRAP) with actin filaments. A A NIH3T3-derived stable transfectant expressing human KRAP hemagglutinin (HA) (NIH3T3-KRAP-HA#1) was established. Cells were fixed and stained with anti-HA antibody (green) and phalloidin (red). Superimposed images are also shown (merge). NIH3T3-KRAP-HA#1 cells were pretreated with 100 ng/ml (B) and 1 μg/ml (C) of cytochalasin E in growth medium for 3 h. Cells were washed three times with PBS and stained with anti-HA antibody (green) and phalloidin (red). Superimposed images are also shown (merge). D COS-7 cells were transfected with pSI-KRAP-HA, and cells were grown on collagen-coated glass coverslips in growth medium. 24 h after transfection, cells were fixed and stained with anti-HA (green) and propidium iodide (PI) (red). Superimposed images are also shown (merge). COS7 cells (E) and HCT116 cells (F) were transfected with pSI-KRAP-HA and were stained with anti-HA antibody (3F10). COS7 cells were also stained with PI
To determine the localization of KRAP in cancer cells, COS7 cells and HCT116 cells were transfected with KRAP-HA and were stained with anti-HA and PI, indicating that KRAP is not localized in the nucleus (Fig. 4D). In COS7 cells, KRAP seemed to be localized near the cell membrane and the leading edge of the cells (Fig. 4E). In HCT116 cells, KRAP seemed to be also localized near the membrane regions (Fig. 4F). To elucidate whether KRAP exists as cell-surface protein, surface biotinylation was done, indicating that KRAP has the outer membrane region in HCT116 cells (Fig. 5).
Ki-ras-induced actin-interacting protein (KRAP) as cell-surface protein. HCT116 cells were surface-biotinylated and then immunoprecipitated with anti-hemagglutinin (HA) antibody. The immunoprecipitated products were detected by anti-HA antibody (left) and streptavidin-horseradish peroxidase (SA-HRP) (right). Arrow indicates biotinylated KRAP
Discussion
We have identified a novel gene, designated KRAP (Ki-ras-induced actin-interacting protein), encoding a protein of 1,259 amino acids with coiled-coil and transmembrane regions, from the cDNA library of human colon cancer HCT116 cells, as one of the genes upregulated by activated Ki-ras.
Although Ki-ras mutation was detected in colon cancer cell lines, including HCT116, DLD-1, HCT-15, LS180, LoVo, and SW620 (Shirasawa 1993), KRAP was not overexpressed in LoVo cells, suggesting that other signals, in addition to the activated Ki-ras-mediated signals, might be necessary for the overexpression of KRAP. On the other hand, the colon cancer cells without Ki-ras mutation, including WiDr and Colo201 (Shirasawa 1993), expressed evident expression of KRAP, suggesting that other signals also could induce deregulated KRAP expression in human colon cancer cells. NIH3T3-derived stable transfectants expressing KRAP did not show tumorigenicity in nude mice nor did they show soft agar colony formation (data not shown), suggesting that overexpression of KRAP itself might not be sufficient for tumorigenicity and that other additional signals might be necessary for tumorigenicity. However, deregulated expression was observed not only in colon cancer cells but also in other kinds of cancer cell lines (data not shown), suggesting that KRAP might play critical roles in tumorigenesis through collaboration with other signals.
Mouse KRAP cDNA had 81% identity with human KRAP at the nucleotides level and 86% similarity at the amino acid level, indicating that KRAP was highly conserved between human and mouse. COOH terminal regions (residues of a.a.1013–1259) of KRAP had been reported as the CS-1 (Javed and Naz 1992). CS-1 was originally identified as a sperm surface antigen of the molecular size of Mr 33,000, and anti-CS-1 polyclonal antibodies inhibited the early cleavage of fertilized eggs without affecting binding between sperm and egg or pronuclear formation (Javed and Naz 1992; Naz 1992).
We prepared the anti-KRAP polyclonal antibodies (FT017–2) raised in rabbits using the peptides of the COOH terminal region of KRAP, which recognized a single band of the molecular size of about Mr 180,000 in HCT116 cells, and processed KRAP was not detected in other human cancer cell lines examined. Although there is a possibility that CS-1 is located in the sperm surface as the molecular size of Mr 33,000, all these results together suggested that KRAP is an actual protein as the molecular size of about Mr 180,000.
XCS-1, Xenopus homologue of CS-1, was reported to be detected on the membrane and in the nucleus of Xenopus blastomeres, on the mitotic spindle in mitotic cells, and on the centrosomes in interphase cells (Nakamura et al. 2000). Immunofluorescence microscopy revealed that neither KRAP nor CS-1-HA, HA-tagged in the COOH terminal region of the original CS-1 (Javed and Naz 1992), was localized in the nucleus in HCT116 cells, COS-7 cells, or NIH3T3 cells (data not shown). On the other hand, immunofluorescence microscopy and cytochalasin E treatment revealed that KRAP was clearly associated with the actin filaments in NIH3T3 cells. Furthermore, immunofluorescence microscopy and surface biotinylation revealed that KRAP exists as a membrane-bound form with the extracellular region in HCT116 cells. All these results together suggest that the localization of KRAP in the cells might vary depending on the cell types or other signals involved in the regulation of KRAP.
One of the critical processes in cancer developments is migration that requires the continuous, coordinated formation and disassembly of adhesions. These processes are complex and require a coordinated interaction of actin (Pollard 2003) or actin-interacting proteins, signaling molecules, structural proteins, integrins, adaptor molecules, and microtubules (Webb et al. 2002). Colocalization of KRAP with actin filaments and localization at the cell membranes with extracellular region suggest that KRAP might be involved in regulation of actin filaments and signals from the outside of the cells.
We have demonstrated previously that activated Ki-ras is involved in the upregulation of c-myc, VEGF, the urokinase-type plasminogen activator receptor, epiregulin, and mig-6, all of which are critically involved in tumorigenesis. Here we have identified a novel actin-interacting protein, KRAP, evidently induced by activated Ki-ras-mediated signals in human colon cancer HCT116 cells. Further elucidation of KRAP function in regulation of cytoskeleton and migration and KRAP-mediated signaling pathways should shed light on the molecular mechanisms of tumorigenesis.
References
Allgayer H, Wang H, Shirasawa S, Sasazuki T, Boyd D (1999) Targeted disruption in an invasive colon cancer cell line down-regulates urokinase receptor expression and plasminogen-dependent proteolysis. Brit J Cancer 80:1884–1891
Almoguera C, Shibata D, Forrester K, Martin J, Arnheim N, Perucho M (1998) Most human carcinomas of the exocrine pancreas contain mutant c-Ki-ras genes. Cell 53:549–554
Baba I, Shirasawa S, Iwamoto R, Okumura K, Tsunoda T, Nishioka M, Fukuyama K, Yamamoto K, Mekada E, Sasazuki T (2000) Involvement of deregulated epiregulin expression in tumorigenesis in vivo through activated Ki-Ras signaling pathway in human colon cancer cells. Cancer Res 60:6886-6889
Barbacid M (1987) ras genes. Annu Rev Biochem 56:779–827
Bos JL (1989) ras oncogene in human cancer: a review. Cancer Res 49:4682–4689
Bos JL, Fearon ER, Hamilton SR, Verlaan-de Vries M, van Boom J, van der Eb AJ, Vogelstein B (1987) Prevalence of ras gene mutations in human colorectal cancers. Nature 327:293–297
Campbell SL, Khosravi-Far R, Rossman KL, Clark GJ, Der CJ (1998) Increasing complexity of Ras signaling. Oncogene 17:1395–1413
Forrester K, Almoguera C, Han K, Grizzle WE, Perucho M (1987) Detection of high incidence of Ki-ras oncogene during human colon tumorigenesis. Nature 327:298–303
Javed AA, Naz RK (1992) Human cleavage signal-1 protein; cDNA cloning transcription and immunological analysis. Gene 112:205–211
Johnson L, Greenbaum D, Cichowski K, Mercer K, Murphy E, Schmitt E, Bronson RT, Umanoff H, Edelmann W, Kucherlapati R, Jacks T (1997) K-ras is an essential gene in the mouse with partial functional overlap with N-ras. Genes Dev 11:2468–2481
Nakamura H, Wu C, Kuang J, Larabell C, Etkin LD (2000) XCS-1, a maternally expressed gene product involved in regulating mitosis in Xenopus. J Cell Sci 113:2497–2505
Naz RK (1992) Effects of antisperm antibodies on early cleavage of fertilized ova. Biol Reprod 46:130–139
Okada F, Rak JW, Croix BS, Lieubeau B, Kaya M, Roncari L, Shirasawa S, Sasazuki T, Kerbel RS (1998) Impact of oncogenes in tumor angiogenesis:mutant K-ras up-regulated of vascular endothelial growth factor/ vascular permeability factor is necessary, but not sufficient for tumorigenicity of human colorectal carcinoma cells. Proc Natl Acad Sci USA 95:3609–3614
Okumura K, Shirasawa S, Nishioka M, Sasazuki T (1999) Activated Ki-Ras suppresses 12-O-tetradecanoylphorbol-13-acetate-induced activation of the c-Jun NH2-terminal kinase pathway in human colon cancer cells. Cancer Res 59:2445-2450
Pollard TD (2003) The cytoskeleton, cellular motility and the reductionist agenda. Nature 422:741–745
Rak J, Mitsuhashi Y, Bayko L, Filmus J, Shirasawa S, Sasazuki T, Kerbel RS (1995) Mutant ras oncogene upregulate VEGF/VPF expression: implications for induction and inhibition of tumor angiogenesis. Cancer Res 55:4575–4580
Shirasawa S (1993) Analysis of molecular mechanism in colorectal tumorigenesis. Fukuoka Igaku Zasshi 84:25–35
Shirasawa S, Furuse M, Yokoyama N, Sasazuki T (1993) Altered growth of human colon cancer cell lines disrupted at activated Ki-ras. Science 260:85–88
Tsunoda T, Inokuchi J, Baba I, Okumura K, Naito S, Sasazuki T, Shirasawa S (2002) A novel mechanism of nuclear factor κB activation through the binding between inhibitor of nuclear factor- κBα and the processed NH2-terminal region of mig-6. Cancer Res 62:5668–5671
Udagawa T, Yuan J, Panigrahy D, Chang Y, Shah J, D’Amato RJ (2000) Cytochalasin E, an epoxide containing Aspergillus-derived fungal metabolite, inhibits angiogenesis and tumor growth. J Pharmacol Exp Ther 294:421–427
Webb DJ, Parsons JT, Horwitz AF (2002) Adhesion assembly, disassembly and turnover in migrating cells—over and over and over again. Nat Cell Biol 4:E97-E100
Zuber J, Tchernitsa OI, Hinzmann B, Schmitz A-C, Grips M, Hellriegel M, Sers C, Rosenthal A, Schafer R (2000) A genome-wide survey of RAS transformation targets. Nat Genet 24:144–152
Acknowledgments
This work was supported in part by a Grant-in-Aid for Scientific Priority Areas from the Ministry of Education, Science, Technology, Sports and Culture, Japan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Inokuchi, J., Komiya, M., Baba, I. et al. Deregulated expression of KRAP, a novel gene encoding actin-interacting protein, in human colon cancer cells. J Hum Genet 49, 46–52 (2004). https://doi.org/10.1007/s10038-003-0106-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-003-0106-3
Keywords
This article is cited by
-
KRAP tethers IP3 receptors to actin and licenses them to evoke cytosolic Ca2+ signals
Nature Communications (2021)
-
Functional non-coding polymorphism in an EPHA2 promoter PAX2 binding site modifies expression and alters the MAPK and AKT pathways
Scientific Reports (2017)
-
Analysis of KRAP expression and localization, and genes regulated by KRAP in a human colon cancer cell line
Journal of Human Genetics (2007)