Abstract
We demonstrated that the rox gene of Nocardia farcinica encodes a rifampicin monooxygenase capable of converting rifampicin to a new compound 2′-N-hydroxy-4-oxo-rifampicin with a markedly lowered antibiotic activity. The deletion mutation (Δrox) of the rox gene gave no significant influence to the rifampicin resistance of N. farcinica. However, transformation with a plasmid containing an overexpressing the rox gene markedly raised the rifampicin resistance in the strain with the deletion mutation of the rpoB2 gene as the principal rifampicin resistance determinant. On the other hand, rifampicin was decolorized by the wild-type strain, whereas it remained intact when incubated with the Δrox strain. Based on these results, it will be conclusive that the rox gene is capable of initiating rifampicin degradation with a new metabolite formation at the first step and having a role as the secondary rifampicin resistance factor in N. farcinica.
Similar content being viewed by others
Introduction
Nocardia species are naturally resistant to wide varieties of antibiotics, for example β-lactams, aminoglycosides, macrolides and rifamycins. Similar to mycobacteria, the hydrophobic nature of the nocardial cell-wall was once believed to act as a permeability barrier for the multidrug resistance of this organism. However, genome sequence analysis of Nocardia farcinica IFM 10152 has since revealed the existence of specific genes which explain resistance to each drug.1 As to rifampicin (RIF), the rpoB2 gene, a duplicated copy of the RNA polymerase β-subunit gene containing RIF-resistant amino acid substitutions, has been predicted to be a resistance determinant and its contribution to the RIF resistance of N. farcinica has been demonstrated experimentally.2 On the other hand, decolorization of RIF by N. farcinica has been reported3, 4 and the sequenced strain IFM 10152 decolorizes RIF in prolonged culture (unpublished observation). This observation implies the participation of decolorization in RIF resistance.
A variety of RIF-modifying enzymes has been identified in Nocardia and related genera, such as enzymes involved in phosphorylation,5 glycosylation4 and ribosylation.6 Therefore, in addition to the rpoB2 gene, we searched the N. farcinica genome for genes which might participate in RIF resistance and rox (formerly denoted as nfa35380) turned out to be a homologous gene to the iri gene that confers a low-level resistance to RIF in Rhodococcus equi.7 The deduced amino acid sequence of rox (473 aa) shows 64% identity and 77% similarity to that of iri (479 aa), thus suggesting that they share similar enzymatic activities. The Iri protein has been believed to be a monooxygenase, attacking the naphthalenyl moiety of RIF, based on its sequence similarity to many monooxygenases acting on phenolic compounds. However, the reaction product of the Iri protein has never been identified so there is no evidence for its proposed action as a monooxygenase. In this study, we determined the chemical structure of the reaction product of Rox protein to discuss plausible functions of rox in resistance to and decolorization of RIF.
Materials and methods
Bacterial strains and plasmids
N. farcinica IFM 10152 was obtained from the Medical Mycology Research Center, Chiba University, Japan, and maintained in our laboratory. Escherichia coli JM109 or BL21(DE3) was used as a host strain for gene cloning and expression, respectively. pKNL027_N05, a plasmid from the N. farcinica IFM 10152 ordered plasmid library (http://nocardia.nih.go.jp/), was used as the source of the rox gene fragment. pNV118 (unpublished), a high copy-number version of pNV18,8 was used for the expression of rox in Nocardia strains. pK18mobsacB9 was obtained from the National Institute of Genetics, Shizuoka, Japan and used for making deletion mutants.
Culture conditions
All strains were incubated at 37 °C unless otherwise indicated. Nocardia strains were grown in the brain–heart infusion (BHI) broth (Becton, Dickinson, Franklin Lakes, NJ, USA) or in a minimal medium (MM) consisting of Na2HPO4˙12H2O 1.5%, KH2PO4 0.3%, sodium citrate 0.05%, NH4Cl 0.02%, MgSO4˙7H2O 0.1%, NaCl 0.05% and glucose 1.5%. E. coli strains were grown in the Luria–Bertani (LB) broth. Minimum inhibitory concentrations (MICs) were determined by a standard microdilution method. For Nocardia strains, the MICs were scored after 48 h of incubation in BHI broth. For other bacterial strains, MICs were scored after 24 h of incubation in Mueller–Hinton II broth (Becton, Dickinson).
Recombinant DNA techniques
A BigDye Terminator v3.1 cycle sequencing kit and a 3130 Genetic Analyzer (Applied Biosystems, Foster City, CA, USA) were used for nucleotide sequencing, in accordance with the manufacturer's instructions. PCR was carried out in 10 μl reaction mixtures with a KOD-Plus-kit (TOYOBO, Osaka, Japan). Bacterial cells were used as templates for the screening of transformants by PCR.10 The PCR amplification program consisted of one cycle of 3 min at 98 °C, followed by 30 cycles of 20 s at 98 °C, 20 s at 55–60 °C and 20 s at 68 °C, with a final extension step at 68 °C for 5 min.
Construction of rox expression plasmid
A 1.6-kb DNA fragment containing rox (Figure 1) was amplified from pKNL027_N05 by PCR with the use of the primers 35380F (CGCTCTAGAGGTGATCGATGTGATCATCG) and 35380R (ATATCTAGATCAGGTGGCGGCGCCGAAC). The amplified fragment was cloned in-frame at the XbaI site of pUC18 and its nucleotide sequence checked, obtaining pUC18rox.
Conversion of RIF using E. coli resting cells
E. coli BL21(DE3)/pUC18rox was grown in 1 l of LB broth containing 50 μg ml−1 of ampicillin for 24 h at 27 °C. Cells were washed with sterile water and suspended in 500 ml of 100 mM potassium phosphate buffer (pH 7.0), supplemented with 1% glucose. RIF was added to the cell suspension at a final concentration of 100 μg ml−1 and incubated for 24 h at 27 °C with vigorous shaking. Cells were removed by centrifugation and the reaction products were recovered from the buffer by adsorption on DIAION HP20 resin (Mitsubishi Chemical, Tokyo, Japan). The resin was washed with water and the reaction products were eluted with methanol. These procedures were repeated and the reaction products were pooled. To monitor the reaction, thin-layer chromatography (TLC) was performed on a silica gel plate (silica gel 60F254, Merck & Co., Whitehouse Station, NJ, USA) using a chloroform:methanol:water (65:15:5, lower phase) solvent system.
Purification of reaction products
The pooled reaction products were purified by silica gel-column chromatography (silica gel 60; Nacalai Tesque, Kyoto, Japan) using stepwise elution with chloroform containing increasing amounts of methanol (1:0, 30:1, 20:1, 15:1, 10:1, 5:1 and 1:1). The fractions (10:1 and 5:1) containing the main compound were pooled and applied to a preparative TLC plate (silica gel 60F254, 0.5 mm thick, Merck & Co.) and developed using a chloroform:methanol:water (65:15:5, lower phase) solvent system. The isolated fraction was further purified by Sephadex LH-20 (GE Healthcare, Uppsala, Sweden) column chromatography using methanol as an elution solvent. The final yield was 50 mg of purified product.
Physicochemical analyses
The 1H-, 13C- and 2D-NMR (1H–1H COSY, heteronuclear multiple quantum coherence (HMQC), 1H–13C and 1H–15N heteronuclear multiple bond coherence (HMBC)) spectra were recorded on a JNM-ECA 500 NMR spectrometer (JEOL, Tokyo, Japan). Electrospray ionization (ESI) MS was carried out with a Platform LC mass spectrometer (Micromass MS Technologies, Manchester, UK). Samples for ESI-MS analysis were dissolved in methanol. IR and UV/VIS spectra were obtained using a Spectrum One FT-IR spectrometer (Perkin Elmer, Waltham, MA, USA) and a V-520 UV/VIS spectrophotometer (JASCO, Tokyo, Japan), respectively. Matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) MS was carried out with a AXIMA-QIT mass spectrometer (Shimadzu, Kyoto, Japan) using 2,5-dihydroxybenzoic acid as an ionization matrix. RIF-6′-N-oxide (RIF-NO) was synthesized by the oxidation of RIF with m-chloroperbenzoic acid and its structure was confirmed by NMR analyses (Supplementary Tables S1–S3).
Construction of Δrox mutant
The 3.1-kb SphI–BamHI fragment of pKNL027_N05 (Figure 1) was cloned into pK18mobsacB,9 yielding pKrox. To make an in-frame deletion of rox, an inverse PCR with the primers rox5R (ATCGATCACGTCAGTCGTCCCAT) and rox3F (TGGTTCGGCGCCGCCACCTGA) was carried out, using pKrox DNA as the template. The amplified fragment was purified by agarose gel electrophoresis, its 5′ end was phosphorylated with T4 polynucleotide kinase (TOYOBO), and then the fragment was self-ligated, obtaining a deletion allele delivery plasmid pKΔrox. Deletion mutants were selected as described in a previous paper.2 Legitimate single and double crossover clones were detected by colony-direct PCR analyses10 with appropriate combinations of the primers rox5F (ATCGGAACGACTGCCTCGGCACGGA), rox3R (AACTGGACGCCCATGTCGTCACCGT), M13-325 (GTGCTGCAAGGCGATTAAGTTGG) and M13-RV507 (TCCGGCTCGTATGTTGTGTGGA). To complement the mutation, the 3.1-kb SphI–BamHI fragment of pKNL027_N05 was cloned into pNV118, yielding pNV118rox.
Decolorization of RIF by Nocardia strains
Nocardia strains were grown in 50 ml of BHI broth for 24 h at 37 °C. Cells were washed twice with sterile water, suspended in 50 ml of MM containing 50 μg ml−1 of RIF and incubated for 24 h at 37 °C. A total of 1 ml portions of cultures were taken and extracted with an equal volume of methanol. After removing cell debris by centrifugation, the extracts were subjected to reverse-phase HPLC using a Prominence system (Shimadzu) with a COSMOSIL 5C18-AR-II column (4.6 by 150 mm; Nacalai Tesque). The mobile phase consisted of 25 mM NaH2PO4:CH3CN (65:35, v/v), running at a flow rate of 1 ml min−1.
Phylogenetic analysis
Phylogenetic analysis was conducted using the MEGA program version 4.11
Results
Identification of monooxygenated RIF
To demonstrate the RIF conversion activity of the protein encoded by rox, an expression plasmid pUC18rox was constructed. After various expression conditions were tested, it turned out that E. coli BL21(DE3)/pUC18rox grown in LB broth at 27 °C without the addition of IPTG yielded reproducible results. A total of 0.5 g of RIF was converted using resting cells of E. coli BL21(DE3)/pUC18rox and the reaction products were analyzed by TLC (Figure 2a). After incubation for 24 h, the RIF spot (Rf 0.6) disappeared, whereas a spot with an RF value of 0.4 was detected instead. This newly detected compound was extracted from the TLC plate and subjected to MALDI-TOF MS analysis. The compound showed a molecular ion peak [M+Na]+ at m/z 861.46. Because this result indicates the addition of one oxygen atom to RIF ([M+Na]+ at m/z 845.46), the compound was considered to be monooxygenated RIF (RIF-O). Therefore, we further purified the compound.
Conversion of rifampicin (RIF) by Rox. (a) RIF was incubated in 100 mM potassium phosphate buffer (pH 7.0) containing 1% glucose with Escherichia coli BL21(DE3) carrying pUC18 (lane 1) or pUC18rox (lane 2) or without E. coli (lane 3). Reaction mixtures were extracted with 50% methanol and subjected to thin layer chromatography (TLC). (b) RIF (50 μg ml−1) was incubated with or without Nicordia farcinica strain in minimal media (MM) at 37 °C. The broths were subjected to high-performance liquid chromatography (HPLC) analysis or photographed after 24 and 72 h, respectively. a, no bacteria; b, wild-type; c, Δrox; and d, Δrox/pNV118rox. The arrow indicated RIF peak.
Structure elucidation of RIF-O
The molecular formula of RIF-O, C43H58N4O13, was confirmed by HRESI-MS (measured m/z 839.4083, calc for C43H59N4O13 [M+H]+, 839.4079, Δ+0.4 mmu), indicating the addition of one oxygen atom to RIF.
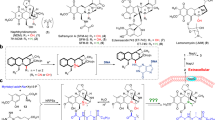
The chemical shift assignment for 1H- and 13C-NMR of RIF-O and RIF was performed by the analysis of NMR experiments (DEPT, 1H–1H COSY, HMQC and HMBC) (Table 1). In RIF-O, all the protons attached to the carbons, including the carbons in the ansa-chain (C15–29-O-C12) and in the piperazine moiety, were assigned successively to the same positions as RIF. The NH proton signals split into two peaks at 7.06 (0.5H) and 7.23 (0.5H), but both signals correlated to the amide-NH in 1H–15N HMBC experiments. These observations indicate that neither a change in the ansa-chain nor the addition of oxygen atom to any carbons occurred during the enzymatic conversion of RIF. On the other hand, the chemical shift of the H3-34 signal (δH 0.69) of RIF-O was remarkably lower than that of RIF (δH −0.27). This implies the oxidation of the naphthoquinol moiety of RIF, because the same downfield shift has been reported in the case of oxidation of rifamycin SV (naphthoquinol form) to rifamycin S (naphthoquinone form).12, 13 This presumption was also supported by the lack of absorption at 470 nm in RIF-O (Supplementary Table S4). In IR spectra, RIF-O showed an additional peak at 1615 cm−1, indicating the existence of an additional carbonyl group in RIF-O. Furthermore, the C-4 signal (δC 184.6) of RIF-O suggested the existence of a quinine-like quaternary carbon (that is, quinone carbonyl).
Next, we tried to synthesize RIF-O by chemical oxidization of RIF with m-chloroperbenzoic acid but yielded RIF-6′-N-oxide (RIF-NO), an RIF metabolite with the same molecular formula as RIF-O.14 Subsequently, we compared the NMR spectra of RIF-NO with those of RIF. The 15N chemical shift (δN) was measured by 1H–15N HMBC (Table 1). The results showed that the chemical shifts at C-5′ and -7′ (α position of N-6′) moved to a lower magnetic field (Table 1). In addition, the chemical shift at N-6′ moved to a higher magnetic field (RIF, δN 38.1; RIF-NO, δN 103.7). These observations indicate that N-oxygenation of RIF resulted in chemical shift changes at the nitrogen, flanking protons and carbons. In the case of RIF-O, the signal changes were as follows. No large chemical shift changes were observed at N-3′ and N-6′ (Table 1 and Supplementary Table S3). Although the chemical shift at N-2′ of RIF-O could not be obtained, but that of a minor component (RIF-Om) was observed (δN 321.2). The chemical shift at N-2′ of RIF-Om was distinct from that of RIF. RIF-Om was always contained in the RIF-O preparation at a very low concentration (data not shown) and considered to be a tautomer of RIF-O. The downfield shifts of 13C-NMR data at C-4′, -5′ and -7′ were minor. In contrast, a remarkable downfield shift was observed at C-1′ (RIF-O, δC 155.0; RIF, δC 137.8). These suggest a change at N-2′ in RIF-O as shown at N-6′ in RIF-NO.
Taken together all the above observations, we conclude that the structure of RIF-O is 2′-N-hydroxy-4-oxo-RIF (Figure 3a), and propose a mechanism involving the enzymatic addition of oxygen to the N-2′ of RIF, resulting in RIF-2′-N-oxide (RIF-Om), followed by RIF-O (Figure 3b).
Antimicrobial activity of RIF-O
The antimicrobial activity of RIF-O was evaluated (Table 2). The MIC values of RIF-O increased by more than 100-fold, thereby indicating a more than 100-fold reduction in antimicrobial activity. However, RIF-O retained its antimicrobial activity against K. rhizophila, S. aureus and B. subtilis. This suggests that RIF was not completely inactivated by Rox protein.
RIF resistance conferred by rox
In order to elucidate the role of rox in the RIF resistance of N. farcinica, we constructed an in-frame unmarked deletion mutant, Δrox, as described in MATERIALS AND METHODS. The MIC value of a Δrox strain was the same as that of the wild-type strain (Table 2). To complement the mutation, we cloned a rox-containing DNA fragment (Figure 1) into pNV118. The resultant plasmid pNV118rox was introduced into the Δrox strain. However, no significant change in the MIC value was observed after the introduction. We next transformed the ΔrpoB2 strain2 with pNV118rox. The ΔrpoB2 strain was susceptible to 0.25 μg ml−1 of RIF, whereas the strain carrying pNV118rox showed elevated resistance to RIF with an MIC of 8 μg ml−1 (a 32-fold increase).
Role of rox in RIF decolorization
We investigated the RIF-converting ability of the Δrox strain. When RIF was incubated with the wild-type N. farcinica strain, neither RIF nor its color was detectable after 24 h and after 72 h, respectively (Figure 2b), suggesting the degradation of RIF. In contrast, RIF seemed to be intact in the Δrox strain after incubation for 24 h. Furthermore, the loss of ability of the Δrox strain to degrade RIF was completely restored by transformation with pNV118rox. These results suggest that monooxygenation by Rox protein is required to decolorize RIF.
Discussion
As described above, we demonstrated that the rox gene encodes an RIF monooxygenase. Monooxygenases constitute a large, divergent protein family and many of them are flavin adenine dinucleotide (FAD)-dependent enzymes. Rox protein is also considered to be a FAD-dependent monooxygenase because it contains the FAD-binding domain (Pfam accession number PF01494). Although the determination of the RIF-O structure demonstrated that Rox protein was a monooxygenase adding an oxygen to the N-2′ position of RIF (Figure 3), Rox protein seems to be different from known FAD-dependent monooxygenases. Phylogenetic analysis of known FAD-dependent monooxygenases and proteins highly homologous to Rox clearly showed that the Rox-containing protein family is distinguishable from those consisting of known FAD-dependent monooxygenases (Figure 4). Little is known about the functions of the Rox-containing family proteins and, therefore, we believe Rox should be the first enzyme in this family for which the real reaction has been experimentally determined. From this point of view, it would be interesting to know the detailed reaction mechanism and structure of this enzyme.
Phylogenetic analysis of known flavin adenine dinucleotide (FAD)-dependent monooxygenases and proteins highly homologous to Rox. An unrooted tree was obtained by the neighbor-joining method with multiple alignment of protein sequence using the MEGA program.10 Shaded areas denote protein families, which include functionally similar known FAD-dependent monooxygenases.
Rifampicin was converted to a reddish compound (RIF-O) by E. coli carrying the rox gene, but was completely decolorized by the wild-type N. farcinica strain (Figure 2b). This indicates the involvement of additional steps in the decolorization of RIF by N. farcinica. Furthermore, the inability of the Δrox strain to decolorize RIF suggests that decolorization is probably initiated by monooxygenation by Rox protein. Further study will be required to determine how RIF is decolorized after monooxygenation.
No difference in RIF resistance was observed between the wild-type and Δrox strains (Table 2). This indicates that the rpoB2 gene is the principal RIF resistance determinant of N. farcinica.2 However, a partial restoration of RIF resistance in the ΔrpoB2 strain carrying pNV118rox suggests that rox plays a role as the secondary RIF-resistance factor in N. farcinica. It is considered that the ΔrpoB2 strain, especially the pNV118rox-carrying strain, was able to convert a relatively low amount of RIF using the Rox protein produced before the addition of RIF and then grew as the RIF concentration decreased. These observations led us to a notion that rpoB2 and rox may have roles like vanguard and rear-guard, respectively, to resist RIF in this microorganism.
References
Ishikawa, J. et al. The complete genomic sequence of Nocardia farcinica IFM 10152. Proc. Natl Acad. Sci. USA 101, 14925–14930 (2004).
Ishikawa, J., Chiba, K., Kurita, H. & Satoh, H. Contribution of rpoB2 RNA polymerase β subunit gene to rifampin resistance in Nocardia species. Antimicrob. Agents Chemother. 50, 1342–1346 (2006).
Tanaka, Y. et al. Different rifampicin inactivation mechanisms in Nocardia and related taxa. Microbiol. Immunol. 40, 1–4 (1996).
Yazawa, K. et al. Inactivation of rifampin by Nocardia brasiliensis. Antimicrob. Agents Chemother. 37, 1313–1317 (1993).
Yazawa, K., Mikami, Y., Maeda, A., Morisaki, N. & Iwasaki, S. Phosphorylative inactivation of rifampicin by Nocardia otitidiscaviarum. J. Antimicrob. Chemother. 33, 1127–1135 (1994).
Quan, S., Venter, H. & Dabbs, E. R. Ribosylative inactivation of rifampin by Mycobacterium smegmatis is a principal contributor to its low susceptibility to this antibiotic. Antimicrob. Agents Chemother. 41, 2456–2460 (1997).
Andersen, S. J., Quan, S., Gowan, B. & Dabbs, E. R. Monooxygenase-like sequence of a Rhodococcus equi gene conferring increased resistance to rifampin by inactivating this antibiotic. Antimicrob. Agents Chemother. 41, 218–221 (1997).
Chiba, K. et al. Construction of a pair of practical Nocardia-Escherichia coli shuttle vectors. Jpn. J. Infect. Dis. 60, 45–47 (2007).
Schäfer, A. et al. Small mobilizable multi-purpose cloning vectors derived from the Escherichia coli plasmids pK18 and pK19: selection of defined deletions in the chromosome of Corynebacterium glutamicum. Gene 145, 69–73 (1994).
Ishikawa, J., Tsuchizaki, N., Yoshida, M., Ishiyama, D. & Hotta, K. Colony PCR for detection of specific DNA sequences in Actinomycetes. Actinomycetologica 14, 1–5 (2000).
Tamura, K., Dudley, J., Nei, M. & Kumar, S. MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol. Biol. Evol. 24, 1596–1599 (2007).
Casey, M. L. & Whitlock, H. W. Determination of the solution conformation of rifamycin S and derivatives by nuclear magnetic resonance. J. Am. Chem. Soc. 97, 6231–6236 (1975).
Walash, M. I., Belal, F., Metwally, M. E. & Hefnawy, M. M. Spectrophotometric determination of rifampin in the presence of its degradation products in pharmaceutical preparations. Anal. Lett. 26, 1905–1917 (1993).
Ripamonti, A., Ferrari, P. & Gallo, G. G. Structure identification of rifampicin N-oxide. Farmaco. Sci. 39, 806–809 (1984).
Acknowledgements
We thank Dr K Ishino for helpful discussion. This work was supported in part by KAKENHI on Priority Areas ‘Applied Genomics’ (no. 17019009) and on Research C (no. 19591196) from the Ministry of Education, Culture, Sports, Science and Technology of Japan and by Research Fund Project on Health Sciences focusing on Drug Innovation (no. KHA3301) from the Japan Human Health Sciences Foundation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The Journal of Antibiotics website (http://www.nature.com/ja)
Supplementary information
Rights and permissions
About this article
Cite this article
Hoshino, Y., Fujii, S., Shinonaga, H. et al. Monooxygenation of rifampicin catalyzed by the rox gene product of Nocardia farcinica: structure elucidation, gene identification and role in drug resistance. J Antibiot 63, 23–28 (2010). https://doi.org/10.1038/ja.2009.116
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2009.116
Keywords
This article is cited by
-
Rifamycin antibiotics and the mechanisms of their failure
The Journal of Antibiotics (2021)
-
The complex resistomes of Paenibacillaceae reflect diverse antibiotic chemical ecologies
The ISME Journal (2018)
-
Counteraction of antibiotic production and degradation stabilizes microbial communities
Nature (2015)
-
Resistance to rifampicin: a review
The Journal of Antibiotics (2014)
-
Metabolic Engineering of Nocardia sp. CS682 for Enhanced Production of Nargenicin A1
Applied Biochemistry and Biotechnology (2012)