Abstract
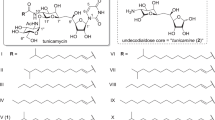
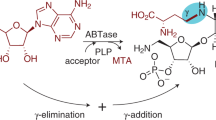
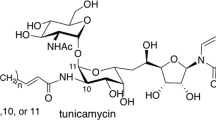
Tunicamycins are nucleotide sugar analogs produced by several Streptomyces species. In eukaryotes, tunicamycins inhibit UDP-N-acetylglucosamine: dolichol phosphate GlcNAc-1-P transferase (GPT) that catalyzes the first step in protein glycosylation. In bacteria they inhibit UDP-N-acetylmuramoyl-pentapeptide: undecaprenol phosphate MurNAc-pentapeptide-1-P transtransferase (MraY) that catalyzes an early stage in peptidoglycan cell wall assembly. Tunicamycins are substrate analog of GPT and MraY, such that the αβ-1″,11′-linked GlcNAc residue of the tunicamycins mimics the transferred GlcNAc-1-phosphate. The unusual structure of tunicamycins, particularly the unique 11-carbon sugar, tunicamine, and the αβ-1″,11′-O-glycosidic linkage, suggest its biosynthesis to be unique. This review discusses potential biosyntheses for tunicamycins via the synthesis and conjugation of uridine-5′-aldehyde and UDP-4-keto-N-acetylgalactosamine-5,6-ene and the subsequent formation of the α,β-1″,11′ glycosidic linkage.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Price, N., Tsvetanova, B. Biosynthesis of the Tunicamycins: A Review. J Antibiot 60, 485–491 (2007). https://doi.org/10.1038/ja.2007.62
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ja.2007.62
Keywords
This article is cited by
-
Modulation of MRSA virulence gene expression by the wall teichoic acid enzyme TarO
Nature Communications (2023)
-
A metabolic modeling approach reveals promising therapeutic targets and antiviral drugs to combat COVID-19
Scientific Reports (2021)
-
Homologous expression of lysA encoding diaminopimelic acid (DAP) decarboxylase reveals increased antibiotic production in Streptomyces clavuligerus
Brazilian Journal of Microbiology (2020)
-
Synergistic enhancement of beta-lactam antibiotics by modified tunicamycin analogs TunR1 and TunR2
The Journal of Antibiotics (2019)
-
Antibiotic sensitivity reveals that wall teichoic acids mediate DNA binding during competence in Bacillus subtilis
Nature Communications (2018)