Abstract
Osteoblast and adipocyte are differentiated from mesenchymal stem cells and dysregulation of the differentiation might result in disease, such as osteoporosis and diabetes. To find small compounds that induce osteoblast differentiation, we screened an in-house natural compounds library with mouse preosteoblastic MC3T3-E1 cells using alkaline phosphatase (ALP) expression as an early osteoblast marker. We found that phenazine-1-carboxylic acid (PCA), one of the major phenazine derivatives produced by Pseudomonas, induced osteoblast differentiation in the cells at micromolar concentrations. PCA acted synergistically with an agonist of hedgehog signaling in inducing ALP activity in the cells. We also found that 2-hydroxy-PCA (2H-PCA) induced osteoblast differentiation in the cells but 2-methoxy-PCA and 1-hydroxy-phenazine did not. Unexpectedly, treatment of mouse pluripotent mesenchymal C3H10T1/2 cells with PCA or 2H-PCA induced an obvious morphological change. Oil Red O staining and real-time reverse-transcription PCR analysis revealed that PCA induced not osteoblast differentiation but adipocyte differentiation in C3H10T1/2 cells. These compounds could allow us to investigate the mechanism of osteoblast and adipocyte differentiation in the two model cell systems through a chemical biology approach.
Similar content being viewed by others
Main
Mesenchymal stem cells (MSCs) are adult stem cells that carry potential for differentiation into various cell lineages, such as osteoblasts and adipocytes.1 These cell lineages are important for bone and lipid homeostasis in the body.2, 3 MSC differentiation is tightly controlled by the local environment in the body and an imbalance of MSC differentiation might result in disease, such as osteoporosis and diabetes. In addition, MSCs can be easily isolated from bone marrow and adipose and are one of the major cell sources for regenerative therapy.1 Therefore, small compounds that affect MSC differentiation such as osteogenesis and adipogenesis could be useful for study of these events and might provide important knowledge for the development of regenerative therapy and novel drugs.
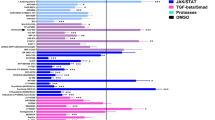
To find small compounds that induce osteoblast differentiation, we screened an in-house natural compounds library with mouse preosteoblastic MC3T3-E1 cells using alkaline phosphatase (ALP) expression as an early osteoblast marker.2 In the course of the screening, we found that phenazine-1-carboxylic acid (PCA), one of the major phenazine derivatives produced by Pseudomonas,4 induced ALP activity in MC3T3-E1 cells at micromolar concentrations (Figure 1b). The induction of ALP activity by PCA was dose-dependent and was weaker than that of purmorphamine, a small-molecule agonist of hedgehog signaling5 (Figure 1c). Real-time reverse-transcription PCR analysis revealed that 7-day treatment with 10 μM PCA induced Alpl mRNA, the gene encoding ALP. PCA also induced the mRNA expression of the osteoblastic genes Ocn, Opn and Runx2 (Figure 1d). Interestingly, we found that PCA acted synergistically with purmorphamine in inducing ALP activity in MC3T3-E1 cells (Figure 1e). These data demonstrated that PCA induces osteoblast differentiation in MC3T3-E1 cells.
Phenzazine-1-carboxylic acid (PCA) treatment induces osteoblastic differentiation of the mouse preosteoblast cell line MC3T3-E1. (a) Chemical structures of phenazine derivatives used in this study. (b) MC3T3-E1 cells were treated with 10 μM of indicated compounds in 96-well plates. After 6 days of treatment, cells were fixed and stained for ALP. Scale bars=200 μm. (c) MC3T3-E1 cells were treated with PCA or purmorphamine (PM) at increasing concentrations as indicated. After 6 days of treatment, ALP activities were measured using AttoPhos substrate. Data are shown as the mean and s.d. of three independent experiments performed in duplicate. *P<0.05, **P<0.01 and ***P<0.005, Student’s t-test, compared with untreated controls. (d) Real-time reverse-transcription PCR (RT-PCR) analysis of osteogenic genes Alpl, Ocn, Osteopontin (Opn) and Runx2. Cells were treated with 10 μM PCA derivatives for 7 days and then total RNA was extracted and subjected to real-time RT-PCR. Data were first normalized to the level of Gapdh and are presented as fold change versus vehicle controls. Data represent the mean and s.d. of three independent experiments. *P<0.05, **P<0.01 and ***P<0.005, Student’s t-test, compared with vehicle controls. (e) ALP activities were measured after 6 days of combination treatment of 10 μM PCA with PM. Data are shown as the mean and s.d. of three independent experiments performed in duplicate. *P<0.05, **P<0.01 and ***P<0.005, Student’s t-test, compared with PCA (−) group. A full colour version of this figure is available at the Journal of Antibiotics journal online.
We next examined whether the three phenazine derivatives, 2-hydroxy-PCA (2H-PCA), 2-methoxy-PCA (2ME-PCA) and 1-hydroxy-phenazine (1H-phenazine), could induce osteoblast differentiation in MC3T3-E1 cells. A 7-day treatment with 10 μM 2H-PCA also induced mRNA expression of the osteoblastic genes Alpl, Ocnand Runx2 (Figure 1d). However, treatment with 2ME-PCA or 1H-phenazine had no effect on these genes (Figure 1d). At this concentration, these compounds did not show any cytotoxic effects on the cells at the cellular density of the differentiation experiments (Supplementary Figure S1). We also performed a time course (4, 8 and 12 days) study about the effect of 2H-PCA treatment on the osteoblast differentiation of MC3T3-E1 cells (Supplementary Figure S2). In 2H-PCA treatment cells, expression of the early osteoblast maker Alpl reached a peak level at day 4 but expression of the late osteoblast marker Ocn increased during the 12-day treatment. Thus, our results demonstrated that the two phenazine derivatives have osteogenic activity on MC3T3-E1 cells and that the functional groups at C-1 and C-2 substantially affect the potency of the osteogenic activity of the phenazine derivatives.
Mouse pluripotent mesenchymal C3H10T1/2 cells, such as MSCs, are able to differentiate into various cells such as osteoblasts and adipocytes.6, 7, 8 Next, we investigated the effect of PCA on the differentiation of C3H10T1/2 cells. After an 8-day treatment with 10 μM PCA, C3H10T1/2 cells showed an obvious morphological change. This morphological change was same as that of C3H10T1/2 cells treated with troglitazone, an activator of nuclear receptor peroxisome proliferator-activated receptor-γ (PPARγ) that induces adipogenesis in the cell line.9 Oil Red O staining supported the conclusion that the morphologically changed cells were adipocytes (Figure 2a). The induction of Oil Red O-positive cells by PCA was dose dependent (Figure 2b). Real-time reverse-transcription PCR analysis demonstrated that 8-day treatment with 10 μM PCA-induced expression of adipogenic genes Fabp4, Lpl, Adiponectin and Cebpa (Figure 2c). We also investigated whether the three phenazine derivatives could induce adipocyte differentiation in C3H10T1/2 cells. The 8-day treatment with 10 μM 2H-PCA induced Oil Red O-positive cells in a dose-dependent manner (Figure 2b). However, treatments with 2ME-PCA or 1H-phenazine did not induce Oil Red O-positive cells (Figure 2b). Treatment with 10 μM 2H-PCA also induced mRNAs of the adipogenic genes, but treatment with 2ME-PCA or 1H-phenazine had no effect on these levels (Figure 2c and Supplementary Figure S2). On the other hand, the early osteoblastic marker ALP activity was not induced by treatment with PCA and its derivatives in C3H10T1/2 cells (Supplementary Figure S4). These results demonstrated that PCA and 2H-PCA induce adipocyte differentiation rather than osteoblast differentiation in C3H10T1/2 cells, and that the functional groups at C-1 and C-2 also substantially affect the potency of the adipogenic activity of phenazine derivatives. This parallel relationship between structures and activities in osteoblast differentiation in MC3T3-E1 cells and adipocyte differentiation in C3H10T1/2 cells could imply the existence of a common target(s) in both cell lines. We also examined whether 2H-PCA could induce differentiation of mouse MSCs derived from the bone marrow of C57BL/6 mice (Supplementary Figures S5 and S6). However, the treatment with 2H-PCA did not induce osteogenic and adipogenic differentiation on the mouse MSCs in our experimental conditions.
PCA treatment induces adipocyte differentiation in the mouse pluripotent mesenchymal cell line C3H10T1/2. (a) C3H10T1/2 cells were treated with 10 μM of the indicated compounds for 8 days in 96-well plates and stained with Oil Red O. The images were taken at × 100 magnification. Scale bars=200 μm. (b) Numbers of Oil Red O (+) cells in each well were quantified by light microscopy. Data are shown as the mean and s.d. of three independent experiments performed in duplicate. (c) Real-time reverse-transcription PCR (RT-PCR) analysis of adipogenic genes Fabp4, Lpl, Adiponectin and Cebpa. Cells were treated with 10 μM PCA derivatives for 8 days and then total RNA was extracted and subjected to real-time RT-PCR. Data were first normalized to the level of Gapdh and are presented as fold change versus vehicle controls. Data represent the mean and s.d. of three independent experiments. *P<0.05, **P<0.01 and ***P<0.005, Student’s t-test, compared with vehicle controls. (d) PPARγ reporter assay. C3H10T1/2 cells and MC3T3-E1 cells were transiently transfected with PPAR-RE-Luc reporter plasmid and PPARγ2 and RXR cDNA expression plasmids. After treatment for 24 h with 10 μM 2H-PCA, the dual luciferase assay was performed. Data are shown as the mean and s.d. obtained from three independent experiments performed in triplicate. *P<0.05, Student’s t-test, compared with vehicle controls. A full colour version of this figure is available at the Journal of Antibiotics journal online.
As both Fabp4 and Lpl are transcriptional target genes of PPARγ, we next investigated whether 2H-PCA could enhance PPARγ-activated transcription using a PPARγ reporter assay in C3H10T1/2 cells and MC3T3-E1 cells (Figure 2d). Treatment with 10 μM troglitazone induced PPARγ reporter activation by approximately twofold compared with control in both cell lines. In contrast, treatment with 10 μM 2H-PCA, at the concentration that induced adipogenesis in C3H10T1/2 cells, did not induce significant reporter activation. These results suggest that 2H-PCA does not function as a PPARγ ligand and other important target(s) for the induction of the differentiation should exist.
Taken together, we found a new biological activity of phenazine derivatives in osteoblast and adipocyte differentiation of the two model cell systems. These natural compounds, which have been researched intensively owing to their various biological activities, could help us to investigate the mechanism of osteogenesis and adipogenesis in the two model cell systems through a chemical biology approach.
Materials and methods
Preparation of the Phenazine derivatives
1H-phenazine was obtained from an in-house chemical library.
PCA was purchased from Princeton Bio-molecular Research (Princeton, NJ, USA). 2H-PCA was prepared as reported10 and identified by comparison of the spectroscopic data.11 For preparation of 2ME-PCA, methyl 2-methoxyphenazine-1-carboxylate was prepared as reported:10 1H-NMR (400 MHz, CDCl3) d 4.10 (3H, s), 4.13 (3H, s), 7.74 (1H, d, J=10 Hz), 7.78 (1H, ddd, J=2, 7, 8 Hz), 7.81 (1H, ddd, J=2, 7, 8 Hz), 8.18 (1H, dd, J=2, 8 Hz), 8.20 (1H, dd, J=2, 8 Hz), 8.31 (1H, d, J=10 Hz); 13C-NMR (100 MHz, CDCl3) d 52.8, 57.0, 117.6, 119.2, 129.5, 129.8, 129.9, 130.9, 132.5, 138.9, 141.6, 142.3, 144.0, 156.6, 167.2; HR-ESI-MS m/z 269.0919 [M+H]+ calcd for C15H13N2O3, 269.0921. Then, methyl 2-methoxyphenazine-1-carboxylate (29 mg, 0.11 mmol) was dissolved in methanol (3 ml) and 4 M sodium hydroxide (6 ml). The solution was stirred at room temperature for 2.5 h. The reaction mixture was cooled, acidified with dilute hydrochloric acid and extracted with ethyl acetate. The organic solution was washed with saturated sodium chloride solution, dried over anhydrous sodium sulfate and concentrated under reduced pressure. The resulting residue was purified by semi-preparative high-performance liquid chromatography (35% acetonitrile water) to give 2ME-PCA (21 mg, 76%) as a yellow solid: 1H-NMR (400 MHz, CDCl3) d 4.29 (3H, s), 7.89 (1H, ddd, J=1, 7, 9 Hz), 7.92 (1H, d, J=10 Hz), 7.97 (1H, ddd, J=1, 7, 9 Hz), 8.21 (1H, dd, J=1, 9 Hz), 8.26 (1H, dd, J=1, 9 Hz), 8.46 (1H, d, J=10 Hz); 13C-NMR (100MHz, CDCl3) d 57.7, 107.9, 120.9, 127.3, 129.8, 130.4, 133.1, 135.8, 139.4 (2C), 141.8, 142.0, 164.5, 165.6; HR-ESI-MS m/z 255.0763 [M+H]+ calcd for C14H11N2O3, 255.0764.
Cell culture
Mouse C3H10T1/2 cells were cultured in Dulbecco’s modified Eagle’s medium (Wako Pure Chemical, Osaka, Japan) containing 10% fetal bovine serum and 1% penicillin/streptomycin. Mouse MC3T3-E1 cells were cultured in MEMα (Wako Pure Chemical) containing 10% fetal bovine serum and 1% penicillin/streptomycin. To induce cellular differentiation, cells were plated in 96-well plates at a density of 10 000 cells per well or in 6-well plates at a density of 400 000 cells per well. After reaching confluence, cells were treated with compounds.
PPARγ reporter assay
C3H10T1/2 cells and MC3T3-E1 cells were transiently transfected with PPAR-RE-Luc reporter plasmid and PPARγ2 and RXR cDNA expression plasmids using FuGeneHD transfection reagent (Promega, Madison, WI, USA) according to the manufacturer’s instructions. One day after transfection, the cells were treated with 10 μM troglitazone or 10 μM 2H-PCA in OPTI-MEM medium (Invitrogen, Carlsbad, CA, USA) containing 5% fetal bovine serum. After 24 h, cells were assayed for firefly and Renilla luciferase activities using the Dual-Glo Luciferase Assay System (Promega) according to the manufacturer’s instructions.
Other experiments such as ALP staining, ALP assay, Oil Red O staining and real-time reverse-transcription PCR were performed as previously described.6, 7
Dedication
This paper is dedicated to Professor Satoshi Ōmura on the occasion of his Nobel Prize in Physiology or Medicine 2015.
References
Nombela-Arrieta, C., Ritz, J. & Silberstein, L. E. The elusive nature and function of mesenchymal stem cells. Nat. Rev. Mol. Cell Biol. 12, 126–131 (2011).
Komori, T. Regulation of osteoblast differentiation by transcription factors. J. Cell. Biochem. 99, 1233–1239 (2006).
Cristancho, A. G. & Lazar, M. A. Forming functional fat: a growing understanding of adipocyte differentiation. Nat. Rev. Mol. Cell Biol. 12, 722–734 (2011).
Pierson, L. S. III & Pierson, E. A. Metabolism and function of phenazines in bacteria: impacts on the behavior of bacteria in the environment and biotechnological processes. Appl. Microbiol. Biotechnol. 86, 1659–1670 (2010).
Wu, X., Walker, J., Zhang, J., Ding, S. & Schultz, P. G. Purmorphamine induces osteogenesis by activation of the Hedgehog signaling pathway. Chem. Biol. 11, 1229–1238 (2004).
Sakamoto, S. et al. Decalpenic acid, a novel small molecule from Penicillium verruculosum CR37010, induces early osteoblastic markers in pluripotent mesenchymal cells. J. Antibiot. 63, 703–708 (2010).
Sakamoto, S. et al. Decalpenic acid induces early osteoblastic markers in pluripotent mesenchymal cells via activation of retinoic acid receptor γ. Biochem. Biophys. Res. Commun. 422, 751–757 (2012).
Huang, H. et al. BMP signaling pathway is required for commitment of C3H10T1/2 pluripotent stem cells to the adipocyte lineage. Proc. Natl Acad. Sci. USA 106, 12670–12675 (2009).
Bäckesjö, C. M., Li, Y., Lindgren, U. & Haldosén, L. A. Activation of Sirt1 decreases adipocyte formation during osteoblast differentiation of mesenchymal stem cells. J. Bone Miner. Res. 21, 993–1002 (2006).
Brooke, P. K. et al. Synthesis of some methoxy- and hydroxy-phenazine-1-carboxylic acids. JCS Perkin1 16, 2248–2251 (1976).
Mehnaz, S. et al. Lahorenoic acids A-C, ortho-dialkyl-substituted aromatic acids from the biocontrol strain Pseudomonas aurantiaca PB-St2. J. Nat. Prod. 76, 135–141 (2013).
Acknowledgements
We thank I Momose, M Igarashi, S Ohba and M Arakawa for their helpful comments and technical support. This work was supported in part by a grant-in-aid from the Ministry of Education, Culture, Sports, Science and Technology of Japan (to SS, number 20790218 and 24102532).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on The Journal of Antibiotics website
Supplementary information
Rights and permissions
About this article
Cite this article
Sakamoto, S., Watanabe, T., Kohda, Y. et al. Phenazine carboxylic acid and its derivative induce osteoblast differentiation in preosteoblastic MC3T3-E1 cells but adipocyte differentiation in pluripotent mesenchymal C3H10T1/2 cells. J Antibiot 70, 1146–1149 (2017). https://doi.org/10.1038/ja.2017.129
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2017.129