Abstract
Extended-spectrum β-lactamase (ESBL)-producing bacteria pose a big challenge in clinical practices, warranting a new therapeutic strategy. In this study, methanol extract of the marine cyanobacterium Oscillatoria acuminata NTAPC05 was fractionated under bioassay guidance and the fractions were tested against three well-characterized ESBL-producing bacteria Escherichia coli U655, Stenotrophomonas maltophilia B929 and Enterobacter asburiae B938. Out of the four HPLC fractions, fraction 2 showed bactericidal activity against all the three ESBL producers much more efficiently (MIC 100 μg ml−1) than the fourth-generation cephalosporin (MIC >125 μg ml−1). The active fraction was subjected to time-kill test at concentrations of 1/2 × MIC, 1 × MIC and 2 × MIC, and the results substantiated the bactericidal property of the fraction against the ESBL producers. Spectral analysis revealed monogalactosyldiacylglycerol containing a palmitoyl (MGDG-palmitoyl), being reported for the first time, as the active fraction, and its bactericidal property against ESBL producers was determined. The active fraction appears to damage the bacterial membrane leading to lysis of the cell, as revealed in confocal laser scanning microscopy (CLSM) analysis, that was confirmed in scanning electron microscopic analysis. Cytotoxicity assay revealed the O. acuminata compound to be safe to a normal cell line HEK293 (human embryonic kidney cell). The in silico analysis of MGDG-palmitoyl revealed two successive H-bonding interactions with Leu198 of TEM1 β-lactamase. Taken together, the MGDG-palmitoyl from O. acuminata NTAPC05 offers potential to develop analogs as a therapeutic for bacteremia caused by ESBL producers.
Similar content being viewed by others
Introduction
Extended-spectrum β-lactamases (ESBLs) are the major basis of resistance of Gram-negative bacteria to broad-spectrum β-lactam antibiotics.1 ESBL-producing organisms are a serious threat to public health and are associated with high rates of morbidity and mortality.2 The third-generation cephalosporins (for example, cefotaxime, ceftazidime and ceftriaxone) were introduced for clinical use during 1980s and are continued to be widely practiced for treatment of infections caused by members of the family Enterobacteriaceae.3 The long-term use of antibiotics has brought about antibiotic selection pressure in bacteria such that the bacteria circumvent the antibiotic treatment through point mutations in the plasmid-mediated β-lactamase temoniera (TEM), sulfhydryl variable (SHV) and cefotaxime (CTX-M) ESBL genes. More than 300 different ESBL variants have been described.4 Though TEM and SHV variants are the most common ESBLs, the strains expressing CTX-M ESBLs are emerging as a serious problem in the clinical scenario.5 ESBLs are reported in many different bacterial species such as Enterobacter aerogenes, Enterobacter cloacae, Proteus mirabilis and Serratia marcescens.6 A vast majority of the ESBL-producing bacteria develop resistance to third-generation antibiotics such as cefotaxime, ceftriaxone, ceftazidime and oxyimino-monobactams over a period of time. However, ESBLs are inhibited by clavulanic acid, sulbactam and tazobactam.4 There are only a few reports with regard to clinical efficacy of therapeutics in treatment of infections caused by ESBL-producing bacteria.7 The ESBL-producing bacteria are multiple drug resistant8 and, thus, pose a serious challenge to treatment of ESBL-related bacteremia.
Microalgae, distributed in ecosystems as varied as fresh water to extreme saline environments, are capable of synthesizing a variety of chemical compounds.9 Several of these compounds are antibiotics with promising antibacterial, antifungal, antiprotozoal antiplasmodial and anticancer activities.10 A wide range of in vitro antifungal activities of extracts of green algae, diatoms and dinoflagellates have also been reported.11, 12 Microalgae are a promising group of organisms for drug discovery research as their metabolites are active against bacteria, fungi and viruses.13, 14 In the present study, we evaluated the HPLC fractions from methanol extract of the marine cyanobacterium Oscillatoria acuminata NTAPC05 for efficacy against three well-characterized ESBL producers, and the most active fraction with potential for therapeutic application was identified.
Materials and methods
Bacterial strains and resistance pattern
Three strains of bacteria that infect urinary tract were isolated and identified at Medwin Hospital (Hyderabad, India). The resistance patterns of these bacterial strains were determined using the antibiotics ceftazidime (30 μg), cepfodoxime (10 μg), amoxicillin (30 μg), novobiocin (30 μg), rifampicin (5 μg), erythromycin (15 μg), amikacin (30 μg), methicillin (5 μg), vancomycin (30 μg) and penicillin (10 units), adopting disk diffusion test (Himedia Laboratories, Mumbai, India) (Table 1). The clinical strains used were KC759521 Escherichia coli U655, KC751004 Enterobacter asburiae B938 and KC759524 Stenotrophomonas maltophilia B929, bacteria that produce ESBL at high levels and infect the urinary tract.
Detection of ESBLs (HEXA G-minus 24 and E-test triple detection strip)
Phenotypic characterization of the clinical isolates was performed by combination disc (HEXA disc, Himedia Laboratories) diffusion test and the results were interpreted according to the Clinical and Laboratory Standards Institute guidelines. Ceftazidime (30 μg), cefotaxime (30 μg) and cepfodoxime (10 μg), alone and in combination with clavulanic acid (10 μg), were used as the indicators.15 Furthermore, the ESBL production of the three isolates was compared using E-test triple detection strip calibrated with MIC reading scales in μg ml−1. ESBL production, if any, was recorded using β-lactamase inhibitor-containing discs with enhanced zone of inhibition (Himedia Laboratories), in which the zone of inhibition for cephalosporin/clavulanic acid was >5 mm (combination method) and >8 mm (E-test) rather than cephalosporin alone.
Isolation and identification of cyanobacteria
The sample of marine cyanobacteria was collected from Mandapam, Ramanathapuram District (Latitude 9°16’51.72”N Longitude 79°10’40.55”E), Tamil Nadu, India. The sample was isolated by plating, purified, cultured in MN medium and maintained in the institute’s microalgal repository. The cyanobacterium of interest, selected based on our preliminary screening of several microalgae, was identified from cell shape and size using standard monograph.16 The species-level identification of O. acuminata NTAPC05 that produced anti-ESBL bacterial substance was confirmed by 16S rDNA sequencing analysis. Briefly, genomic DNA was extracted as previously described.17 Universal cyanobacteria 16S rDNA primer was used for amplification of DNA. The 16S rDNA sequence was obtained and compared with other cyanobacterial sequences using NCBI-BLAST with a sequence query for their pairwise identities. The evolutionary distances were computed using maximum composite likelihood method18 and are expressed in units of number of base substitution per site that was computed using MEGA 5.0 softwaren (http://www.megasoftware.net/).
Bioassay-guided fractionation of O. acuminata extract
Filaments of the cyanobacterium were harvested and homogenized in methanol using a mortar and pestle. The homogenate was centrifuged at 10 000 r.p.m. for 10 min at 4 °C and the supernatant was separated. This process was repeated until the pellet turned gray or the supernatant turned colorless. The supernatant was pooled and filtered through ordinary filter paper, followed by Whatman No. 1 filter paper (Himedia Laboratories) and then concentrated using a rotary vacuum evaporator. The methanol extract was subjected to TLC separation. The chromatogram was developed using different solvent systems including different ratios of hexane in ethyl acetate and methanol in chloroform. The extract was also fractionated in HPLC system (Perkin-Elmer, Shelton, CT, USA) consisting of a Perkin Elmer Series 2000 pump, a Gilson FC203B fraction collector (Middleton, WI, USA) and a Perkin Elmer Series 200 UV/VIS detector set at 238 nm. The samples were separated using a reverse phase analytical column (Chromolith performance RP-18 endcapped, 4.6 mm × 100 mm, pore size 2 μm to 13 nm, monolithic; Merck, Darmstadt, Germany) equipped with a guard column (Chromolith guard 5–4.6 mm cartridge, RP-18; Merck). The mobile phase consisted of a mixture of acetonitrile and water operated on a gradient basis starting from a ratio of 9:1 to 100% acetonitrile within 20 min at a flow rate of 2 ml min−1. This separation yielded four fractions.
Assay for antibacterial activity
The antibacterial activities of the O. acuminata NTAPC05 methanol extract fractions from bioassay-guided fractionation were determined by agar diffusion method against the growth of ESBL-producing E. coli U655, E. asburiae B938 and S. maltophilia B929, with the fourth-generation cephalosporin cefepime as the reference drug,19 and the inhibition zones around the spotted fractions were measured. Inhibition zones ⩾8 mm were considered indicative of inhibitory activity.
Determination of MIC and MBC
The determination of MIC was performed in 96-well microplates by the microdilution method in Mueller–Hinton broth (Himedia) medium, according to the Clinical and Laboratory Standards Institute (CLSI).20 ESBL-producing E. coli U655, E. asburiae B938 and S. maltophilia B929 (104 CFUs ml−1) were inoculated in the broth with cefepime (50–250 μg ml−1) and the active methanol fractions (50–500 μg ml−1) and incubated at 37 °C for 18 h. The MBC was determined after the MIC assays. In wells in which MIC results revealed no bacterial growth, an aliquot of 0.01 ml was seeded in Mueller–Hinton agar without addition of drugs, and bacterial growth was evaluated for MBC determination. After 18 h at 37 °C, if the MIC=MBC or the MBC was one, two or three dilutions above the MIC, the activity was considered bactericidal.21 This approach helped to narrow down to the most active fraction.
Time-kill assay
The rate of killing of bacteria by the O. acuminata active fraction was assayed using a modified plating technique of Eliopoulos and Moellering.22 The active fraction was incorporated into 10 ml Mueller–Hinton broth at 1/2 × MIC, 1 × MIC and 2 × MIC. Two controls (1) Mueller–Hinton broth without the compound but inoculated with the test organisms and (2) Mueller–Hinton broth with the compound at the test concentrations but the test organisms, not included, were maintained. Inoculum density of ∼105 CFUs ml−1 verified from total viable count was used to inoculate 10 ml volumes of both test and control tubes. The tubes were incubated at 37 °C on an orbital shaker at 120 r.p.m. Then, 100 μl aliquots at 0, 3, 6, 9 and 12 h were removed from the culture medium for determination of CFUs ml−1, adopting plate count technique by plating out 25 μl of each dilution. After incubation for 24 h, the emergent bacterial colonies were counted, CFUs ml−1 calculated and compared with the count of the control culture without the compound.
Characterization of active fraction using analytical HPLC
The most active fraction (that is, F2) of O. acuminata NTAPC05 was further analyzed in a Perkin Elmer series HPLC system consisting of an analytical column within 20 min at a flow rate of 2 ml min−1.
FTIR analysis of active fraction
The IR spectrum of the most active fraction was recorded on Perkin Elmer Spectrum (Germany). The spectra were scanned in the 400–4000 cm−1 range. The spectra were plotted as intensity versus wave number.
NMR studies of active fraction
The most active fraction of O. acuminata was subjected to 1H and 13C NMR analysis. NMR spectrum was recorded on a Bruker Avance III spectrometer (Bruker GmbH, Karlsruhe, Germany) with a 5 mm probe. The anti-ESBL E. coli fraction was dissolved in deuterated chloroform, CDCl3, at a concentration of 20 mg ml−1. For 1H NMR the frequency was 400 MHz, and for 13C NMR the frequency was 100 MHz.
Confocal laser scanning microscopy
The active fraction (final concentration 1/2, 1 and 2 × MIC values) from O. acuminata NTAPC05 was added to ESBL E. coli, S. maltophilia and E. asburiae cells at the early log growth phase in MHA broth medium, and the control group contained bacteria only. The cultures were then maintained at 37 °C for 12 h and the optical density was determined at 600 nm. To analyze a possible mode of action of the active fraction on ESBL-producing bacterial cells, ∼10 μl of each sample was collected for acridine orange–ethidium bromide staining and subsequent observation by confocal laser scanning microscopy (CLSM). The CLSM images were obtained using a Carl Zeiss CLSM 710 (Carl Zeiss, Jena, Germany) equipped with a 100 × oil-immersion objective lens.
Scanning electron microscopic analysis of bacteria
After the exposure, bacteria were fixed with 2% glutaraldehyde for 24 h. The samples were postfixed with 1% osmic acid for 2 h, and then dehydrated in an acetone gradient (35, 50, 70, 80, 95 and 100%). Finally, the air-dried samples were analyzed by scanning electron microscopy (model: VEGA3 TESCAN, Brno, Czech Republic).
Cytotoxicity bioassays
A microassay for cytotoxicity in HEK293 (human embryonic kidney) normal cell line was performed using the MTT method.23 The adherent noncancerous cells were seeded in 96-well microplates at a concentration of 0.3 × 106 cells ml−1 and incubated for 24 h for them to attach. The active fraction was added to the cell culture at concentrations of 1/2 × MIC, 1 × MIC and 2 × MIC, and incubated for 24 h. The MTT solution was added 3 h before the end of the incubation time. Cell survival was evaluated using a multiwell scanning spectrophotometer at 540 nm. The dose and time-dependence of cellular toxicity were derived from MTT results.
Iodometric assay
The modified method of Ahmad and Yadava24 was used for detecting the inhibition of β-lactamase in the ESBL E. coli. Exponentially growing culture of ESBL E. coli was inoculated on to starch agar plate and incubated overnight at 37 °C. Well-developed colonies of ESBL E. coli on starch agar plates were then isolated and flooded with freshly prepared phosphate-buffered saline containing potassium iodide (15 mg ml−1), iodine (3 mg ml−1) and MGDG-palmitoyl (100 μg ml−1). No growth was observed on the starch agar plate that indicated inhibition of β-lactamase production. A negative control with ampicillin (10 mg ml−1) supplemented to ESBL E. coli in starch agar plate was also run.
Molecular docking
Crystal structure of ESBL E. coli TEM1 β-lactamase was retrieved from Protein Data Bank (PDB ID: 1BTL) with the resolution of 1.8 Å. The bioactive compound from O. acuminata NTAPC05 was identified as MGDG-palmitoyl, and confirmed by FTIR and NMR spectroscopic methods. The compound was drawn using Chemdraw software, (CambridgeSoft Corporation, Waltham, MA, USA) and the 3D structure thus generated was converted into pdb files using Discovery Studio (version 2.5.5, Accelrys, San Diego, CA, USA). Furthermore, the ligand was energy-minimized and optimized using Swiss PDB viewer for molecular docking study. Docking of MGDG-palmitoyl to E. coli β-lactamase was carried out using Autodock 4.0.25 Receptor–ligand interaction was visualized using Pymol software (http://www.pymol.org).
Results
ESBL typing by combination disc and E-test strip
CLSI ESBL detection, in the combination disc method, showed that three isolates produced ‘positive’ result (Table 2). According to CLSI ESBL detection (triple detection strip), MIC criteria showed similar results (Table 2), although the confirmatory test was positive for the three ESBL-producing strains. Instead of cepfodoxime, fourth-generation cefepime was used in the strip test as marker for high-level ESBL production.
Identification of marine cyanobacterium
The trichome of O. acuminata was straight, 3.2 μm long and 6.8 μm wide, with curved ends and slightly constricted along the length. Molecular characterization studies were carried out adopting 16S rRNA gene sequence analysis. The amplified 16S rRNA gene sequence (650 bp) of O. acuminata was submitted to GenBank with accession no. KF469280. Blast results showed that KF469280 O. acuminata was highly similar (that is, 99%) to O. acuminata NTDM04 (accession no. GU812860). The evolutionary history was inferred using Tamura–Nei method. The tree with the highest log likelihood (−153.4067) is shown in Supplementary Figure S1. The percentage of trees in which the associated taxa clustered together is shown next to the branches. Initial tree(s) for the heuristic search were obtained automatically by applying neighbor-joining and BioNJ algorithms to a matrix of pairwise distances estimated using the maximum composite likelihood approach, and then selecting the topology with superior log likelihood value. The tree was drawn to scale, with branch length measured in the number of substitutions per site. The analysis involved 16 nucleotide sequences. All positions containing gaps and missing data were eliminated. There were totally 21 positions in the final data set.
Bioassay-guided fractionation and antibacterial activity
The crude extract of O. acuminata NTAPC05 was separated by TLC (Supplementary Figure S2). The spots 1 and 2 were separated using hexane in ethyl acetate (10:90 and 30:70 ratios), and spots 3 and 4 were separated using methanol in chloroform at the same ratios. The four fractions were separated by the preparative HPLC (Supplementary Figure S3). Among the four, fraction 2 showed inhibitory activity against ESBL E. coli U655, E. asburiae B938 and S. maltophilia B929, whereas the other three fractions exhibited only moderate to no inhibitory activity. Assay with cefepime, a fourth-generation cephalosporin, showed no inhibitory effect when compared with O. acuminata active fraction (Supplementary Figure S4). The MIC and MBC as determined for the separated fractions and cefepime against E. coli, E. asburiae and S. maltophilia are listed in Table 3. As the MBC and MIC values were the same, our results indicate bactericidal activity of the active fraction (that is, fraction 2).
Time-kill test
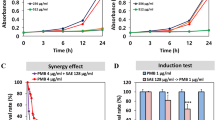
The results of the time-kill tests are presented in Figure 1. Data are presented as the log10 CFUs per ml change and are based on the conventional bactericidal activity standard, that is, 3 log10 CFUs per ml or greater reduction in the viable colony count. Average log reduction in viable cell count in time-kill assay ranged from 0.015 log10 to 3.180 log10 CFUs per ml after 12 h of incubation, and from 1.527 log10 to 3.230 log10 CFUs per ml after 12 h of incubation in 1 × MIC and 2 × MIC, respectively, of the O. acuminata NTAPC05 active fraction. The order of reduction in cell density on treatment with the extract was E. coli (−1.527 log10) >E. asburiae (−1.473 log10) and >Stenotrophomonas sp (−0.735 log10). The significant reduction in the bacterial populations at these concentrations suggests that the compound is efficiently bactericidal on 12 h of incubation. The efficacy of growth inhibition of the active fraction from O. acuminata NTAPC05 was found to be dose as well as time dependent, producing distinct time-kill profiles for the tested bacteria.
Analytical HPLC and FTIR analysis
The reverse phase HPLC analysis of the active fraction showed a single peak with the retention time 2.809, revealing no impurities in the active fraction (Supplementary Figure S5). The FTIR spectrum of the active fraction showed peaks in between 3325 and 1036 cm−1. The peaks of fingerprints were also observed within the range 500 to 400 cm−1, whereas no peak was observed in the range 300–260 cm−1, clearly indicating that the fraction did not contain protein or nucleic acid impurities. Functional peaks at 3325 and 2923 cm−1 revealed the existence of O–H and C–H stretch bands, respectively. Similarly, C–H stretch in CH3 was also observed at 2853 and 1404 cm−1. Presence of carboxyl group and C–O stretch in the active fraction was observed at 1629 and 1036 cm−1 region, respectively (Supplementary Figure S6).
NMR studies
The 1H and 13C NMR analysis of O. acuminata revealed peaks as listed in Table 4. The proton and carbon NMR spectra of active fraction are shown in Supplementary Figure S7a and b. Based on the results of FTIR and NMR analysis, the structure of the active fraction was determined as MGDG-palmitoyl (Figure 2).
The structure of monogalactosyldiacylglycerol (MGDG)-palmitoyl (3-(3,4,5-trihydroxy-6-(hydroxymethyl) tetrahydro-2H-pyran-2-yloxy) propane-1,2-diyl dipalmitate) from O. acuminata NTAPC05 that showed antibacterial activity against the three extended-spectrum β-lactamase (ESBL) producers. A full color version of this figure is available at the Journal of Antibiotics journal online.
Confocal microscopic study
CLSM was performed to investigate the possible mechanism of action of O. acuminata NTAPC05 active fraction on samples of ESBL E. coli, S. maltophilia and E. asburiae at early log phase growth. It was observed that the active fraction could effectively deal with the ESBL-producing bacterial cells at the concentrations of 1/2, 1 and 2 × MIC for 12 h in a dose-dependent manner (Figure 3). The overlay images clearly demonstrated that the active fraction initiated the detrimental effect on all three bacteria at 1/2 × MIC exposures (Figures 3b) and the bacteria were killed at 1 × MIC (Figures 3c). Exposure up to 2 × MIC resulted in disintegration of the bacterial cells (Figures 3d), whereas the control cells (Figures 3a) were intact up to 12 h of incubation.
Confocal laser-scanning microscopic (CLSM) images of extended-spectrum β-lactamase (ESBL)-producing bacteria treated with Oscillatoria acuminata fraction. (a, e, i) Control cells of ESBL Escherichia coli, Stenotrophomonas maltophilia and Enterobacter asburiae. (b, f, j) ESBL E. coli, S. maltophilia and E. asburiae cells treated with the active fraction at 1/2 × MIC (50 μg ml−1) concentration for 12 h. (c, g, k) ESBL E. coli, S. maltophilia and E. asburiae cells treated with the active fraction at 1 × MIC (100 μg ml−1) concentration for 12 h. (d, h, l) ESBL- E. coli, S. maltophilia and E. asburiae cells treated with the active fraction at 2 × MIC (200 μg ml−1) concentration for 12 h. Green color indicates live cells; yellow and red colors indicate dead cells (excitation: 48 nm; emission: 570–620 nm). A full color version of this figure is available at the Journal of Antibiotics journal online.
Cell morphology by SEM analysis
Figures 4b clearly indicate that exposure of the three bacterial strains to MGDG-palmitoyl at MIC brought about structural changes in the outer membrane of the cells, whereas the untreated cells did not suffer similar damage (Figures 4a. The loss of viability of bacteria correlated with disruption of cell membrane that is highly consistent with the damage of cell walls of all three strains of bacteria revealed in the SEM analysis.
Scanning electron microscopic (SEM) images of control and treated cells of extended-spectrum β-lactamase (ESBL) producers. (a, c, e) Control cells E. coli, S. maltophilia and E. asburiae without addition of monogalactosyldiacylglycerol (MGDG)-palmitoyl from O. acuminata NTAPC05. (b, d, f) E. coli, S. maltophilia and E. asburiae cells were treated with 1 × MIC of MGDG-palmitoyl from O. acuminata NTAPC05. A full color version of this figure is available at the Journal of Antibiotics journal online.
Dose-dependent cytotoxicity of MGDG-palmitoyl
HEK293 human embryonic kidney cells were exposed to MGDG-palmitoyl at 1/2 × MIC, 1 × MIC and 2 × MIC concentrations. Cell viability, observed as the function of dose, was not affected to any significant level, and ⩾90% cells were viable for 24 h (Figure 5). The compound was clearly nontoxic to the normal cell line HEK293.
Morphological observation of HEK293 cell by phase-contrast microscopy. Cells were exposed to different concentrations (1/2 × MIC, 1 × MIC and 2 × MIC) of the most active fraction from Oscillatoria acuminata NTAPC05 for 24 h. (a) Control. (b) Cells treated with 1/2 × MIC concentration of O. acuminata active fraction. (c) Cells treated with 1 × MIC concentration of O. acuminata active fraction. (d) Cells treated with 2 × MIC concentration of O. acuminata active fraction. The images clearly indicate that the HEK293 cells treated with the active fraction of O. acuminata NTAPC05 do not show morphological changes such as cell shrinkage, irregular shapes and nuclear condensation. A full color version of this figure is available at the Journal of Antibiotics journal online.
Iodometric assay
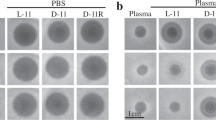
Inhibition studies revealed that the enzyme β-lactamase of ESBL E. coli and the growth of the bacterium were uniformly inhibited by MGDG-palmitoyl (100 μg ml−1), whereas ESBL E. coli growth was not inhibited by ampicillin (Supplementary Figure S8).
In silico docking study
The results of docking of MGDG-palmitoyl in orientation of blaTEM-1 interface are shown in Figure 6. The results reveal that the ligand has good interaction toward the targets with highest scores because of the lowest binding energy, H-bond distances and hydrophobic surface interaction toward the crystal structure of the molecule (Table 5).
Molecular docking analysis of Escherichia coli TEM1 extended spectrum β-lactamase with monogalactosyldiacylglycerol (MGDG)-palmitoyl complex (receptor–ligand). (a, b) Inner and outer view of receptor–ligand interaction. (c) Receptor–ligand was docked with highest binding energy and two hydrophobic interactions with Leu 198. Binding affinity of receptor–ligand complex was −4.9. A full color version of this figure is available at the Journal of Antibiotics journal online.
Discussion
During the past three decades, a great lot of information about antibiotic resistance has emerged and has been disseminated all over the world. Bacteria have developed resistance to almost all antibiotics in practice, whereas only a very few new antibiotics have been discovered for treatment of bacterial infections.26 Overuse of broad-spectrum β-lactams antibiotics in clinical practice is considered as the possible basis of development of resistance.27, 28 The spread of multidrug-resistant ESBL-producing pathogens has become a serious threat to public health, and a major concern for infection control practitioners.29
Some recent developments in respect of research on secondary metabolites based on bioassay-guided screening of prokaryotic and eukaryotic microalgae offer viable scientific and commercial potentials.30 However, until now these potentials have not been exploited to any great extent. In the present study, we focused to triumph over the ESBL-producing Gram-negative infection. Hence, in the light of antibacterial activities having been attributed to filamentous strains belonging to a wide range of genera of cyanobacteria31 we investigated the antibacterial effect of the methanol extract of O. acuminata NTAPC05 against ESBL-producing E. coli U655, S. maltophilia B929 and E. asburiae B938. In an earlier study, the extracts of O. princeps were shown as active against Bacillus subtilis, Staphylococcus aureus, E. coli and Brucella bronchiseptica32 but the active compound was not identified.
However, to date there is no authenticated report of antibacterial activity of cyanobacterial compounds on ESBL-producing bacteria. Therefore, this is the first study in this respect and it shows O. acuminata NTAPC05 fractions to inhibit ESBL producers, a rapidly growing Gram-negative infection. For the isolation of the antibacterial substance from O. acuminata NTAPC05, the methanol extract was fractionated under bioassay guidance, and assays were conducted to examine the effect of O. acuminata fractions on the growth of ESBL E. coli, S. maltophilia and E. asburiae. The differences in the susceptibility of different ESBL-producing bacteria may be because of the differences in their cell wall composition and/or the genetic constitution of their plasmids.33
Furthermore, we sought to determine the single peak of the active fraction from O. acuminata NTAPC05 by reverse phase HPLC. Data obtained from FTIR spectral analysis and 1H and 13C NMR study confirmed the functional groups as hydroxyl, ether and alkyl long-chain fatty acids that led to the conclusion that in O. acuminata NTAPC05 active fraction the palmitoyl in MGDG is 3-(3,4,5-trihydroxy-6-(hydroxymethyl) tetrahydro-2H-pyran-2-yloxy) propane-1,2-diyl dipalmitate that has not been reported previously.
Glycolipids represent a poorly studied class of metabolites with growing interest recently. Seaweeds biosynthesize three major types of glycolipids; viz, MGDGs, digalactosyldiacylglycerides and sulfoquinovosyldiacylglycerides. These glycoglycerolipids are present in chloroplasts of eukaryotic algae where MGDGs and digalactosyldiacylglycerides are the most abundant lipids of the thylakoid membrane and appear to play a crucial role in photosynthesis.34 For instance, cyanobacterial fatty acid and glycerolipid compositions closely resemble those of the inner envelope and thylakoid membranes of chloroplasts.35 The structure shows a glycosidic linkage between the sugar and C-3 of glycerol and ester linkage between fatty acids and two hydroxyls of glycerol, where galactose is the common sugar.36 A previous study showed that the MGDG and digalactosyldiacylglyceride isolated from Phormidium tenue37 and Chlorella vulgaris38 inhibit tumor-promoting stage of Epstein–Barr virus-associated early antigen (EBV-EAbhn). The fatty acyl chain length, its position and the nature of sugar moiety influence the activity. MGDGs, containing (7Z, 10Z)-hexadecadienoic acyl group, obtained from the green alga C. vulgaris, have been reported to exhibit antitumor property.38 Similarly, the medium-chain fatty acids of 8–12 carbon atoms exhibited antibacterial and antifungal properties that were enhanced when these fatty acids were esterified with glycerol.39 The composition of monogalactolipid mixture was determined for O. trichoide and it was found to inhibit HIV-1 reverse transcriptase enzyme activity.40 In our study, MGDG containing 3-(3,4,5-trihydroxy-6-(hydroxymethyl) tetrahydro-2H-pyran-2-yloxy) propane-1,2-diyl dipalmitate, with antibacterial activity against ESBL producers, has been identified in the marine cyanobacterium O. acuminata NTAPC05 that is being reported for the first time.
Although the exact mechanism underlying the antibacterial activity remains to be understood, it is generally known that bactericidal activities of antimicrobial compounds are due to their ability to penetrate and disrupt the integrity of the plasma membrane.41 In this study the results of CLSM and SEM analyses showed that the active fraction at MIC disrupted the membrane of ESBL-producing bacterial cells, leading to their death and disintegration. The antibacterial actions are thought to be linked to interactions of the biocides with the cell membrane of the microorganisms. The agents then enter the cell and finally act at various target sites. It is an established fact that disruption of the membrane by undesired or foreign substances can cause loss of integrity of the membrane that would lead to malfunction of the permeability barrier.42
As the ultimate objective of this study is to find a strategy to deal with the multidrug-resistant pathogenic ESBL-producing bacteria, it is essential to exonerate the possibility of the active compound inflicting toxicity to the humans. Pending an animal study, herein we tested the MGDG-palmitoyl against a normal cell line HEK293 and it does not affect the viability of this cell, indicating that the compound is not toxic to normal cells.
The TEM1 β-lactamase was docked with the MGDG-palmitoyl ligand that convincingly showed two hydrophobic interactions. Dhara et al.43 studied the docking interaction of TEM1 β-lactamase with the third-generation cephalosporins; viz., ceftazidime (ZINC ID: 03830469), cefotaxime (ZINC ID: 04468780) and cepfodoxime (ZINCID: 14235259). The data reveal that these third-generation cephalosporins require higher binding energy than MGDG-palmitoyl. An increase in the number of hydrophobic atoms in the active core of antimicrobial–target interface further increases the binding affinity between protein–antimicrobial interfaces. The palmitoylated compound attaches to long-chain fatty acid moieties that were identified and tested successfully both in vivo and in silico for the treatment of inflammation.44 The in silico analysis revealed that MGDG-palmitoyl from O. acuminata NTAPC05 efficiently interacts with the macromolecule that complemented and supported the observation in the iodometric assay. Thus, we have come up to the stage of finding a tangible lead to the effect that MGDG-palmitoyl from the marine cyanobacterium O. acuminata NTAPC05 offers potential to be used as an antibacterial agent against ESBL-producing pathogenic bacteria.
Conclusion
Herein we demonstrated the antibacterial activity of the MGDG-palmitoyl, isolated for the first time from O. acuminata NTAPC05, against urinary tract infection-causing ESBL producers. The compound is nontoxic to HEK293 normal cell line that ensures safety and, at the same time, offers a lead for chemical modifications that could allow its use as a therapeutic. Our results indicate that MGDG-palmitoyl is a new candidate for research on antibacterial substance (antibiotic) for the control of bacteremia caused by ESBL-producing multidrug-resistant bacteria.
References
Bradford, P. A. Extended-spectrum β-lactamases in the 21st century: characterization, epidemiology, and detection of this important resistance threat. Clin. Microbiol. Rev. 14, 933–951 (2001).
Rishi, H. P. D. & John, C. ESBLs: a clear and present danger. Critical Care Res. Pract. 2012, 625170 (2012).
Sanders, C. C. Chromosomal cephalosporinases responsible for multiple resistances to newer beta-lactam antibiotics. Ann. Rev. Microbiol. 41, 573–593 (1987).
Paterson, D. L. & Bonomo, R. A. Extended-spectrum b-lactamases: a clinical update. Clin. Microbiol. Rev. 18, 657–686 (2005).
Bonnet, R. Growing group of extended-spectrum b-lactamases: the CTX-M enzymes. Antimicrob. Agents Chemother. 48, 1–14 (2004).
Albertini, M. T. et al. Surveillance of methicillin-resistant Staphylococcus aureus (MRSA) and Enterobacteriaceae producing extended-spectrum beta-lactamase (ESBLE) in Northern France: a five-year multicentre incidence study. J. Hosp. Infect. 52, 107–113 (2002).
Teng, C. P., Chen, H. H., Chan, J. & Lye, D. C. Ertapenem for the treatment of extended-spectrum beta-lactamase-producing gram-negative bacterial infections. Int. J. Antimicrob. Agents 30, 356–359 (2007).
Rasheed, M. U., Thajuddin, N., Ahamed, P., Teklemariam, Z. & Jamil, K. Antimicrobial drug resistance in strains of Escherichia coli isolated from food sources. Rev. Inst. Med. Trop. São Paulo 56, 341–346 (2014).
Tandeau-de-Marsac, H. J. Adaptation of cyanobacteria to environmental stimuli: new steps towards molecular mechanisms. FEMS Microbiol. Lett. 104, 119–190 (1993).
Mayer, A. M. S. & Hamann, M. T. Marine pharmacology in 2001–2002: marine compounds with anti helmintic, antibacterial, anticoagulant, anti diabetic, antifungal, anti-inflammatory, anti malarial, anti platelet, antiprotozoal, anti tuberculosis, and antiviral activities, affecting the cardiovascular, immune and nervous systems and other miscellaneous mechanisms of action. Comp. Biochem. Physiol. C 140, 265–286 (2005).
Moreau, J. D., Pasando, P., Bernand, P., Caram, B. & Pinnat, J. C. Seasonal variation in the production of antifungal substrates by dictyotales (brown algae) from the French Mediterranean coast. Hydobiologia 2, 1097–1132 (1988).
Renu, A. Antibacterial activities of freshwater algae Chlorella ellipsoidea. J. Basic Appl. Biol. 4, 22–26 (2010).
Abed, R. M. M. et al. Cyanobacterial diversity and bioactivity of inland hyper saline microbial mats from a desert stream in the Sultanate of Oman. Fottea 11, 215–224 (2011).
Rama murthy, V., Raveendran, S., Thirumeni, S. & Krishnaveni, S. Antimicrobial activity of heterocytic cyanobacteria. Int. J. Adv. Lif. Sci. 1, 32–39 (2012).
CLSI Performance Standards for Antimicrobial Susceptibility Testing: 22nd Informational Supplement M100-S22, (Clinical and Laboratory Standard Institute, Wayne, PA, (2012).
Desikachary, T. V. Cyanophyta, (ICAR Publication, New Delhi, (1959).
Smoker, J. A. & Barnum, S. R. Rapid small scale DNA isolation from filamentous cyanobacteria. FEMS Microbiol. Let. 56, 119–122 (2006).
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Cheesbrough, M. inMedical Laboratory Manual for Tropical Countries Vol. 2, 2–392 (Tropical Health Technology Publications and Butterworth–Heinemann, Cambridge, UK, (2002).
Wikler, M. A. Performance Standards for Antimicrobial Susceptibility Testing; Eighteenth Informational Supplement; M100-S18. C.L.S.I. (Clinical and Laboratory Standard Institute), Pennsylvania, PA, USA 28, 46–52 (2008).
Weigand., I., Hilpert, R. & Hancock, R. E. W. Agar and broth dilution methods to determine the minimum inhibitory concentration (MIC) of antimicrobial substance. Nat. Protoc. 3, 163–175 (2008).
Eliopoulos, G. M. & Moellering, R. C. inAntibiotics in Laboratory Medicine 4thedn(ed.Lorain V) 330–396 The Williams & Wilkins, Baltimore, MD, USA, (1996).
Denizot, F. & Lang, R. Rapid colorimetric assay for cell growth and survival: modifications to the tetrazolium dye procedure giving improved sensitivity and reliability. J. Immunol. Methods 89, 271–277 (1986).
Ahmad, S. & Yadava, J. N. S. Rapid detection of b-lactam antibiotic resistance among clinical isolates of Escherichia coli. India Vet. Med. J. 3, 256–259 (1979).
Morris, G. M. et al. AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J. Comput. Chem. 30, 2785–2791 (2009).
Payne, D. J. Desperately seeking new antibiotics. Science 321, 1644–1645 (2008).
Lipsitch, M., Bergstrom, C. T. & Levin, B. R. The epidemiology resistance in hospital; Paradoxes and prescription. Proc. Natl Acad. Sci. USA 97, 1938–1943 (2000).
Pitout, J. D., Hanson, N. D. & Church, D. L. Population-based laboratory surveillance for Escherichia coli producing extended spectrum beta lactamases: Importance of community isolates with blaCTX-M genes. Clin. Infect. Dis. 38, 1736–1741 (2004).
Sanders, C. C. & Sanders, W. E. β-Lactamase resistance in Gram-negative bacteria: global trends and clinical impact. Clin. Infect. Dis. 15, 824–839 (1992).
Kerby, N. N. & Rowell, P. inPhotosynthetic Prokaryotes (eds Mann, N H & Car,N.G.) 233–254 (Plenum Press, New York, (1992).
Gupta, A. B. & Shrivastava, G. C. On antibiotic properties of some fresh water algae. Hydrobiologia 25, 285–288 (1965).
Schlegel, I., Doan, N. T., Chazal, N. & Smith, G. D. Antibiotic activity of new cyanobacterial isolates from Australia and Asia against green algae and cyanobacteria. J. Appl. Phycol. 10, 471–479 (1999).
Harriman, A. (Photo) isomerization dynamics of merocyanine dyes in solution. J. Photochem. Photobiol. A 65, 79–93 (1992).
Holzl, G. & Dormann, P. Structure and function of glycoglycerolipids in plants and bacteria. Prog. Lipid Res. 46, 225–243 (2007).
Heinz, E. & Roughan, P. G. Similarities and differences in lipid metabolism of chloroplasts isolated from 18:3 and 16:3 plants. Plant Physiol. 72, 273–279 (1983).
Dormann, P. & Benning, C. Galactolipids rule in seed plants. Trends Plant Sci. 7, 112–118 (2002).
Shirahashi, H. et al. Isolation and identification of anti-tumor-promoting principles from the fresh-water cyanobacterium Phormidium tenue. Chem. Pharm. Bull. 41, 1664–1666 (1993).
Morimoto, T. et al. Antitumor promoting glycerol-glycolipids from the green alga Chlorella vulgaris. Phytochemistry 40, 1433–1437 (1995).
Frentzen, M., Weier, D. & Feussner, I. Reports on symposia and congresses. Eur. J. Lipid Sci. Technol. 105, 784–792 (2003).
Vered, R. et al. New acylated sulfoglycolipids and digalactolipids and related known glycolipids from cyanobacteria with a potential to inhibit the reverse transcriptase of HIV-1. J. Nat. Prod. 60, 1251–1260 (1997).
Shai, Y. Mode of action of membrane active antimicrobial peptides. Biopolymers 66, 236–248 (2002).
Hameed, A. S. et al. In vitro antibacterial activity of ZnO and Nd doped ZnO nanoparticles against ESBL producing Escherichia coli and Klebsiella pneumoniae. Sci. Rep. 6, 24312 (2016).
Dhara, L., Tripathi, A. & Pal, A. Molecular characterization and in silico analysis of naturally occurring TEM β-lactamase variants among pathogenic Enterobacteriaceae infecting Indian patients. Biomed. Res. Int. 2013, 783540 (2013).
Bhagavat, R., Saqib, A. & Karigar, C. Molecular docking studies of novel palmitoyl- ligands for cyclooxygenase-2. Chem. Biol. Drug Des. 79, 1043–1048 (2012).
Acknowledgements
We are grateful to Dr Hemalatha Rao, Medwin Hospital (Hyderabad, India) for providing the clinical isolates. We thank the Deanship of Scientific Research at King Saud University for funding the work through the research group project (Ref No RGP-VPP-332). We thank DST- PURSE scheme (Project Ref No- SR/FT/LS-113/2009) of the Department of Science and Technology (DST), New Delhi, India, for the Confocal Laser Scanning Microscopy facility. This work was otherwise supported by the Project (Ref No BT/PR4815/AAQ/3/587/2012, BT/PR6619/PBD/26/310/2013, BT/IN/Indo-UK/SuBB/23/NT/2013 and BT/PR7005/PBD/ 26/357/2015) sanctioned by the Department of Biotechnology (DBT), New Delhi, India.
Author contributions
AP and NT performed most of the experiments and drafted the manuscript; MUR carried out the sample collection from humans and identified the bacterial strains; KPMN investigated the antibiotic resistance pattern and identified ESBL producers; NR contributed to the amplification of ESBL genes and interpretation of results; SS obtained the microphotograph and identification of cyanobacterium; MYMI contributed in 16S DNA amplification in ESBL producers; NSA, CA and SAA equally contributed in the analysis and interpretation of data; MAA carried out the cell line study, coordinated the manuscript drafting and language editing. All authors approved the final version of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on The Journal of Antibiotics website
Supplementary information
Rights and permissions
About this article
Cite this article
Parveez Ahamed, A., Rasheed, M., Peer Muhamed Noorani, K. et al. In vitro antibacterial activity of MGDG-palmitoyl from Oscillatoria acuminata NTAPC05 against extended-spectrum β-lactamase producers. J Antibiot 70, 754–762 (2017). https://doi.org/10.1038/ja.2017.40
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2017.40
This article is cited by
-
Nanosynthesis, phycochemical constituents, and pharmacological properties of cyanobacterium Oscillatoria sp.
Naunyn-Schmiedeberg's Archives of Pharmacology (2024)
-
Assessment of Antimicrobial and Antioxidant Potential of Oscillatoria sancta and Oscillatoria proteus Isolated from Chilika Lake
Current Microbiology (2024)
-
Synthesis and Characterization of Novel Schiff Bases Derived from 2-Butyl-4-chloro Imidazole for the Enhanced Antimicrobial Property
Applied Biochemistry and Biotechnology (2023)
-
Anti-quorum Sensing and Anti-biofilm Effect of Nocardiopsis synnemataformans RMN 4 (MN061002) Compound 2,6-Di-tert-butyl, 1,4-Benzoquinone Against Biofilm-Producing Bacteria
Applied Biochemistry and Biotechnology (2023)
-
In vitro antimicrobial and antioxidant activities of certain brackish water cyanobacteria from Chilika Lake, India
Vegetos (2022)