Abstract
Manipulation of pathway regulation is an efficient strategy to increase specific secondary metabolite production. In this study, we successfully improved the production of both the enediyne antitumor antibiotic C-1027 and a heptaene, an early metabolite of the C-1027 pathway, by manipulating the three regulatory genes, sgcR1, sgcR2 and sgcR3, within the C-1027 biosynthetic gene cluster. SgcR3 has previously been established as an activator, and we now propose that SgcR1 and SgcR2 are also positive regulators based on their upregulation effects on titer and/or timing of heptaene and C-1027 production in Streptomyces globisporus. Specifically, overexpression of sgcR1 significantly improved the production of heptaene (about fivefold) and C-1027 (two- to threefold) compared with the wild-type strain. However, the titers of heptaene and C-1027 were not increased by overexpressing all the three activators together, underscoring the complexity of C-1027 biosynthetic pathway regulation. The possibility of exploiting the heptaene as a readily identifiable and unique indicator for rapidly detecting enediyne production was also assessed.
Similar content being viewed by others
Introduction
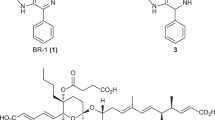
The enediyne family of natural products shares a characteristic unsaturated core containing two acetylenic groups conjugated to a double bond within either a 9- or 10-membered ring.1, 2 Benefiting from their extreme cytotoxicity, the 9-membered enediyne neocarzinostatin and the 10-membered enediyne calicheamicin have already been developed into anticancer drugs,2 and another 9-membered enediyne C-1027 (Figure 1), which shows even higher cytotoxicity than the former two, is currently in phase II clinical trials.3
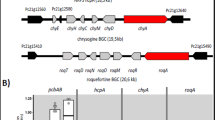
Similar to most of the nine-membered enediynes, C-1027 is a chromoprotein antibiotic consisting of an enediyne chromophore and an apoprotein.1 As its biosynthetic gene cluster was cloned from Streptomyces globisporus in 2002,4, 5 C-1027 biosynthesis has been extensively studied,6, 7, 8 and the unique enediyne core structure was proposed to be made by an iterative type I enediyne polyketide synthase (PKSE) SgcE together with several accessory enzymes.5 When SgcE and its cognate thioesterase SgcE10 were coexpressed in different hosts, a 15-carbon heptaene was produced (Figure 1).9 Significantly, this heptaene was produced by all combinations of PKSEs and thioesterases from five cloned enediyne clusters,10, 11 and has been found in all four examined 9- or 10-membered enediyne wild-type producers, suggesting that the heptaene may be a universal indicator of enediyne production.11
Improvement of specific secondary metabolite titers by manipulating biosynthetic pathway regulatory machinery has been demonstrated by numerous successful examples.12, 13 In this study, we set out to manipulate C-1027 pathway regulation for two purposes: (1) to improve C-1027 production, as the low titer has hampered both mechanistic studies and clinical development, as well as the efforts toward understanding C-1027 biosynthesis, and (2) to compare the production of C-1027 and heptaene in different mutant strains and therefore to assess the utility of heptaene as a phenotypic reporter of enediyne production. Among the three proposed regulators (SgcR1, SgcR2 and SgcR3) identified within the C-1027 biosynthetic gene cluster, only SgcR3 has been characterized as an activator; C-1027 titer can be modestly enhanced (about 30%) by overexpressing sgcR3.14 The other two uncharacterized regulators SgcR1 and SgcR2 provide opportunities to further improve C-1027 production.
In this paper, we report that both SgcR1 and SgcR2 are positive regulators, and significant improvements of both C-1027 (two- to threefold) and heptaene (approximately fivefold) production were achieved by sgcR1 overexpression. Moreover, the titer correlation between C-1027 and heptaene in selected S. globisporus wild-type and mutant strains suggested that the latter can be used as a convenient phenotypic indicator for enediyne production.
Materials and methods
Bacterial strains and plasmids
S. globisporus wild-type and mutant strains were grown on ISP4 agar medium or cultured in A9 liquid medium as described.15 Escherichia coli DH5α and ET1256716 were cultured in Luria-Bertani broth.17 Micrococcus luteus ATCC 9431 was grown on Luria-Bertani agar.17 Cosmid pBS10064 and plasmid pWHM125018 were previously described. Vector pGEM-T Easy was purchased from Promega (Madison, WI, USA).
DNA manipulations
General DNA manipulations were carried out according to standard procedures.17 PCRs were performed using Pfx DNA polymerase according to the manufacturer’s instructions (Invitrogen, Carlsbad, CA, USA). The PCR fragment obtained using Pfx was treated with Taq DNA polymerase from Invitrogen for 10 min at 72 °C to add the terminal ‘A’ overhangs. Streptomyces protoplasts preparation and transformation were performed as previously described.19 Homologous sequence database searching was executed with BLAST.
Overexpression of sgcR1, sgcR2 and sgcR3 in wild-type S. globisporus
Cosmid pBS1006 was used as a template to clone the three regulators for C-1027 production by PCR. The sgcR1 gene was amplified by PCR using primers 5′-CGAGGATCCGCGGATGGATTACACG-3′ (sgcR1 upstream, BamHI) and 5′-CCTCTAGAAGTAGGCCGTCAACGGC-3′ (sgcR1 downstream, XbaI), and was inserted into pGEM-T Easy after adding terminal ‘A’ overhangs by Taq DNA polymerase. After confirming its identity by DNA sequencing, the sgcR1 gene was excised as a 1.1-kb BamHI/XbaI fragment and inserted into the same sites of pWHM1250 to generate pBS1107, in which the expression of sgcR1 is under the control of the constitutive ErmE* promoter. Similarly, the sgcR2 gene was amplified by PCR using primers 5′-CTTCTAGAGGGAGTTACGGTGACCACG-3′ (sgcR2 upstream, XbaI) and 5′-TTCTGCAGGGCCCTGGTCACCC-3′ (sgcR2 downstream, PstI), and was inserted into pGEM-T Easy after adding terminal ‘A’ overhangs by Taq DNA polymerase; its identity was confirmed by DNA sequencing. Plasmid pBS1097 with sgcR2 under the control of ErmE* was then constructed by inserting the 0.9-kb PstI/XbaI sgcR2 fragment into the same sites of pWHM1250. For sgcR3 overexpression, the sgcR3 gene was amplified by PCR using primers 5′-CGAAAGCTTGCCCAGGAGAAGGGAAAC-3′ (SgcR3 upstream, HindIII) and 5′-GGGACGCGTCGCTCAGCTCC-3′ (SgcR3 downstream, MluI), inserted into pGEM-T Easy after adding terminal ‘A’ overhangs by Taq DNA polymerase, and confirmed its identity by DNA sequencing. After removal of the terminator between the ErmE* promoter and the HindIII site in pWHM1250 as a 0.5-kb SphI fragment, the sgcR3 gene was cloned as a 1.2-kb HindIII/MluI fragment into the same sites of the modified pWHM1250 to afford pBS1108, in which the expression of sgcR3 is under the control of ErmE*. Finally, pBS1107, pBS1097 and pBS1108 were introduced into the wild-type S. globisporus strain by protoplast transformation to afford recombinant S. globisporus strains SB1014, SB1015 and SB1016, which overexpress sgcR1 (SB1014), sgcR2 (SB1015) and sgcR3 (SB1016), respectively.4
Plasmid pBS1109, in which sgcR2 and sgcR3 are expressed in tandem under the control of ErmE*, was constructed by inserting the 1.2-kb HindIII/MluI fragment of sgcR3 into the same sites of pBS1097. Similarly, insertion of the 1.1-kb BamHI/XbaI fragment of sgcR1 into the same sites of pBS1109 yielded pBS1110, in which ErmE* controlled the expression of all three genes sgcR1, sgcR2 and sgcR3. Introduction of pBS1109 and pBS1110 into the wild-type S. globisporus strain by protoplast transformation finally afforded the recombinant strains SB1017 (sgcR2R3 overexpressing) and SB1018 (sgcR1R2R3 overexpressing), respectively, which overexpress the selected regulatory genes in tandem.4
Production and isolation of C-1027 and heptaene
S. globisporus wild-type and recombinant strains were cultured in A9 medium using a two-step fermentation for C-1027 and heptaene production as described4, 15 and then treated to obtain three fractions as follows: (fraction 1) the cells were pelleted by centrifugation; (fraction 2) the supernatant was adjusted to pH 4 and centrifuged to collect the precipitate; and (fraction 3) the remaining supernatant was saturated to 70% with ammonium sulfate, incubated at 4 °C for 3 h and the precipitated proteins were collected by centrifugation. The C-1027 chromophore was isolated from fraction 3 by acetone extraction as previously described.15 Fractions 1 and 2 were combined and extracted with acetone to isolate the heptaene metabolite.11 The acetone extracts were detected directly by HPLC after being concentrated.5, 11
Analysis of C-1027 by bioassays
C-1027 production was assayed against M. luteus ATCC 9431 as previously described.4, 15 For the liquid A9 medium cultures, the ammonium sulfate-precipitated fraction 3 containing the C-1027 chromoprotein complex was dissolved in the same volume of water and 5 μl of the resultant solution was used for each bioassay. For the solid ISP4 plates (25 ml ISP4 in a 9-cm Petri dish) growing S. globisporus strains, a round agar block of 0.5 cm diameter was placed directly onto an M. luteus-seeded Luria-Bertani agar plate for antibacterial assay.
Analytical and spectroscopic procedures for the C-1027 chromophore and heptaene
HPLC analysis was performed with a C18 column (5 μm, 250 mm × 4.6 mm, Alltech, Lexington, KY, USA) on a Varian HPLC system with an in-line Prostar 330 PDA detector (Woburn, MA, USA). HPLC detection of the C-1027 chromophore was the same as previously described.15 For HPLC analysis of the heptaene, the column was eluted at a flow rate of 1 ml min−1 with a gradient of acetonitrile with 0.1% trifluoroacetic acid from 50 to 100% for 15 min and maintained at 100% for a further 10 min.
Results and discussion
Manipulation of pathway regulation is an efficient strategy to increase specific secondary metabolite production.12, 13 Among the three regulators identified within the C-1027 biosynthetic gene cluster, SgcR3 has been characterized as a TylR-like activator,14 and SgcR1 and SgcR2 have been proposed to be positive regulators based on bioinformatic analyses. SgcR2 belongs to the AraC/XylS activator family,20 whereas SgcR1 is an StrR-like protein; the only two studied StrR-like regulators, StrR (36.7% identity)21 and NovG (35.1% identity),22 are both activators. On the basis of the premise that all three C-1027 regulators are potential activators, the three regulatory genes were overexpressed individually under the control of the constitutive ErmE* promoter on the high-copy-number plasmid pWHM1250 in S. globisporus C-1027 to generate strains SB1014 (sgcR1 overexpressing), SB1015 (sgcR2 overexpressing) and SB1016 (sgcR3 overexpressing).
Evaluation of the metabolic profiles of each of the recombinant strains revealed that SgcR1 was a particularly effective activator for the production of a heptaene, an early metabolite of the enediyne biosynthetic machineries.11 The sgcR1-overexpressing strain S. globisporus SB1014 produced 46±4 mg l−1 of the heptaene at day 2, which was about ninefold higher than that in wild-type (5±0.4 mg l−1), and by day 5 yielded 66±5 mg l−1, which is about fivefold higher than the 14±3 mg l−1 observed in wild type. The titers of heptaene in SB1015 (18±1 mg l−1) and SB1016 (24±1 mg l−1) were also higher than wild type at day 5 (Figure 2), supporting a positive regulatory role for all the three regulators.
Production of the heptaene in selected S. globisporus recombinant strains with the wild-type strain as a control. (a) Time course of heptaene production in S. globisporus wild-type (□) and recombinant strains with different regulatory genes overexpressed: SB1014 (▿), SB1015 (♦), SB1016 (⋄), SB1017 (▾) and SB1018 (▪). (b) Representative HPLC traces of heptaene production in the selected S. globisporus strains at day 5. The heptaene peak is indicated with (•).
The production of C-1027 in each strain was detected using a bioassay against M. luteus ATCC 9431. In the case of A9 liquid medium, all the three regulator-overexpressing recombinant strains produced C-1027 earlier than did the wild-type control (Figure 3). The S. globisporus wild-type or recombinant strains achieved highest C-1027 production at day 4 (SB1016) or day 5 (wild-type, SB1014 and SB1015), and all three recombinant strains exhibited larger inhibition zones compared with that observed with wild type at their highest titer (Figure 3b), confirming the upregulating characteristics of SgcR1, SgcR2 and SgcR3. The sgcR1-overexpressing mutant SB1014 produced the highest titer of C-1027 at day 5, which was two to three times the C-1027 titer in wild type when analyzed by HPLC (Figure 3c). In the case of ISP4 agar medium, similar results were obtained for SB1014. SB1015 and SB1016 also produced C-1027 earlier, but their largest inhibition zones were comparable in size to those of the S. globisporus wild type (Figure 3a). In summary, overexpression of each individual regulator induced similar titer increases for both C-1027 and heptaene in A9 medium, indicating a common regulatory system controlling the biosyntheses of both compounds. Moreover, SgcR1 overexpression was more effective at titer improvement than overexpression of either SgcR2 or SgcR3, implying that one or more genes controlled by SgcR1 are bottlenecks in C-1027 production.
Production of C-1027 in the selected S. globisporus recombinant strains with the wild-type strain as a control. (a) Time course of C-1027 production in the selected S. globisporus strains growing on ISP4 plates as evaluated by bioassays against M. luteus. (b) Time course of C-1027 production in the selected S. globisporus strains cultured in A9 media as evaluated by bioassays against M. luteus. (c) HPLC traces of the C-1027 chromophore produced by the selected S. globisporus strains in A9 media at day 5. C-1027 chromophore and the aromatized C-1027 chromophore are marked as (▿) and (○), respectively.
Given that SgcR1, SgcR2 and SgcR3 behave as activators, we reasoned that their co-overexpression in the S. globisporus wild-type strain could further improve the production of C-1027 and related compounds. Two pWHM1250-derived constructs, containing sgcR2R3 (pBS1109) or sgcR1R2R3 (pBS1110) in tandem under the control of the ErmE* promoter, were transformed into S. globisporus wild-type to afford SB1017 and SB1018, respectively. SB1017 produced more heptaene (31±5 mg l−1) at day 5 than SB1015 and SB1016, but less than SB1014. To our surprise, SB1018 produced heptaene with an efficiency comparable to that of the wild type (12±1 mg l−1) (Figure 2). For C-1027 titers in A9 liquid media, SB1017 reached optimum yield at day 4, and the inhibition zone was larger than those of C-1027 wild type. In contrast, the C-1027 titer in SB1018 was not improved (Figure 3). In the case of ISP4 agar media, C-1027 was produced earlier in both strains. However, the largest inhibition zones were comparable in size to those found for C-1027 wild type. The counterintuitive observation that overexpression of all three activators together in S. globisporus SB1018 did not further improve C-1027 or heptaene production implies the multifaceted regulation of C-1027 biosynthesis.
Owing to their low titers and inherent reactivity, HPLC analysis of enediyne production is laborious, often requiring a lengthy process of sample preparation, particularly for the nine-membered enediynes produced as chromoprotein complexes. Although antibacterial bioassays provide a convenient detection method, they are inevitably limited by a lack of specificity as other antibiotics may also inhibit the tested bacterial growth. Moreover, some newly discovered enediyne analogs such as the sporolides from Salinispora tropica have no detectable antibacterial activity,23 which limits the utility of antibacterial bioassays for discovering novel enediynes. The heptaene metabolite was previously shown to be produced by all 9- or 10-membered PKSEs and their cognate thioesterases thus far tested using in vivo and in vitro approaches.9, 10, 11 Subsequent discovery of this heptaene in fermentations with all enediyne producers inspired its use as a unique and convenient phenotypic indicator for enediyne production.11 As both the heptaene and final enediyne natural products are biosynthesized by PKSEs, production of heptaene indicates that the PKSE is functionally expressed and capable of synthesizing the polyketide precursor of the enediyne core. Previous study on S. globisporus showed that production of both C-1027 and heptaene was abolished in an sgcE PKSE null mutant, but could be restored by complementation with a functional copy of the sgcE gene in trans.11 In this study, we demonstrate that in all five S. globisporus recombinant strains, C-1027 and hepatene production was similarly affected. These data complement previously reported genotypic screenings for new enediyne producers.24, 25 More importantly, the use of heptaene production as an indicator of enediyne production presents novel opportunities for fermentation optimization efforts and the identification of cryptic and otherwise undetectable enediyne producers.
In conclusion, we have demonstrated that, in addition to SgcR3, the other two regulators, SgcR1 and SgcR2, identified in the C-1027 biosynthetic gene cluster also have positive regulatory roles in C-1027 production. The titers of both C-1027 and the heptaene metabolite were significantly increased in the sgcR1-overexpressing S. globisporus strain SB1014. Moreover, data presented here support the application of heptaene production as a convenient phenotypic indicator for enediyne production.
References
Smith, A. L. & Nicolaou, K. C. The enediyne antibiotics. J. Med. Chem. 39, 2103–2117 (1996).
Van Lanen, S. G. & Shen, B. Biosynthesis of enediyne antitumor antibiotics. Curr. Top. Med. Chem. 8, 448–459 (2008).
Shao, R. G. & Zhen, Y. S. Enediyne anticancer antibiotic lidamycin: chemistry, biology and pharmacology. Anticancer Agents Med. Chem. 8, 123–131 (2008).
Liu, W. & Shen, B. Genes for production of the enediyne antitumor antibiotic C-1027 in Streptomyces globisporus are clustered with the cagA gene that encodes the C-1027 apoprotein. Antimicrob. Agents Chemother. 44, 382–392 (2000).
Liu, W., Christenson, S. D., Standage, S. & Shen, B. Biosynthesis of the enediyne antitumor antibiotic C-1027. Science 297, 1170–1173 (2002).
Christenson, S. D., Liu, W., Toney, M. D. & Shen, B. A novel 4-methylideneimidazole-5 -one-containing tyrosine aminomutase in enediyne antitumor antibiotic C-1027 biosynthesis. J. Am. Chem. Soc. 125, 6062–6063 (2003).
Van Lanen, S. G., Lin, S. J. & Shen, B. Biosynthesis of the enediyne antitumor antibiotic C-1027 involves a new branching point in chorismate metabolism. Proc. Natl Acad. Sci. USA 105, 494–499 (2008).
Lin, S. J., Van Lanen, S. G. & Shen, B. A free-standing condensation enzyme catalyzing ester bond formation in C-1027 biosynthesis. Proc. Natl Acad. Sci. USA 106, 4183–4188 (2009).
Zhang, J. et al. A phosphopantetheinylating polyketide synthase producing a linear polyene to initiate enediyne antitumor antibiotic biosynthesis. Proc. Natl Acad. Sci. USA 105, 1460–1465 (2008).
Belecki, K., Crawford, J. M. & Townsend, C. A. Production of octaketide polyenes by the calicheamicin polyketide synthase CalE8: implications for the biosynthesis of enediyne core structures. J. Am. Chem. Soc. 131, 12564–12566 (2009).
Horsman, G. P., Chen, Y., Thorson, J. S. & Shen, B. Polyketide synthase chemistry does not direct biosynthetic divergence between 9- and 10-membered enediynes. Proc. Natl. Acad. Sci. USA; (e-pub ahead of print 7 June 2010; doi:10.1073/pnas.1003442107).
Olano, C., Lombó, F., Méndez, C. & Salas, J. A. Improving production of bioactive secondary metabolites in actinomycetes by metabolic engineering. Metab. Eng. 10, 281–292 (2008).
Chen, Y., Smanski, M. J. & Shen, B. Improvement of secondary metabolite production in Streptomyces by manipulating pathway regulation. Appl. Microbiol. Biotechnol. 86, 19–25 (2010).
Wang, L. et al. Role of sgcR3 in positive regulation of enediyne antibiotic C-102 production of Streptomyces globisporus C-1027. BMC Microbiol. doi:10.1186/1471-2180-9-14 (2009).
Lin, S. et al. Characterization of the SgcF epoxide hydrolase supporting an (R)-vicinal diol intermediate for enediyne antitumor antibiotic C-1027 biosynthesis. J. Am. Chem. Soc. 131, 16410–16417 (2009).
MacNeil, D. J. et al. Analysis of Streptomyces avermitilis genes required for avermectin biosynthesis utilizing a novel integration vector. Gene 111, 61–68 (1992).
Sambrook, J., Fritsch, E. F. & Maniatis, T. Molecular cloning, a laboratory manual. (Cold Spring Harbor Laboratory Press, New York, 1989).
Madduri, K. et al. Production of the antitumor drug epirubicin (4′-epidoxorubicin) and its precursor by a genetically engineered strain of Streptomyces peucetius. Nat. Biotechnol. 16, 69–74 (1998).
Kieser, T., Bibb, M. J., Buttner, M. J., Chater, K. F. & Hopwood, D. A. Practical Streptomyces genetics (The John Innes Foundation, Norwich, 2000).
Gallegos, M. T., Schleif, R., Bairoch, A., Hofmann, K. & Ramos, J. L. AraC/Xyls family of transcriptional regulators. Microbiol. Mol. Biol. Rev. 61, 393–410 (1997).
Retzlaff, L. & Distler, J. The regulator of streptomycin gene expression, StrR, of Streptomyces griseus is a DNA binding activator protein with multiple recognition sites. Mol. Microbiol. 18, 151–162 (1995).
Eustáquio, A. S., Li, S. M. & Heide, L. NovG, a DNA-binding protein acting as a positive regulator of novobiocin biosynthesis. Microbiology 151, 1946–1961 (2005).
Buchanan, G. O. et al. Sporolides A and B: structurally unprecedented halogenated macrolides from the marine actinomycete Salinisporatropica. Org. Lett. 7, 2731–2734 (2005).
Liu, W. et al. Rapid PCR amplification of minimal enediyne polyketide synthase cassettes leads to a predictive familial classification model. Proc. Natl Acad. Sci. USA 100, 11959–11963 (2003).
Zazopoulos, E. et al. A genomics-guided approach for discovering and expressing cryptic metabolic pathways. Nat. Biotechnol. 21, 187–190 (2003).
Acknowledgements
We thank Dr Y Li, Institute of Medicinal Biotechnology, Chinese Academy of Medical Sciences, Beijing, China, for the wild-type S. globisporus strain. This work is supported in part by the NIH Grants CA78747 and CA113297. GPH is the recipient of an NSERC postdoctoral fellowship.
Author information
Authors and Affiliations
Corresponding author
Additional information
We celebrate Dr Arnold Demain's 60-year distinguished scientific career and his pioneering work and long-lasting contributions to the study of antibiotics.
Rights and permissions
About this article
Cite this article
Chen, Y., Yin, M., Horsman, G. et al. Manipulation of pathway regulation in Streptomyces globisporus for overproduction of the enediyne antitumor antibiotic C-1027. J Antibiot 63, 482–485 (2010). https://doi.org/10.1038/ja.2010.55
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2010.55
Keywords
This article is cited by
-
Challenges and opportunities for natural product discovery, production, and engineering in native producers versus heterologous hosts
Journal of Industrial Microbiology and Biotechnology (2019)
-
Crystal structure of SgcJ, an NTF2-like superfamily protein involved in biosynthesis of the nine-membered enediyne antitumor antibiotic C-1027
The Journal of Antibiotics (2016)