Abstract
A convenient and efficient method was established for isolating antifungal antibiotic-producing fungi from soil samples. In this method, soil samples were diluted and directly plated in agar medium by the standard fungi-isolating method, and the plates were cultured at 27 °C for 2–3 days to permit the growth of fungal colonies. Then, the suspension of pathogenic Candida albicans in saline (40 μl, 5–10 × 105 CFU ml−1) was overlaid by spraying on the plates under controlled conditions in the safety cabinet. After 1-day incubation, fungal colonies showing an antagonistic effect with the inhibition zone against sprayed C. albicans were selected. Among 151 isolates, 26 strains were found to reproduce anti-C. albicans activity in liquid medium, yielding a higher selection rate (17.2%) than that (3.1%) by the traditional method. This new method can be applied for isolation of microorganisms (fungi and actinomycetes) that produce antibiotics active against pathogenic microorganisms.
Similar content being viewed by others
Introduction
It is well known that fungi remain one of the most important resources for the discovery of new bioactive compounds.1 It is thought that fungi rank as the second biggest kingdom of organisms in nature and that as many as 1.5–5.1 million fungal species exist.2, 3, 4 From the history of drug discovery from microorganisms, fungal secondary metabolites have provided a number of important drugs, such as the antibiotic penicillin,5 the immunosuppressant cyclosporine6 and the antihypercholesterolemic agents lovastatin7 and compactin.8, 9 Recent DNA sequencing technology has markedly advanced, revealing whole-genome sequences of a number of organisms. The first whole-genome sequence was published for a free-living organism, Haemophilus influenza.10 Since then, a number of organism genome sequences, including bacteria (more than 1500 species), archaea (more than 100 species) and eukaryotes (more than 100 species, including Homo sapiens), have been determined.11 From genomic information about microorganisms, Streptomyces species have 7–9-Mb-long genomes containing 20–30 genes of secondary metabolites biosynthesized via pathways of polyketide synthases or non-ribosomal peptide synthases,12, 13, 14 while fungi such as Aspergillus species have 30–40-Mb-long genomes containing 30–80 polyketide synthase and non-ribosomal peptide synthase genes.15, 16, 17 Interestingly, only a few genes can be expressed to produce the corresponding secondary metabolites, but most of them remain silent.18, 19 Thus, these microorganisms were proved to have high potential to produce many secondary metabolites.
Many antibiotics have been discovered from microorganisms. The process of discovering antibiotics usually includes several steps: (1) isolation of microorganisms mainly from soil (usually 2–3 days are needed), (2) pure culture of isolated microorganisms (3–5 days), (3) identification of isolated microorganisms mainly by morphological characteristics to eliminate duplicate strains (14–21 days), (4) testing whether or not these culture broths or extracts of microorganisms show antimicrobial activity to select candidate microorganisms that produce antibiotics (7–8 days) and (5) re-culture of candidate microorganisms to observe the reproducibility of antimicrobial activity (6–8 days). This routine process (named the traditional method in this study) usually takes 5–7 weeks. In this study, we show a more efficient and faster method to select candidate fungi that can produce anti-Candida albicans antibiotics.
Results
Media for fungal isolation
As C. albicans was usually cultured in GY medium, the growth of C. albicans on media 1–4 was investigated. C. albicans was found to grow normally in media 1–4; therefore, all the media were used throughout this study.
Preparation of soil plates
By plating method A, in which a soil sample (15 mg) was directly dispensed into each medium (media 1–4), fast-growing fungi such as Trichoderma, Aspergillus and Penicillium species predominantly appeared on the plate and inhibited the growth of other fungi. Colonies of slow-growing fungi such as Virgaria and Humicola species were isolated on the plates of highly diluted soil samples by plating method B. Thus, both plating methods A and B were used to prepare soil plates. Soil plates were incubated at 27 °C for 2–3 days to form fungal colonies.
Inoculation of C. albicans to soil plates
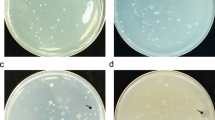
The fundamental aim of this study was to directly observe fungal colonies derived from soil that show antagonistic effects on the growth of C. albicans. Five C. albicans-inoculating methods were tested. In inoculating method I, C. albicans and the soil sample were mixed and incubated at the same time. In inoculating method II, C. albicans-containing agar was first prepared, and soil-containing agar was overlaid on the plate, which was then incubated. Thus, in inoculating methods I and II, pathogenic C. albicans and soil-derived fungi began to grow almost at the same time. As a result, the plate was covered with C. albicans (Figure 1a), because C. albicans grew much faster than soil-derived fungi. By these methods, soil-derived fungi could not form colonies and antagonistic effects were not observed on the plates. In inoculating method III, C. albicans was spread on soil plates after soil-derived colonies were formed on the plates (2–3-day incubation at 27 °C); however, the spreader shaved soil-derived fungal colonies, which were spread on the plate. Unfortunately, C. albicans could not grow evenly on the plate (Figure 1b). In inoculating method IV, C. albicans-containing agar was overlaid on the soil plates after the formation of soil-derived fungal colonies (2–3-day incubation at 27 °C); however, soil-derived fungal colonies were also diffused on the plate when C. albicans-containing agar was overlaid (Figure 1c).
Inoculation of C. albicans into soil plates. (a) Inoculating method I: the plate was covered with C. albicans, and soil-derived fungi could not form colonies. (b) Inoculating method III: The spreader-shaved soil-derived fungal colonies and C. albicans could not grow evenly on the plate. (c) Inoculating method IV: soil-derived fungal colonies were diffused on the plate by C. albicans-containing agar.
Thus, C. albicans-inoculating methods I–IV were not suitable for our purpose; therefore, we tested another method of overlaying C. albicans on the soil plate by spraying (inoculating method V). In this method, it is expected that soil-derived fungal colonies remain intact, and that C. albicans can be overlaid evenly on the soil plate. To establish this method, the concentration of C. albicans suspension for spraying and the spraying time after formation of soil-derived fungal colonies were investigated.
Establishment of inoculating method V
Using the atomizer in this study, a 40-μl solution can be sprayed. Five concentrations of C. albicans suspension (40 μl, 1.0 × 105–5.0 × 106 CFU ml−1) were tested for spraying on the soil plate and the growth of C. albicans was compared after 1–3-day incubation at 27 °C. Figure 2 shows C. albicans colonies sprayed on the agar plate (without soil) at the five concentrations 1 day after inoculation. C. albicans grew rapidly, so 1-day incubation was enough for C. albicans to form colonies. As shown in Figure 2, about 1000 and 2000 colonies of C. albicans had formed on the plates 1 day after inoculation by spraying at (c) 5.0 × 105 and (d) 1.0 × 106 CFU ml−1, respectively. These two concentrations appeared appropriate to observe antagonistic effects.
The conditions were established accordingly; that is, in inoculating method V, soil plates were prepared by plating method A or B and incubated at 27 °C. After 2- or 3-day incubation to form soil-derived fungal colonies, C. albicans suspension (40 μl, 5–10 × 105 CFU ml−1) was sprayed on the soil plates in the plastic bag installed in the biological safety cabinet (Figure 3). After 1-day incubation at 27 °C, fungal colonies showing antagonistic effects on the growth of C. albicans were selected (arrowed colony (a) in Figure 4).
Selection of soil-derived fungi showing antagonistic effects on C. albicans. Soil plates were prepared by plating method A. After 2-day incubation to form colonies, C. albicans suspension (40 μl, 1.0 × 106 CFU ml−1) was sprayed on the soil plates by Inoculating method V. (a) A fungal colony showing antagonistic effect on C. albicans was selected. (b) A fungal colony showing no antagonistic effect on C. albicans. (c) Colonies of C. albicans.
Screening result
From 18 soil samples, 151 fungal strains that showed antagonistic effects on the growth of C. albicans were selected (Table 1); therefore, they were re-cultured in the liquid medium to confirm whether their culture broths exhibited an inhibition zone against C. albicans. Among them, the culture broths of 26 strains were reproduced to show an inhibition zone against C. albicans.
Taxonomy of selected 26 strains
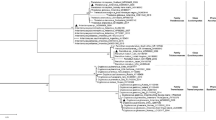
The results of the taxonomy of 26 selected fungi by morphological observation are summarized in Table 1. They were identified as Penicillium sp. (16 strains), Trichoderma sp. (3), Cylindrocarpon sp. (2), Aspergillus sp. (1), Metarhizium sp. (1), Humicola sp. (1), Acremonium sp. (1) and Beauveria sp. (1).
Discussion
In this study, we established an efficient and rapid method for directly isolating fungi from soil samples that can produce anti-C. albicans antibiotics. As described in inoculating method V, after soil-derived colonies had formed on the plates (on days 2–3), C. albicans (40 μl, 5–10 × 105 CFU ml−1) was overlaid by spraying on the plates. After 1-day incubation, soil-derived fungi that showed antagonistic effects on the growth of C. albicans were selected. By this method, 151 candidate fungi from 18 soil samples showing antagonistic effects were selected within a week and, among them, 26 fungi exhibited anti-C. albicans activity in their re-culture broths. Thus, this method permits us to obtain antifungal antibiotic-producing fungi from soil samples in a shorter time (2 weeks vs 5–7 weeks) and at a higher rate (17.2% vs 3.1%) than the traditional method. The 151 candidate fungi were re-cultured in one liquid medium; therefore, if they were cultured in other media, the hit rate would increase. As summarized in Table 1, Penicillium was most highly selected as an antifungal-producing genus (16/26), followed by Trichoderma (3/26). Using the traditional method carried out by our group in 2008, 730 fungi were isolated from 18 soil samples. Penicillium was the most highly isolated genus (173), followed by Paecilomyces (100) and Trichoderma (93) (unpublished data); therefore, it is intriguing that two Cylindrocarpon strains were selected as anti-C. albicans antibiotic-producing fungi by this method. In the accompanying study,20 we describe that a known antifungal antibiotic, ascochlorin, was isolated from the culture broth of one Cylindrocarpon strain. It is expected that this method will be applicable for isolating soil-derived microorganisms (fast-growing/slow-growing fungi and actinomycetes) showing antagonistic effects on other pathogenic microorganisms.
Materials and methods
Soil samples
We used 18 soil samples. Eighteen samples (No. 1–18) were collected in Kanagawa and Tokyo (Japan).
Media for fungal isolation
Four media were used for isolation of fungi from samples (soil). Kanamycin (Meiji Seika Pharma, Tokyo, Japan) was added to all media to protect against bacteria growth in soil samples.
Medium 1 contains sucrose 2.0%, glucose 1.0%, corn steep powder (Marcor Development, Carlstadt, NJ, USA) 0.5%, bonito extract (Kyokuto Pharmaceutical Industrial, Tokyo, Japan) 0.5%, KH2PO4 0.1%, CaCO3 0.3%, kanamycin 0.1 mg ml−1 and agar 1.5%, adjusted to pH 6.0 before sterilization.
Medium 2 contains soluble starch 3.0%, glycerol 1.0%, soybean meal 2.0%, dry yeast (Fermipan, GB Ingredients, Dordrecht, Zuid-Holland, Netherlands) 3.0%, KCl 0.3%, CaCO3 0.2%, MgSO4·7H2O 0.05%, KH2PO4 0.05%, kanamycin 0.1 mg ml−1 and agar 1.5%, adjusted to pH 6.5 before sterilization.
Medium 3 contains non-glutinous brown rice (Cigario Co., Ltd., Tokyo, Japan) 3.0%, glucose 0.5%, glycerol 0.5%, yeast extract (Oriental Yeast, Tokyo, Japan) 0.2%, potato dextrose broth (Becton Dickenson, Sparks, MD, USA) 0.2%, ammonium acetate 0.2%, NaNO3 0.03%, KCl 0.005%, MgSO4·7H2O 0.005%, FeSO4·7H2O 0.0001%, ZnSO4·7H2O 0.0001%, CuSO4·5H2O 0.00005%, kanamycin 0.1 mg ml−1 and agar 1.5%, adjusted to pH 6.0 before sterilization.
Medium 4 contains sucrose 3.0%, soluble starch 3.0%, malt extract (BD) 1.0%, brewer’s yeast (Asahi Food and Healthcare, Tokyo, Japan) 0.3%, KH2PO4 0.5%, MgSO4·7H2O 0.05%, kanamycin 0.1 mg ml−1 and agar 1.5%, adjusted to pH 6.0 before sterilization.
Soil plating methods
Two soil (or sample) plating methods, A and B, were tested on each medium.
Plating method A: A soil sample (15 mg) was directly placed in a plastic plate (90 φ × 15 mm; Kanto Chemical, Tokyo, Japan). Each melting (at 45–50 °C) medium (15 ml, media 1–4) was poured into the plate and mixed well to disperse the sample. The soil plate was incubated at 27 °C for 2–3 days to form fungal colonies.
Plating method B: A soil sample (1.0 g) was suspended in Winogradsky buffer (10 ml) (K2HPO4 0.38%, KH2PO4 0.12%, MgSO4·7H2O 0.51%, NaCl 0.25%, Fe2(SO4)3·nH2O 0.005% and MnSO4·5H2O 0.005%) by shaking vigorously to make a series of dilutions (1:10–1:1000). Each diluted suspension (200 μl) was placed on the surface of each medium (15 ml, media 1–4 in a plastic plate) (90 φ × 15 mm) and spread evenly over the surface with a sterilized bacteria spreader. The soil plates were incubated at 27 °C for 2–3 days to form fungal colonies.
C. albicans-inoculating methods
C. albicans ATCC 64548 was used as a pathogenic fungus. A loopful of C. albicans from potato dextrose agar (BD) slant (7.5 ml, in a tube with screw cap, 9 × 16 × 150 mm3; Asahi Glass, Tokyo, Japan) was inoculated in Waksman medium (10 ml, peptone 0.5%, meat extract 0.5%, NaCl 0.3% and glucose 2.0%, adjusted to pH 7.0 before sterilization) and incubated at 27 °C for 2 days, reaching 1.0 × 109 CFU ml−1 concentration. This original culture was directly used as a C. albicans suspension. Five C. albicans-inoculating methods, I–V, were tested.
Inoculating method I: The original C. albicans culture (15 μl, 1.0 × 109 CFU ml−1) was directly added to each soil plate at the same time when they were prepared according to plating method A.
Inoculating method II: The original C. albicans culture (15 μl, 1.0 × 109 CFU ml−1) was directly placed in plastic plates (90 φ × 15 mm). Each melting (at 45–50 °C) medium (15 ml, media 1–4) was poured into the plate and mixed well to disperse the C. albicans. Soil samples were overlaid according to plating method B.
Inoculating method III: The original C. albicans culture was diluted with sterilized saline to adjust to 1.0 × 105 CFU ml−1. The suspension (100 μl) was placed on the surface of the soil plates previously prepared according to plating methods A and B, and spread evenly over the surface with a sterilized bacteria spreader.
Inoculating method IV: The original C. albicans culture was diluted with 0.8% agar at 45–50 °C to adjust to 1.0 × 104 CFU ml−1. Two milliliters of the diluted C. albicans suspension was overlaid on soil plates previously prepared according to plating methods A and B.
Inoculating method V: The original C. albicans culture was diluted with sterilized saline to adjust to 5–10 × 105 CFU ml−1. The suspension (40 μl) was overlaid by spraying from heights of about 150 mm onto soil plates prepared by plating methods A and B (Figure 4). The atomizer was purchased from Ryohin Keikaku (Tokyo, Japan). With one push of the atomizer, a 40-μl solution was sprayed. Inoculated plates were incubated at 27 °C for 1 day. Fungi showing antagonistic effects (inhibition zone) on the growth of C. albicans were selected.
Taxonomical identification of fungi
Taxonomical identification21, 22 of fungi was carried out by observing the precise arrangement of the conidiophores and the way in which spores were produced (conidial ontogeny) by fungi grown on potato dextrose agar and potato carrot agar23 (potato 2.0%, carrot 2.0% and agar 2.0%). For this, the slide culture method was adopted.24 Potato dextrose agar and potato carrot agar blocks (5 × 5 mm2) of the cultures were cut out aseptically, and placed on sterile microscope slides (26 × 76 mm2; Toshinriko, Tokyo, Japan). Four edges of the agar block were inoculated with fungal strains. Inoculated agar blocks were covered with a sterile micro cover glass (18 × 18 mm2; Matsunami Glass, Osaka, Japan) and incubated in moist chambers at 25 °C for 5 days. Cover slips were carefully removed from the agar blocks and mounted in a drop of Shear’s Mounting Medium (potassium acetate 1.0%, glycerol 20% and EtOH 30%) on a clean glass slide. The slide was sealed with transparent nail polish. The structure and branching pattern of conidiophores were observed under a Vanox-S AH-2 microscope (Olympus, Tokyo, Japan).
Fermentation
Fungi from soil samples were maintained on Miura’s medium (LCA) (glycerol 0.10%, KH2PO4 0.08%, K2HPO4 0.02%, MgSO4·7H2O 0.02%, KCl 0.02%, NaNO3 0.2%, yeast extract 0.02% and agar 1.5%, adjusted to pH 6.0 before sterilization). To confirm the antifungal antibiotics produced by selected strains, a loopful of strains from LCA slant was inoculated into a test tube containing one of the media (10 ml, media 1, 2, 3 or 4 without kanamycin and agar). The tube was incubated on a rotary shaker (300 r.p.m.) at 27 °C for 6 days. After treatment of the culture broth (10 ml) with EtOH (10 ml), the mixture was centrifuged to obtain the EtOH extracts. Anti-C. albicans activity was investigated by the agar diffusion method using paper disks.
Assay of anti-C. albicans activity
Anti-C. albicans activity was measured by the established agar diffusion method25, 26, 27 using paper disks (8 mm, Advantec, Tokyo, Japan) containing EtOH extracts of isolated fungi (60 μl). The original C. albicans culture (15 μl, 1.0 × 109 CFU ml−1) was directly placed on plastic plates (90 φ × 15 mm). Melting (at 45–50 °C) GY agar (15 ml, glucose 1.0%, yeast extract 0.5% and agar 0.8%, adjusted to pH 6.0 before sterilization) was poured into the plate and mixed well to disperse C. albicans. The C. albicans plate was incubated at 27 °C for 24 h.
Traditional isolation method
The traditional isolation method was carried out by using the established method.28 Soil plates were prepared by the same method (plating methods A and B) using other media. Soil-derived fungi, isolated from the soil plates after 2- or 3-day incubation at 27 °C, were re-cultured on LCA and YpSS (soluble starch 1.5%, yeast extract 0.4%, K2HPO4 0.1%, MgSO4·7H2O 0.05% and agar 2.0%, adjusted to pH 6.0 before sterilization) plated for 2 weeks. After duplication of the fungal strains, fungi were re-cultured in four liquid media (media 1–4) by using the same method as described above.
References
Pelaez, F. in Handbook of Industrial Mycology (ed. Zhiqiang An, ) Vol. 22, 49–92 (Marcel Dekker, New York, 2005).
Blackwell, M. The fungi: 1, 2, 3 … 5.1 million species? Am. J. Bot. 98, 426–438 (2011).
Hawksworth, D. L. The magnitude of fungal diversity: the 1.5 million species estimate revisited. Mycol. Res. 105, 1422–1432 (2001).
Bull, A. T., Goodfellow, M. & Slater, J. H. Biodiversity as a source of innovation in biotechnology. Annu. Rev. Microbiol. 46, 219–252 (1992).
Fleming, A. On the antibacterial action of cultures of a Penicillium, with special reference to their use in the isolation of B. influenzae. Brit. J. Exp. Pathol. 10, 226–236 (1929).
Rüegger, A. et al. [Cyclosporin A, a peptide metabolite from Trichoderma polysporum (Link ex Pers.) Rifai, with a remarkable immunosuppressive activity]. Helv. Chim. Acta 59, 1075–1092 (1976).
Endo, A., Kuroda, M. & Tsujita, Y. ML-236A, ML-236B, and ML-236C, new inhibitors of cholesterogenesis produced by Penicillium citrinium. J. Antibiot. 29, 1346–1348 (1976).
Endo, A. & Monacolin, K. A new hypocholesterolemic agent produced by a Monascus species. J. Antibiot 32, 852–854 (1979).
Alberts, A. W. et al. Mevinolin: a highly potent competitive inhibitor of hydroxymethylglutaryl-coenzyme A reductase and a cholesterol-lowering agent. Proc. Natl Acad. Sci. USA 77, 3957–3961 (1980).
Fleischmann, R. D. et al. Whole-genome random sequencing and assembly of Haemophilus influenzae Rd. Science 269, 496–512 (1995).
Relman, D. A. Microbial genomics and infectious diseases. N. Engl. J. Med. 365, 347–357 (2011).
Bentley, S. D. et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 417, 141–147 (2002).
Ohnishi, Y. et al. Genome sequence of the streptomycin-producing microorganism Streptomyces griseus IFO 13350. J. Bacteriol. 190, 4050–4060 (2008).
Omura, S. et al. Genome sequence of an industrial microorganism Streptomyces avermitilis: deducing the ability of producing secondary metabolites. Proc. Natl Acad. Sci. USA 98, 12215–12220 (2001).
Galagan, J. E. et al. Sequencing of Aspergillus nidulans and comparative analysis with A. fumigatus and A. oryzae. Nature 438, 1105–1115 (2005).
Payne, G. A. et al. Whole genome comparison of Aspergillus flavus and A. oryzae. Med. Mycol. 44, S9–S11 (2006).
Sanchez, J. F., Somoza, A. D., Keller, N. P. & Wang, C. C. Advances in Aspergillus secondary metabolite research in the post-genomic era. Nat. Prod. Pep. 29, 351–371 (2012).
Gross, H. Strategies to unravel the function of orphan biosynthesis pathways: recent examples and future prospects. Appl. Microbial. Biotechnol. 75, 267–277 (2007).
Brakhage, A. A. & Schroeckh, V. Fungal secondary metabolites - strategies to activate silent gene clusters. Fungal. Genet. Biol. 48, 15–22 (2011).
Kawaguchi, M. et al. A new ascochlorin derivative from Cylindrocarpon sp. FKI-4602. J. Antibiot. 66, 23–29 (2013).
von Arx, J. A. The Genera of Fungi Sporulating in Pure Culture 3rd edn. ( Verlag J Cramer: Vaduz, (1981) ).
Seifert, K., Morgan-Jones, G., Gams, W. & Kendrick, B. The Genera of Hyphomycetes. CBS Biodiversity Series 9 ( CBS-KNAW Fungal Biodiversity Centre: Utrecht,, (2011) ).
Atlas, R. M. Handbook of Microbiological Media 4th edn. ( CRC Press: Boca Raton, (2010) ).
Zalar, P. et al. Phylogeny and ecology of the ubiquitous saprobe Cladosporium sphaerospermum, with descriptions of seven new species from hypersaline environments. Stud. Mycol. 58, 157–183 (2007).
De Beer, E. J. & Sherwood, M. B. The paper-disc agar-plate method for the assay of antibiotic substances. J. Bacteriol. 50, 459–467 (1945).
Humphrey, J. H. & Lightbown., J. W. The general theory for plate assay of antibiotics with some practical applications. J. Gen. Microbiol. 7, 129–143 (1952).
Iwatsuki, M. et al. Lariatins, novel anti-mycobacterial peptides with a lasso structure, produced by Rhodococcus jostii K01-B0171. J. Antibiot. 60, 357–363 (2007).
Takada., M. in The Isolation Method of Microorganisms. (eds Yamasato K., Morichi T., Udagawa S., Kodama T.,) 31–43 ( R&D Planning: Tokyo, (2001) in Japanese.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kawaguchi, M., Nonaka, K., Masuma, R. et al. New method for isolating antibiotic-producing fungi. J Antibiot 66, 17–21 (2013). https://doi.org/10.1038/ja.2012.79
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2012.79
Keywords
This article is cited by
-
Microbial Nanoculture as an Artificial Microniche
Scientific Reports (2016)
-
Better visualization and photodocumentation of zone of inhibition by staining cells and background agar differently
The Journal of Antibiotics (2015)
-
A new ascochlorin derivative from Cylindrocarpon sp. FKI-4602
The Journal of Antibiotics (2013)