Abstract
A fraction of methicillin-resistant Staphylococcus aureus (MRSA) shows resistance to vancomycin (VCM) in the presence of β-lactam antibiotics (BIVR) at low concentrations. We hypothesized that the BIVR phenomenon might be exerted by a peptidoglycan derivative(s) generated as a consequence of β-lactam antibiotic action. To verify this hypothesis, we isolated the fraction that mimicked the effect of β-lactam antibiotics by the enzymatic treatment of the crude cell wall. The active components were purified by a combination of reverse phase chromatographies, mass spectrum and amino-acid analyses, and were identified to be a muropeptide with the following formula: N-acetyglucosamyl-N-acetylmuramyl--Ala-D-isoGln-L-Lys-(ɛ-NH-4Gly)-D-Ala-2Gly. This is the very first identification of the active component, which induces VCM resistance in MRSA. We found that the BIVR cells are highly sensitive to this compound rendering the cells resistant to VCM compared with non-BIVR MRSA.
Similar content being viewed by others
Introduction
Infection by methicillin-resistant Staphylococcus aureus (MRSA) is problematic in hospitals throughout the world, ranging from 60 to 80% of all S. aureus infections in Japan.1, 2 Patients who suffer from MRSA infection are often coinfected with Gram-negative bacteria, such as Pseudomonas aeruginosa.1 These patients are likely to be treated with a combination of vancomycin (VCM) and β-lactam antibiotics, such as carbapenem, because Gram-negative bacteria are intrinsically resistant to VCM. Accordingly, this combination therapy has been recommended and practiced for the treatment of MRSA infection in Japan for over 20 years.1, 3 The problem has arisen, however, that combination therapy with VCM and β-lactam antibiotics has caused the emergence of VCM-resistant MRSA,4, 5 designated as β-lactam antibiotic-induced VCM-resistant MRSA (BIVR).6, 7 The isolation frequency of BIVR in Japanese University hospitals has been reported to be about 6–12% among MRSA isolates.7, 8 In vitro studies have revealed that the BIVR phenomenon can be induced by any β-lactam antibiotic so far tested without exception.9
The mode of VCM action is understood to be as follows. VCM inhibits the peptidoglycan synthesis of Gram-positive bacteria by binding to the D-Ala-D-Ala terminals of newly synthesized peptidoglycan and of a membrane-bound murine monomer intermediate (C55-isoprenoid-PP-murein peptide or lipid-II).10, 11, 12 It has also been reported that a VCM-dimer crosslinks two D-Ala-D-Ala terminals of newly synthesized peptidoglycan and lipid-II.13 Thus, VCM blocks the access of penicillin-binding proteins to their substrate, consequently inhibiting the transpeptidation and transglycosylation reactions.
The accepted mode of β-lactam antibiotic action, based on many empirical studies, is that the antibiotics covalently bind with and inactivate penicillin-binding proteins, consequently inhibiting transpeptidation, but otherwise, β-lactam antibiotics are harmless to the intracellular metabolisms. Cells exposed to β-lactam antibiotics continue to biosynthesize macromolecules needed for cell division except for the peptidoglycan networks, and cells with the defective peptidoglycan network eventually rupture. Inhibition of peptidoglycan synthesis promotes the expression of autolysin, resulting in the release of large amounts of peptidoglycan fragments into the medium.14, 15
Three mechanisms of VCM resistance have been reported. (i) Alteration of the muropeptide structure such that peptidyl-D-Ala-D-Ala was modified to peptidyl-D-Ala-D-lactate or peptidyl-D-Ala-D-Ser by the products of the vanA, vanB, vanC, vanD and van-E genes, respectively.16 However, these genes were mainly found in Enterococcus species. VCM-resistant MRSA strains harboring vanA were reported in North America and designated as VCM-resistant S. aureus.17 However, vanA-positive BIVR has not yet been reported. (ii) VCM intermediate-resistant S. aureus has been reported and has shown elevated peptidoglycan synthesis and cell-wall thickening.18 The vanA gene is undetectable in VCM intermediate-resistant S. aureus-type cells. A mainly proposed mechanism for VCM resistance in VCM intermediate-resistant S. aureus cells is that the thickened peptidoglycan layers adsorb free VCM, thereby blocking the free passage of VCM through the peptidoglycan layers, thus, preventing the access of VCM to its target.19, 20 (iii) We reported recently that the BIVR cells rapidly remove the free VCM from medium in the presence of β-lactam antibiotics. When the medium VCM concentration was down to the critical level, growth of the BIVR cells resumed.21
This observation let us to hypothesize that the factor(s), which induces VCM resistance in the presence of β-lactam antibiotics, might be released into the medium and promotes the biosynthesis of peptideglycan precursor such as lipid II. We reported earlier that exogenously added UDP-N-acetylmuramyl-L-alanyl-D-isoglutamyl-L-lysine (UDP-MurNAc-L-Ala-D-isoGlu-L-Lys) induces the BIVR phenomenon.7 However, this compound is an intermediate precursor of peptidoglycan synthesis only present within the cells. Therefore, it is unlikely that this intracellular material induces the BIVR phenomenon. We assumed that unidentified peptidoglycan fragment(s) generated by the action of β-lactam antibiotics promotes the synthesis of lipid-II. A large amount of lipid-II translocated to the cell surface would trap unbound VCM, lower the free VCM concentration and restore cell growth.
This paper reports on the purification and identification of the peptideglycan fragment, which mimics the role of β-lactam antibiotics in the induction of the BIVR phenomenon.
Materials and methods
Bacterial strain, culture medium and antimicrobial agents
BIVR and non-BIVR strains were obtained from clinical sources by a modified method described previously,8 using Brain Heart Infusion (BHI) agar (Nippon Becton-Dickinson, Tokyo, Japan) in the presence of 4 μg ml−1 of VCM. The K744 strain isolated from the blood of a septicemia patient was used as a representative BIVR cell. MICs of VCM, teicoplanin, arbekacin, linezolid, ceftizoxime (CZX) and imipenem in K744 were 1, 1, 2, 2, 8192 and 128 μg ml−1, respectively. The K1179 strain obtained from the septicemia blood is a typical non-BIVR MRSA, which shows MICs of the above antibiotics as 1, 1, 1, 2, 512 and 16 μg ml−1, respectively. The FDA209P strain is a reference strain for methicillin-susceptible S. aureus (MSSA) and the MIC of the above antibiotics was 0.5, 0.5, ⩽0.25, 2, 4 and <0.25 μg ml−1, respectively. The culture media used in this study were nutrient broth, Mueller–Hinton broth, BHI broth (Nippon Becton-Dickinson) and BHI agar. VCM and CZX were purchased from Sigma-Aldrich (Saint Louis, MO, USA) and Astellas Pharma (Tokyo, Japan), respectively.
Determination of free peptidoglycan fragment(s)
The amount of peptidoglycan fragment in the culture supernatant was determent by silkworm larva plasma test (Wako Pure Chemical Industries, Osaka, Japan) according to manufacturer’s instruction.15 Bacterial cells were grown in nutrient broth overnight at 35 °C. The cells were harvested by centrifugation for 15 min at 3000 × g, washed twice with deionized water, and absorption at 578 nm was adjusted to 0.3 (about 108 c.f.u. ml−1) with fresh nutrient broth. The culture medium was supplemented with the following: (i) 1 μg ml−1 of CZX, (ii) 4 μg ml−1 of VCM, (iii) 1 μg ml−1 of CZX plus 4 μg ml−1 of VCM and incubated at 35 °C. An aliquot was withdrawn at the desired time, filtered through a membrane filter of 0.45 μm pore size. The filtrate was diluted a 100-fold by deionized water and mixed with an equal amount of silkworm larva plasma reagent. The absorption at 630 nm was recorded every 30 s at 30 °C with a plate reader (EL808 Ultra microplate reader; Biotek Instruments, Winooski, VT, USA). A calibration curve was drawn using reference peptidoglycan from Micrococcus luteus (Wako Pure Chemical Industries), and the curve appeared linear in a range of 0.08–1000 ng ml−1. At least three independent experiments were executed.
Preparation and purification of the cell-wall fragment
The experiment was conducted at 4 °C unless otherwise stated. An overnight culture of K744 cells was diluted with a 500-fold pre-warmed fresh BHI broth, and incubated overnight at 35 °C under aeration. The cells were harvested by centrifugation for 15 min at 3000 g, washed twice with deionized water and heated in a boiling water bath for 20 min. Insoluble materials were collected by centrifugation for 15 min at 3000 g and washed twice with deionized water. The suspension was then subjected to ultrasonic oscillation at 360 watts by a SONICATOR 3000 (Misonix, Farmingdale, NY, USA) in the presence of φ=0.4 mm glass beads for a total of 30 cycles by 20-s exposure and 30-s intermittent cooling in icy water. The ultrasonic-treated material was centrifuged for 15 min at 3000 g at 24 °C, then the pellets were suspended in 100 mM Tris-HCl (pH 8.0) and treated with 0.2 mg ml−1 of trypsin (Sigma-Aldrich) overnight at 37 °C. This material was heated in a boiling water bath for 5 min to inactivate trypsin, and the insoluble materials were washed twice with deionized water by centrifugation for 15 min at 3000 g. The insoluble material in 100 mM Tris-HCl (pH 7.0) was treated with 20% of hydrofluoric acid for 18 h at 4 °C to any liberate bound form of teichoic acid (cell-wall fraction) and successively treated with 4 μg ml−1 of mutanolysin (from Streptomyces globisporus, Sigma-Aldrich) and 4 μg ml−1 lysostaphin (Wako Pure Chemical Industries) at 37 °C for 24 h to release cell-wall components. The mixture was heated in a boiling water bath for 5 min, centrifuged at 14 000 g for 20 min and the soluble fraction was lyophilized. This material was dissolved in a small amount of distilled water, and applied to an octadecylsilyl column (15 × 30 mm, Senshu Scientific, Tokyo, Japan) activated with acetonitrile and equilibrated with deionized water. The column was washed with 20 ml of deionized water, and eluted stepwise with 20 ml of 5 and 10% methanol and the fractions were lyophilized (peptidoglycan fragment). The 10% methanol fraction was dissolved in 3 ml of distilled water, and a 5 μl aliquot was applied to the analytical HPLC column, PEGASIL C18 (4.6 × 250 mm, Senshu Scientific). The column was eluted at the rate of 0.7 ml min−1 with a solution containing 13% methanol and 50 mM sodium phosphate buffer (pH 2.5) and absorption at 206 nm was recorded. The remaining aliquot (2.995 ml) was applied to a preparative PEGASIL C18 column (20 × 250 mm, Senshu Scientific), which was then eluted at the rate of 7 ml min−1 under the same protocol as described above. The pH of the fractions was adjusted to neutrality with 0.5 M sodium hydroxide, the residual methanol was evaporated, and then the fractions were lyophilized. This material was dissolved in a small amount of deionized water and applied to a SepPak C18 cartridge (Waters, Milford, MA, USA) desalting column equilibrated with deionized water, and after washing with 10 ml of deionized water, the column was eluted with 10 ml of 20% methanol. The materials were lyophilized after evaporation of the residual methanol (muropeptide).
1H NMR analysis
NMR spectra were measured on the Inova 600 spectrometer (Varian, Palo Alto, CA, USA) with 1H NMR at 600 MHz in deuterium oxide. The chemical shifts were expressed in p.p.m. and are referred to as deuterium oxide (4.76 p.p.m.) in the 1H-NMR spectra.
Mass spectrometric analysis
Electro-spray ionization time of flight mass spectrometry was carried out with the JMS T100LP mass spectrometer (JEOL, Tokyo, Japan) according to manufacturer's instructions. The reference used was 0.2% sodium trifluoro acetate in methanol.
Amino-acid analysis
The sample was hydrolyzed in the gas phase of 6 N hydrochloride (198 μl) and phenol (2.0 μl) at 110 °C for 18 h in a reaction vial of which the gas-phase was replaced with an N2 gas using a Pico-Tag workstation (Waters).
The absolute configuration and quantities of amino acids were determined by the SUMICHIRAL OA5000 HPLC column, 4.6 × 150 mm, (Sumitomo Chemical, Tokyo, Japan) according to manufacturer's manual. To configure and quantify D-Lys, L-Lys, Gly, D-Ala and L-Ala, the column was eluted with 2 mM copper sulfate at the rate of 1.0 ml min−1 at 40 °C and A254 nm was recorded. The retention times of L-Lys, D-Lys, Gly, L-Ala and D-Ala were 2.46, 3.00, 4.16, 4.82 and 6.56 min, respectively. For D-Gln and L-Gln, the column was eluted with a solution of 15% methanol in 2 mM copper sulfate at the rate of 1.0 ml min−1 at 25 °C and A254 nm was recorded. The retention times of L-Gln and D-Gln were 19.10 and 21.15 min, respectively.
The aminopeptidase M treatment
The purified muropeptide (50 μg) was dissolved in 100 μl of 20 mM Tris-HCl (pH 7.2), treated with 50 μl (2 U ml−1) of aminopeptidase M in the same buffer (Takara Bio, Shiga, Japan) and incubated at 37 °C overnight. The reaction was terminated by adding 850 μl of methanol and the insoluble material was removed by centrifugation for 10 min at 10 000 g. The soluble materials were subjected to the electro-spray ionization time of flight mass spectrometry analysis.
Induction of the BIVR phenomenon on the agar plate
K744 cells were grown in Mueller–Hinton broth at 35 °C for 18 h. Absorption at 578 nm was adjusted to 0.3 (about 108 c.f.u. ml−1) and a 100-μl aliquot was streaked on BHI agar containing 3 μg ml−1 of VCM. Paper discs with the diameter of 8 mm were impregnated with an appropriate amount of CZX, highly purified muropeptide, and then placed on the above-described agar plate. The plates were incubated at 35 °C for 22 h.
Assay for the reversal of VCM-mediated growth inhibition in the liquid medium
K744 cells were grown in Mueller–Hinton broth at 35 °C for 18 h, and the absorption at 578 nm was adjusted to 0.03 (about 107 c.f.u. ml−1) with fresh BHI broth. The culture medium was supplemented with the following materials: (i) 4 μg ml−1 of VCM, (ii) 4 μg ml−1 of VCM plus 1 μg ml−1 of CZX and (iii) 4 μg ml−1 of VCM plus various amounts of purified muropeptide. Tubes were aerated by rocking at 20 strokes per min at 35 °C, and the cell density was monitored every 30 min at 578 nm by use of the TVS062 Bio-Photorecorder (Advantec, Tokyo, Japan).
Results
Peptidoglycan fragments released into the medium
Hypothesizing that peptidoglycan fragments were released into the culture medium in the presence of β-lactam antibiotics and VCM, we first determined the amount of peptidoglycan in the culture supernatant. In the presence of 4 μg ml−1 of VCM alone, the FDA209P, K1179 and K744 cells released 0.23, 0.44 and 2.24 μg ml−1 of peptidoglycan fragments into medium, respectively, at 6 h of the culture (Figure 1). When 1 μg ml−1 of CZX was supplemented in addition to VCM, the amount of peptideglycan fragments in the culture supernatant at 6 h appeared to be 11.4, 0.9 and 1.3 μg ml−1 in the BIVR (K744), non-BIVR MRSA (K1179) and MSSA (FDA209P), respectively. These results revealed that the BIVR cells released the peptidoglycan about 10 times more than the MRSA and MSSA cells. Experiments were carried out in the presence of 1 μg ml−1 of CZX only. The results showed that MSSA, non-BIVR MRSA and BIVR produced 55.0, 85.0 and 40.9 μg ml−1 of peptidoglycan fragments. However, the interpretation of the results was difficult because the MIC of CZX in the FDA209P, K1179 and K744 were extremely diverse, such as 4, 512 and 8192 μg ml−1, respectively. Thus, it was firmly established that all the Staphylococcus strains tested so far produced the peptidoglycan fragments in the presence of β-lactam antibiotic and the BIVR cells produce largest amount of extracellular peptidoglycan fragment in the presence of both CZX and VCM.
Release of peptidoglycan from the cells into medium. S. aureus cells (108 c.f.u. ml−1) were cultured in nutrient broth (NB) at 35 °C and then sampled every 2 h. NB was supplemented with either 4 μg ml−1 of VCM, 1 μg ml−1 of CZX plus 4 μg ml−1 of VCM. The supernatant fraction was filtrated through a 0.45 μm membrane filter. Then measured amount of peptideglycan by the silkworm larva plasma test. X axis is the sampling time (h). Y axis is the amount of peptidoglycan released (μg ml−1). Symbols:▵, 4 μg ml−1 of VCM (FDA209P); □, 4 μg ml−1 of VCM (K1179); ◊, 4 μg ml−1 of VCM (K744); ▴, 1 μg ml−1 of CZX plus 4 μg ml−1 of VCM (FDA209P); ▪, 1 μg ml−1 of CZX plus 4 μg ml−1 of VCM (K1179); ♦, 1 μg ml−1 of CZX plus 4 μg ml−1 of VCM (K744).
Purification and identification of an active compound(s) that induces VCM resistance in BIVR cells
The above results suggested a possibility that peptidoglycan fragments released from the cells is likely to elicit the BIVR phenomenon. Accordingly, we attempted to purify the active component from enzymatically degraded peptidoglycan fragments.
K744 cells were disintegrated by ultrasonic oscillation, and the insoluble materials obtained by centrifugation were treated successively with trypsin and hydrofluoric acid to degrade the proteins and to liberate the bound form of teichoic acids, respectively. Crude peptidoglycan fragments were released into the medium by lysostaphin and mutanolysin treatments of the crude cell-wall fraction. These materials were subjected to the PEGASIL C18 HPLC. The final elution profile shown in Figure 2a revealed two prominent peaks. Rechromatography of peak-1 yielded two peaks at the peak-1 and peak-2 positions, and that of peak-2 yielded two peaks at the peak-1 and peak-2 positions. These results suggested that peaks 1 and 2 are likely anomers of the same compound. NMR analysis revealed the anomeric proton in the compounds at peaks 1 and 2. The ESI-TOF mass spectrum of peaks-1/2 revealed four distinct peaks between 1000 and 1500 mass/charge-Na+ values at 1259, 1202, 1145 and 1088 (Figure 2b), corresponding to the mass values of N-acetylglucosaminyl(GlcNAc)-N-acetylmuraminyl(MurNAc)-L-Ala-D-isoGln-L-Lys-D-Ala+6Gly+Na, (muropeptide), GlcNAc-MurNAc-L-Ala-D-isoGln-L-Lys-D-Ala+5Gly+Na, GlcNAc-MurNAc-L-Ala-D-isoGln-L-Lys-D-Ala+4Gly+Na and GlcNAc-MurNAc-L-Ala-D-isoGln-L-Lys-D-Ala+3Gly+Na, respectively. The muropeptides with 5-, 4- and 3-glycine molecules were the products appearing during ESI-TOF mass analysis. Therefore, these compounds were not present in peaks-1/2. Similarly, peaks at 853, 796, 739, 682 and 429 mass/charge were detected and the mass/charge-Na+ values corresponded to the mass value of D-lactate-L-Ala-D-isoGln-L-Lys-D-Ala+6Gly+Na, D-lactate-L-Ala-D-isoGln-L-Lys-D-Ala+5Gly+Na, D-lactate-L-Ala-D-isoGln-L-Lys-D-Ala+4Gly+Na, D-lactate-Ala-isoGln-Lys-Ala+3Gly+Na and GlcNAc-MurNAc (lacking lactate)+Na, respectively. These molecules were the products formed when GlcNAc-MurNAc was split from the peptide moiety. The molecular formula of peaks 1/2 was determined by HR-electro-spray ionization time of flight mass spectrometry to be C48H80N14O24 (calculated, for [M–H]−: m/z 1235.53916, found 1235.53858). The relative amount of L-Ala, D-Glu, L-Lys, D-Ala and Gly appeared to be 1.1 : 1 : 1.1 : 0.8 : 7.1, respectively, as normalized to D-Glu=1.0. Thus, the molar ratio of L-Ala, D-isoGln, L-Lys, D-Ala and Gly was determined to be 1 : 1 : 1 : 1 : 6.
Purification and identification of the active component by HPLC and ESI-TOF mass. (a) The PEGASIL C18 HPLC profile of the crude peptidoglycan fragment. Details are given in the Materials and methods section. (b) ESI-TOF mass spectrum of the muropeptide from peaks-1/2. m/z, represents the mass/charge value.
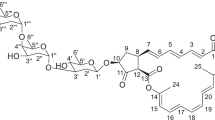
The process of linkage of six glycine molecules is still not clear. Assuming that some of the glycine molecules are linked to the ɛ-amino terminal of L-lysine molecule, the above muropeptide was treated with aminopeptidase M and the product was again analyzed by ESI-TOF mass. The result showed a single mass/charge peak with the mass value of 1009, which corresponded to the value of GlcNAc-MurNAc-L-Ala-D-isoGln-L-Lys-D-Ala+2Gly (data not shown). The result suggested that four glycine molecules linked to ɛ-amino group of the L-lysine were cleaved off by the aminopeptidase and two glycine molecules linked to carboxylterminal end of D-alanine was kept intact. On the basis of these data, we concluded that peaks-1/2 contained only one species of peptidoglycan fragment with the following formula, GlcNAc-MurNAc-L-Ala-D-isoGln-L-Lys-(ɛ-amino-4Gly)-D-Ala-2Gly. Figure 3 shows the chemical structure of the identified compound.
Chemical structure of the active component is identified based on the ESI-TOF mass and amino-acid analyses. Numbers represent mass values of the respective parts of the muropeptide. 1259, whole muropeptide+Na; 1202, muropeptide lacking 1Gly+Na; 1145, muropeptide lacking 2Gly+Na; 1088, muropeptide lacking 3Gly+Na; 853, whole peptide moiety+lactate+Na; 429, GlcNAc-MurNAc lacking lactate+Na; 204, GlcNAc lacking hydroxyl.
Effect of purified cell-wall fragment (muropeptide) on the induction of the BIVR phenomenon
Peaks-1/2 contained muropeptides as demonstrated in the above-described experiments, yet whether or not these components mimicked the role of β-lactam antibiotics in the induction of the BIVR phenomenon was not established. Therefore, we carried out growth experiments with the BIVR cells, K744, in the presence of various concentrations of muropeptide and 4 μg ml−1 of VCM. When approximately 107 cells were inoculated in the absence and presence of 4 μg ml−1 of VCM, the former culture soon started to grow and reached A578 ∼2 in about 5 h of incubation, but the latter culture took about 42 h (Figure 4). The β-lactam antibiotic, CZX, at 1 μg ml−1 shifted the VCM alone curve to about 13.5 h earlier, thereby confirming previous results.21
Induction of VCM resistance by the purified muropeptide in broth culture. A preculture of the K744 strain grown overnight was diluted with fresh BHI broth, adjusting the cell number to 107 c.f.u. ml−1 (A578=0.03) and the growth of the cells was monitored spectrophotometrically at A578 at 30 min intervals. BHI broth was supplemented with either 4 μg ml−1 of VCM, 1 μg ml−1 of CZX plus 4 μg ml−1 of VCM or 3.13∼100 μg ml−1 of purified muropeptide plus 4 μg ml−1 of VCM. Symbols: •, drug free; ♦, 4 μg ml−1 of VCM; ▪, 4 μg ml−1 of VCM plus 1 μg ml−1 of CZX; ○, 100 μg ml−1 of muropeptide plus 4 μg ml−1 of VCM; □, 50 μg ml−1 of muropeptide plus 4 μg ml−1 of VCM; ◊, 25 μg ml−1 of muropeptide plus 4 μg ml−1 of VCM; ▵, 12.5 μg ml−1 of muropeptide plus 4 μg ml−1 of VCM;  , 6.25 μg ml−1 of muropeptide plus 4 μg ml−1 of VCM; ▿, 3.13 μg ml−1 of muropeptide plus 4 μg ml−1 of VCM.
, 6.25 μg ml−1 of muropeptide plus 4 μg ml−1 of VCM; ▿, 3.13 μg ml−1 of muropeptide plus 4 μg ml−1 of VCM.
Muropeptide having the structure of GlcNAc-MurNAc-L-Ala-D-isoGln-L-Lys-(ɛ-amino-4Gly)-D-Ala-2Gly at concentrations of 100 μg ml−1 down to 3.13 μg ml−1 was added to the medium containing 4 μg ml−1 of VCM and the culture was monitored. When 100 and 50 μg ml−1 of the muropeptides were added, the growth curves reached A578 ∼2 in about 18 and 25.5 h, respectively. Growth of the cells in the presence of 100 and 50 μg ml−1 of muropeptide appeared 24 and 16.5 h earlier than that containing 4 μg ml−1 of VCM only. Purified muropeptide alone exerted no detectable effect on the cell growth. The effect of these concentrations of muropeptides on promoting the BIVR phenomenon was much stronger than that of 1 μg ml−1 of CZX. As the muropeptide concentrations were lowered to 25, 12.5 and 6.25 μg ml−1, the effect of the muropeptides diminished accordingly, appearing less effective than that of 1 μg ml−1 of CZX, yet the cell growth appeared much faster than that in the muropeptide-free VCM culture. Muropeptide at the concentration of 3.13 μg ml−1 was not effective showing a growth curve close to that in the VCM-only curve. These results clearly demonstrated that purified muropeptide promoted VCM resistance by mimicking the effect of β-lactam antibiotics.
The effect of purified muropeptide on induction of the BIVR phenomenon was tested using various types of S. aureus on agar plates. The paper discs contained muropeptides in the amounts of 75, 37.5, 18.7, 9.3 and 0 μg. A hollow growth zone around each paper disc was measured. K744 cells showed hollow diameters of 17.5, 14.2, 11.8 and 8.6 mm at muropeptide concentration 75, 37.5, 18.7 and 9.3 μg per disc, respectively (Figure 5). The results confirmed the data of the broth assay shown above. The growth zone of the K1179 cells was 10.9 and 9.2 mm in the presence of 75 and 37.5 μg per disc, respectively, of the muropeptide, which was undetectable at 18.7 μg per disc. Similarly, FDA2097 cells showed 10.5 and 8.2 mm growth zones in the presence of 75 and 37.5 μg per disc, respectively, of the muropeptide. These results revealed that the BIVR phenomenon was not only limited to BIVR cells, but could be induced even in non-BIVR MRSA and MSSA. The only apparent difference was that the BIVR cell was more susceptible to the muropeptide than non-BIVR MRSA and MSSA.
Induction of VCM resistance by the purified muropeptide on agar plate. The K744 cell suspension (100 μl of 108 c.f.u. ml−1) was streaked on BHI agar containing 3 μg ml−1 of VCM. The paper discs impregnated with 75 μl of muropeptide solution were placed on the agar and the plate was then incubated at 35 °C for 22 h. Zone of growth around the disc was measured. Symbols: ▴, the K744 cells; ▪, the K1179 cells; •, the FDA209P cells.
Discussion
This study was conducted to search for an active component that could replace β-lactam antibiotics in the induction of the BIVR phenomenon. The released peptidoglycan fragments in the presence of 1 μg ml−1 of CZX only appeared to be 55.0, 85.0 and 40.9 μg ml−1 in the FDA209P, K1179 and K744 cells, respectively, and that was lowered to 1.3, 0.9 and 11.4 μg ml−1, respectively, in the presence of CZX and VCM. Low-level production of peptidoglycan fragment in the presence of VCM in addition to CZX may be due to that the CZX-mediated activation of peptidoglycan metabolism was inhibited by VCM. Yet, the level of inhibition was the smallest in the K744 cells. We hypothesized that the released peptidoglycan was recycled and that promoted the cell-wall synthesis. In this experiment, we purified and identified the active component from the enzymatically degraded cell-wall components of the K744 cells. The chemical structure of the active component eventually identified was GlcNAc-MurNAc-L-Ala-D-isoGln-L-Lys-(ɛ-amino-4Gly)-D-Ala-2Gly. The number of glycine molecules identified in the compound was six with a total mass value of 1236. The structure of this compound coincided with the previously reported muropeptide molecule,22 yet its biological activity was not reported. As the ɛ-amino-4Gly moiety was cleaved off with aminopeptidase M, the product obtained had the structure of GlcNAc-MurNAc-L-Ala-D-isoGln-L-Lys-D-Ala-2Gly. The BIVR phenomenon inducing activity of this compound was comparable with that of the compound having ɛ-amino-4Gly. Therefore, ɛ-amino-4Gly is not essential for the activity. Although the identified compound had the mass value of 1009, whether or not this compound is an optimum structure for inducing the BIVR phenomenon remains unclear. This is the first report that identified the BIVR-inducible compound. The activity of the muropeptide identified in this study seems to be roughly comparable with that in UDP-MurNAc-L-Ala-D-isoGlu-L-Lys. It must be noted that the active component purified from enzymatically degraded peptidoglycan of the K744 cells may not be unique to the K744 cells but also present in most, if not all, S. aureus peptidoglycans. Thus, it is likely that all S. aureus produce the similar compound in the presence of β-lactam antibiotics as described above. The response to such compound is strongest in BIVR cells compared with that in non-BIVR MRSA and MSSA cells.
The question to be asked first would be whether or not these compounds can bind directly to the VCM molecule and activate it. As the muropeptides identified had no free D-Ala-D-Ala structure, this prediction is extremely unlikely. Nevertheless, we tested the binding and inactivation of VCM with purified muropeptide and found no evidence to support this hypothesis. The second question to be asked would be how the muropeptide acts to interfere with the antibiotic activity of VCM. We demonstrated very recently that free VCM in the BIVR culture medium rapidly decreased in the presence of β-lactam antibiotics. We assumed that muropeptides with a similar or identical structure to that investigated in this study were released into the medium as the β-lactam antibiotics exerted their effect on the cells. As the muropeptide identified in this study did not bind directly to VCM, it is most likely that the peptide induced a compound having a terminal D-Ala-D-Ala structure to which VCM could bind. It is still premature to speculate the mechanism by which the muropeptide acts in the induction of the BIVR phenomenon. However, it is of great interest to predict the possible mechanism(s), as the results may promote the advancement of this field of science.
We have assumed that muropeptides with a similar or identical structure to that investigated in this study may be released into the culture medium by the action of autolysin induced by the action of β-lactam antibiotics.14, 15 The muropeptide may be actively transported into the cell as the signal of the β-lactam-mediated peptidoglycan damage and activates the synthesis of lipid-II, a course that is reminiscent of Escherichia coli peptidoglycan recycling and β-lactamase induction.23, 24 The large amount of lipid-II exported to the cell surface traps VCM and reduces the free VCM in the medium promoting the growth of the cells. It is still necessary to identify genuine active component(s), which promotes the BIVR phenomenon from the culture supernatant of the BIVR cells. Such study is on progress in this laboratory.
References
Shimizu, K. et al. Clinical studies on vancomycin in the treatment of MRSA infection. Jpn. J. Antibiot. 49, 782–799 (1996).
Sunada, A. & Asari, S. Report of questionnaire survey for methicillin-resistant Staphylococcus aureus and penisillin-resistant Streptococcus pneumoniae between 1998 and 2000 in the Kinki district (article in Japanease). Kansenshogaku. Zasshi. 77, 331–339 (2003).
Totsuka, K., Shiseki, M., Kikuchi, K. & Matsui, Y. Combined effects of vancomycin and imipenem against methicillin-resistant Staphylococcus aureus (MRSA) in vitro and in vivo. J. Antimicrob. Chemother. 44, 455–460 (1999).
Haraga, I. et al. Emergence of vancomycin resistance during therapy against methicillin-resistant Staphylococcus aureus in a burn patient—importance of low-level resistance to vancomycin. Intl. J. Infect. 6, 302–308 (2002).
Takayama, Y. et al. Investigation of methicillin-resistant Staphylococcus aureus showing reduced vancomycin susceptibility isolated from a patient with infective endocarditis. Intl. J. Antimicrob. Agents. 22, 567–573 (2003).
Hanaki, H., Yamaguchi, Y., Barada, K., Sakai, H. & Sunakawa, K. Improved method of detection of β-lactam antibiotic-induced VCM-resistant MRSA (BIVR). Intl. J. Antimicrob. Agents. 23, 311–313 (2004).
Hanaki, H. et al. Investigation of β-lactam antibiotic-induced vancomycin-resistant MRSA (BIVR). J. Infect. Chemother. 11, 104–106 (2005).
Hanaki, H. et al. Method of detecting β-lactam antibiotic induced vancomycin resistant MRSA (BIVR). Intl. J. Antimicrob. Agents. 23, 1–5 (2004).
Hanaki, H., Inaba, Y., Sasaki, K. & Hiramatsu, H. A novel method of detecting Staphylococcus aureus heterogeneously resistant to vancomycin (Hetero-VRSA). Jpn. J. Antibiot. 51, 521–530 (1998).
Barna, J C. & Williams, D H. The structure and model of action of glycopeptide antibiotics of the vancomycin group. Annu. Rev. Microbiol. 38, 339–357 (1984).
Perkins, H R. Specificity of combination between mucopeptide precursors and vancomycin or ristocetin. Biochem. J. 111, 195–205 (1969).
Perkins, H R. & Nieto, M. The chemical basis for the action of the vancomycin group of antibiotics. Ann. NY. Acad. Sci. 235, 348–363 (1974).
Beauregard, D A., Williams, D H., Gwynn, M N. & Knoweles, DJ. Dimerization and membrane anchors in extracellular targeting of vancomycin group antibiotics. Antimicrob. Agents. Chemother. 39, 781–785 (1995).
Tuomanen, E., Cozens, R., Tosch, W., Zak, O. & Tomasz, A. The rate of killing of Escherichia coli by β-lactam antibiotics is strictly proportional to the rate of bacterial growth. J. Gen. Microbiol. 132, 1297–1304 (1986).
van Langevelde, P. et al. Antibiotic-induced release of lipoteichoic acid and peptidoglycan from Staphylococcus aureus: quantitative measurements and biological reactivities. Antimicrob. Agents. Chemother. 42, 3073–3078 (1998).
Courvalin, P. Vancomycin resistance in Gram-positive cocci. Clin Infect Dis 42 (Suppl 1), S25–S34 (2006).
Sievert, D M., Rudrik, J T., Patel, J B., McDonald, LC. & Wilkins, MJ. Vancomycin-resistant Staphylococcus aureus in the United States, 2002-2006. Clin Infect Dis 46, 668–674 (2008).
Hanaki, H. et al. Activated cell-wall synthesis is associated with vancomycin resistance in methicillin-resistant Staphylococcus aureus clinical strains Mu3 and Mu50. J. Antimicrob. Chemother. 42, 199–209 (1998).
Cui, L. et al. Novel mechanism of antibiotic resistance originating in vancomycin-intermediate Staphylococcus aureus. Antimicrob. Agents. Chemother. 50, 428–438 (2006).
Cui, L. et al. Cell wall thickening is a common feature of vancomycin resistance in Staphylococcus aureus. J. Clin. Microbiol. 41, 5–14 (2003).
Yanagisawa, C. et al. Rapid depletion of free vancomycin in medium in the presence of β-lactam antibiotics and growth restoration in Staphylococcus aureus strain with β-lactam-induced vancomycin resistance. Antimicrob. Agents. Chemother. 53, 63–68 (2009).
de Jonge, B L., Chang, Y S., Gage, D. & Tomasz, A. Peptidoglycan composition of a highly methicillin-resistant Staphylococcus aureus strain. J. Biol. Chem. 267, 11248–11254 (1992).
Jacobs, C., Huang, L., Bartowsky, E., Normark, S. & Park, JT. Bacterial cell wall recycling provides cytosolic muropeptides as effectors for β-lactamase induction. EMBO J. 13, 4684–4694 (1994).
Jacobs, C. et al. AmpD, essential for broth β-lactamase regulation and cell wall recycling, is a novel cytosolic N-acetylmuramyl-L-alanine amidase. Mol. Microbiol. 15, 533–539 (1995).
Acknowledgements
We are grateful to Dr K Nagai, Ms A Nakagawa and Ms N Sato, School of Pharmacy, Kitasato University, for their assistance in the mass and NMR analyses. This work was supported by grants-in-aid from the Food Safety Commission, Japan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ikeda, S., Hanaki, H., Yanagisawa, C. et al. Identification of the active component that induces vancomycin resistance in MRSA. J Antibiot 63, 533–538 (2010). https://doi.org/10.1038/ja.2010.75
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2010.75