Abstract
Antarctic sea-ice bacterial community composition and dynamics in various developmental stages were investigated during the austral winter in 2013. Thick snow cover likely insulated the ice, leading to high (<4 μg l–1) chlorophyll-a (chl-a) concentrations and consequent bacterial production. Typical sea-ice bacterial genera, for example, Octadecabacter, Polaribacter and Glaciecola, often abundant in spring and summer during the sea-ice algal bloom, predominated in the communities. The variability in bacterial community composition in the different ice types was mainly explained by the chl-a concentrations, suggesting that as in spring and summer sea ice, the sea-ice bacteria and algae may also be coupled during the Antarctic winter. Coupling between the bacterial community and sea-ice algae was further supported by significant correlations between bacterial abundance and production with chl-a. In addition, sulphate-reducing bacteria (for example, Desulforhopalus) together with odour of H2S were observed in thick, apparently anoxic ice, suggesting that the development of the anaerobic bacterial community may occur in sea ice under suitable conditions. In all, the results show that bacterial community in Antarctic sea ice can stay active throughout the winter period and thus possible future warming of sea ice and consequent increase in bacterial production may lead to changes in bacteria-mediated processes in the Antarctic sea-ice zone.
Similar content being viewed by others
Introduction
Antarctic sea ice is an important component of the Southern Ocean ecosystem from both the geophysical and biogeochemical standpoints. It covers vast areas from 3 × 106 km2 during the sea-ice minimum in February up to 18.3 × 106 km2 in the September maximum (Comiso, 2010). In contrast to the diminishing sea ice extent in the Arctic (Perovich et al., 2014), Antarctic sea ice is globally expanding (IPCC, 2013). However, this encompasses a large spatial heterogeneity meaning that there are indeed areas (for example, the Antarctic Peninsula) in which the surface ice temperatures are increasing and ice extent decreasing (Stammerjohn et al., 2008; Comiso, 2010). Also, the inter-annual variation in sea ice is high: for example, in 2017 during February minimum sea ice extent was 2.35 × 106 km2, which was 24.4%, below the 1981–2010 average (NOAA, 2017).
In the Southern Ocean, dynamic sea-ice growth through the pancake ice (PCI) cycle and deformed growth (for example, rafting) is common (Lange et al., 1989; Haas, 2010; Petrich and Eicken, 2010). In addition, the thick snow cover and dynamic growth cause flooding, which introduces additional nutrients from seawater to the surface and internal ice horizons (Lange et al., 1990; Eicken et al., 1994; Fritsen et al., 1994). These physical events induce a more distributed algal biomass than in the Arctic, that is, with chlorophyll-a (chl-a) peaks in the upper and middle horizons of the sea-ice cover, in addition to the more prevalent bottom-ice maxima (Meiners et al., 2012; Arrigo, 2014). The highest chl-a concentrations in Antarctic sea ice occur during spring and summer. However, unlike in the Arctic, peaks in chl-a are also observed in autumn (Fritsen et al., 1994; Delille et al., 2002; Meiners et al., 2012).
Sea-ice algae, together with sea-ice bacteria are the most productive and abundant organisms in ice. As in the water column, bacteria in sea ice drive an active microbial loop, in which dissolved organic matter is recycled back to the upper trophic levels as bacterial biomass (Kottmeier et al., 1987; Smith and Clement, 1990; Laurion et al., 1995; Kaartokallio, 2004; Cowie et al., 2014). In addition, bacteria decompose particulate organic matter and remineralize nutrients (Sullivan and Palmisano, 1984; Deming, 2010). Sea-ice bacteria and their community structure are governed by abiotic and biotic factors such as temperature, salinity, light, substrate and nutrient availability, as well as grazing and viral lysis (Kaartokallio, 2004; Mock and Thomas, 2005; Kuosa and Kaartokallio, 2006; Riedel et al., 2007; Collins et al., 2008, 2010; Piiparinen and Kuosa, 2011; Collins, 2015). As the fluctuation of these factors is highly seasonal, they also induce the seasonal dynamics of bacterial abundance and community composition in sea ice.
Most sea-ice bacterial community studies have been conducted during spring and summer, when the sea-ice algal mass growth provides ample autochthonous substrate for bacteria. During the spring and summer seasons, copiotrophic (that is, bacteria adapted to growing best in nutrient-rich environments) Gammaproteobacteria (for example, genera Glaciecola and Colwellia) dominate the bacterial communities, together with copiotrophic Flavobacteriia (for example, genera Polaribacter and Flavobacterium) and Alphaproteobacteria (for example, genus Octadecabacter) both in first-year ice (FYI) and multiyear ice (Bowman et al., 1997, 2012; Brown and Bowman, 2001; Junge et al., 2002; Brinkmeyer et al., 2003; Kaartokallio et al., 2008; Deming, 2010; Hatam et al., 2014, 2016; Eronen-Rasimus et al., 2015).
In contrast to spring and summer, bacterial communities during winter are poorly known. Members of the oligotrophic Alphaproteobacteria of the SAR11 clade dominate the sea-ice bacterial communities in the upper ice column during the Arctic winter and remain nearly unchanged until the algal mass accumulation in spring (Collins et al., 2010). In newly formed Antarctic sea ice, bacterial activity is temporarily suppressed and later restored after consolidation of the sea ice (Grossmann and Dieckmann, 1994; Helmke and Weyland, 1995). With the consolidation of ice, psychrophilic (that is, bacteria that grow best at temperatures close to freezing point) predominante over psychrotolerant (that is, bacteria that tolerate cold temperatures) bacteria, which indicates change in community structure during winter (Helmke and Weyland, 1995). However, the Antarctic wintertime bacterial community structure has not been described to date, because the available information is limited to isolated bacterial strains (Brinkmeyer et al., 2003).
In this study, we describe the bacterial community dynamics in the different developmental stages of winter-pack ice in Antarctica. Understanding the community dynamics allows us to assess the biogeochemical role played by bacteria during periods of ice cover and provide insights on the potential consequences of climate change on the biogeochemical cycles and food webs of ice-covered seas.
Materials and methods
Study site, sampling and environmental variables
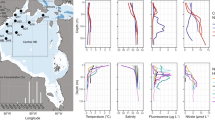
The samples were collected from 10 pack-ice stations along the Weddell and Lazarev Seas during the Antarctic Winter Ecosystem Climate Study (AWECS, leg ANT-XXIX/6)-expedition aboard the R/V Polarstern during the austral winter in June–July 2013 (Figure 1, Supplementary Table 1). The samples included different sea ice developmental stages: PCI, FYI and older fast ice/second-year ice (hereafter called SYI, Tison et al., 2014). Based on the preliminary interpretation of physical properties (Tison et al., 2014), in the Eastern Weddell Sea the ice was mostly frazil, deformed FYI with indication of rafting and signs of flooding (for example, station 500). At stations 515 and 517 close to the Antarctic Peninsula, the ice was predominantly columnar; however, 515 was SYI whereas 517 was FYI (Tison et al., 2014). All FYI and SYI stations were covered with a thick snow cover (mean 22.1 cm, range 14.5–35.6 cm), Tison et al., 2014) with generally increasing snow cover along the developmental stages of the ice (PCI<FYI<SYI), as previously described (Supplementary Table 1, Haas, 2010). At station 515, three sibling ice cores were collected two of which were SYI and one was 3 m long, from which only the middle horizons were sampled. The middle section had strong odour of H2S, suggesting anoxic conditions (hereafter called ANOX).
Map of the study area in the Weddell Sea, showing the sampling stations during the Antarctic Winter Ecosystem Climate Study (AWECS, leg ANT-XXIX/6) expedition. The map was generated with Ocean Data View (https://odv.awi.de; Schlitzer, 2016).
The ice cores were drilled with a trace-metal-clean (electropolished steel) ice auger (Lannuzel et al., 2006, 2007), 14 cm in diameter. Two ice cores were collected and pooled at each FYI station for the microbiological analyses. We emphasised careful sampling and subsequent sample processing to avoid contamination. The ice cores collected were cut with an ethanol-wiped handsaw into two to seven pieces, depending on the ice thickness (each horizon 10–30 cm), crushed and placed in sterile plastic containers at 4 °C over night in darkness after which the rest of the ice was quickly melted in a water bath with constant stirring. The melted samples were immediately filtered after becoming fully melted. For the ANOX, only ice sections from 130 to 160 cm were sampled. Before melting, part of the ice core was taken for the bacterial production measurements. For the bacterial diversity analyses, direct melting (Helmke and Weyland, 1995; Kaartokallio, 2004) was used to avoid external DNA or nutrient contamination of the samples.
Bacterial abundance (flow cytometer)
The samples for bacterial abundance were fixed with paraformaldehyde (final concentration 1%). The cells were stained with SYBR Green I (Sigma-Aldrich Inc, St Louis, MO, USA) at a final dilution of 1:10 000 for 15 min in the dark and analysed with a CyFlow Cube8 flow cytometer (Partek GmbH, Münster, Germany) using a 488-nm laser (Gasol and Del Giorgio, 2000) within 30 min of staining.
Bacterial production
Bacterial production was measured as the incorporation of 3H-labelled thymidine (Fuhrman and Azam, 1982) immediately after sampling, following the protocol described by Kaartokallio (2004). The ice sections were crushed, and from each section two aliquots and a formaldehyde-killed absorption blank (final conc. 1.85%) of approximately 7.5 g of crushed ice were taken. The samples were incubated in the dark at approximately –0.8 °C for 18 h with 2 ml of 0.2-μm-filtered and autoclaved seawater (salinity 33.9‰, measured on a unitless practical salinity scale; however, as it is essentially equal to ‰, it is used here; UNESCO, 1981), amended with [methyl-3H]-thymidine (specific activity 20 Ci mmol−1; PerkinElmer Inc, Waltham, MA, USA) at a final concentration of 20 nM. The incubations were stopped by adding formaldehyde (final conc. of 1.85%). The unincorporated 3H-thymidine was removed with the standard cold-trichloroacetic acid extraction method (Fuhrman and Azam, 1980, 1982). The samples were filtered onto 0.2-μm mixed cellulose ester filters (Advantec Mfs Inc, Dublin, CA, USA), the filters dissolved in Insta-Gel Plus scintillation cocktail (PerkinElmer) and the radioactivity measured with a Tri-Carb 2900TR scintillation counter (PerkinElmer) aboard the R/V Polarstern.
DNA extraction
For the DNA extractions, approximately 500 ml of the melted sea-ice were filtered onto sterile 0.22-μm membrane filters (Ø 47 mm; Whatman GE Healthcare, Little Chalfont, Kent, UK) and frozen in liquid nitrogen and later transferred to –80 °C.
The bacterial community DNA was extracted from the filters with a PowerSoil DNA Isolation Kit (Mo Bio Laboratories Inc, Carlsbad, CA, USA), as described by Eronen-Rasimus et al. (2014), 6 months after the cruise. In addition to the samples, negative controls without the sample were extracted.
Bacterial community analysis
For MiSeq sequencing, the 16S ribosomal RNA gene region from V1 to part of the V3 was amplified with a polymerase chain reaction, using the universal bacterial primers F8 (Chung et al., 2004) and R492 (Edwards et al., 1989). A two-step polymerase chain reaction and Illumina MiSeq (Illumina Inc, San Diego, CA, USA) paired-end multiplex sequencing were performed at the Institute of Biotechnology, University of Helsinki, Finland.
Bioinformatics for the MiSeq data
In all, approximately 6.2 million raw reads covering the V1–V3 region of the 16S ribosomal RNA gene were obtained with the Illumina MiSeq platform. Primer removal was done with Cutadapt (v. 1.7.1; Martin, 2011). The paired-end reads were merged with Paired-End reAd mergeR (Pear, v. 0.9.6; Zhang et al., 2014). Quality filtering (> 400 bp, maximum expected error 2), chimera checking (UCHIME; Edgar et al., 2011) and operational taxonomic unit clustering (Edgar, 2013) were done according to the UPARSE pipeline (Edgar, 2013). After quality filtering, a total of 1 298 455 sequences remained for further analyses. Taxonomic classification of the operational taxonomic units was done with Silva (v. 119, 60% confidence threshold; Quast et al., 2013) in Mothur (v. 1.36.1; Schloss et al., 2009). Before the statistical analyses, chloroplasts, mitochondria and singletons were removed based on the phylogenetic classification with Silva, and the libraries were normalised with metagenomeSeq (Paulson et al., 2013) in R (v. 3.2.4) (R Development Core Team, 2011). In all, 1273 operational taxonomic units including 717 232 sequences were obtained. Raw reads were deposited into the Sequence Read Archive of National Center for Biotechnology Information under study accession number SRP094398.
Statistical analysis
All multivariate analyses were performed on the Bray–Curtis dissimilarity matrix derived from square-root-transformed, normalised values. Square-root transformation was applied to reduce the contribution of dominant species, because the Bray–Curtis resemblance measure does not scale individual sequences by the total values throughout all samples, and our samples showed large differences in abundance.
The bacterial community dynamics among all samples (n=50) was visualised with non-metric multidimensional scaling (50 iterations). To determine whether the sea-ice bacterial communities differed significantly between different ice types (fixed factor; PCI=7, FYI: n=29, SYI: n=9, ANOX: n=5), a permutational multivariate analyses of variance (PERMANOVA) with pairwise comparisons (Anderson, 2001; McArdle and Anderson, 2001) was performed. As the data were unbalanced, all the tests were performed using type III sums of squares. A total of 9999 permutations, using unrestricted permutation of raw data (Manly, 1997) were performed, which is recommended for one-way designs (Anderson et al., 2008). The homogeneity of dispersion was tested with permutational multivariate analysis of dispersion (Anderson, 2006), using the distance to the centroids. Homogeneity of dispersion (that is, homogeneity of variance) is an assumption in PERMANOVA and thus needed to discriminate whether the location, dispersion or both explains the variation in the bacterial communities.
To determine the association between the chl-a and bacterial community composition, canonical analysis of principal coordinates (canonical correlation analysis; Anderson and Willis, 2003) based on the Bray–Curtis dissimilarity matrix derived from square-root-transformed, normalised values of the bacterial 16S ribosomal RNA gene sequences was performed (test statistics with 9999 permutations).
The correlation between chl-a and bacterial production (measured as thymidine incorporation) and abundance were calculated with two-way Spearman’s rank-sum rho with a base package of R software (v. 3.2.4; R Development Core Team, 2011).
For the multivariate analyses, Plymouth Routines In Multivariate Ecological Research (PRIMER) v. 6 software (Clarke and Gorley, 2006) with the add-on package permutational ANOVA/MANOVA+ (PERMANOVA+) (Anderson et al., 2008) were used.
Results and discussion
The bacterial communities in the different developmental stages of sea ice, that is, PCI, FYI, SYI and ANOX, were significantly different. Bacterial community dynamics was linked with the age of the ice and increasing chl-a concentrations, suggesting that as in spring and summer sea ice, the sea-ice bacteria and algae may be also coupled during the Antarctic winter. In addition, predominance of sulphate-reducing bacteria together with the odour of H2S suggests that under suitable conditions reduction of sulphur compounds may occur in sea ice.
Based on the preliminary interpretation of physical properties (Tison et al., 2014), all FYI and SYI stations were covered with a thick snow layer (mean 22.1 cm, range 14.5–35.6 cm), efficiently insulating the ice from cold air leading to warmer ice (Tison et al., 2014). The ice was, in most cases, permeable (relative brine volume above 5%, calculated as a function of temperature and salinity—equation 2.6 of Petrich and Eicken (2010); Golden et al. (1998); Tison et al. (2014)). Ice permeability allows brine transport and therefore potentially favourable conditions for the sea-ice bacteria and algae.
The maximum chl-a concentrations, that is, a proxy for algal biomass, globally increased along with the age of ice from young PCI to thick SYI (Figure 2). The variation in chl-a concentrations in FYI was wide, ranging from stations with low (<4 μg l−1) chl-a concentrations to stations (493, 496, 498, 500 and 506b) that showed chl-a concentrations up to 18.1 μg l−1 (Tison et al., 2014). The highest chl-a (113.15 μg l−1) concentrations were observed in the middle of the SYI (110–120 cm), suggesting that it originated from the sea-ice-algal bottom ‘bloom’ from the previous summer. Unlike the typical bottom-ice-dominated chl-a peaks in the Weddell Sea (Meiners et al., 2012), the chl-a peaks in our study were vertically distributed along the ice cores. This was the likely result of flooding, rafting and the predominance of frazil ice (preliminary ice-type observations from Tison et al., 2014), which enhance scavenging of algae (Garrison et al., 1983; Arrigo et al., 2010). High chl-a concentrations in FYI were arguably preserved from the autumnal growth of sea-ice algae as the day length was very short and solar angle and light levels low. Low incident irradiation combined with thick snow cover likely prevented any noticeable photosynthetic activity. In addition, high cell-specific chl-a content because of the low-light adaptation may have contributed to the high chl-a concentrations. However, at the early-winter PCI stations with thin or no snow cover in the Eastern Weddell Sea, the growth of sea-ice algae could have been possible, as reported also earlier (Melnikov et al., 1998).
Bacterial production (measured as thymidine incorporation), bacterial abundance and chl-a concentrations in the sea-ice samples, Weddell Sea, Antarctica. Chl-a results are redrawn from Tison et al. (2014). Note the different scales.
Strong association between the bacterial community composition and chl-a was observed in canonical correlation analysis (canonical analysis of principal: 0.92, P=0.0001; Anderson and Willis, 2003), indicating that the development of the bacterial community was associated with the availability of the algal-derived substrate (Figure 3). In addition, both the bacterial production and abundance correlated with the chl-a concentrations (Spearman’s ρ: 0.63, P: 2.291 × 10−6, ρ: 0.83, P: 1.51 × 10−11), further supporting the coupling between the algal and bacterial communities. The coupling of the sea-ice bacterial community composition and chl-a has been also previously reported in early spring and summer sea ice (Cowie et al., 2014; Eronen-Rasimus et al., 2016). However, a previous mid-winter study showed no correlation between chl-a and bacterial biomass likely because of the low chl-a concentrations in the ice (Stewart and Fritsen, 2004). Our results suggest that warmer winter ice temperatures and associated high algal biomass in the ice may also sustain bacterial activity during the winter months, which may affect the bacteria-mediated biogeochemical processes in the ice. Mid-winter algal peaks and associated bacterial activity have also been described in warm and nutrient-rich Baltic Sea ice (Kaartokallio, 2004).
Canonical analysis of principal coordinates (CAP canonical correlation analysis; Anderson and Willis, 2003) based on the square-root transformed Bray–Curtis dissimilarity matrix of the bacterial 16S ribosomal RNA (rRNA) gene sequences. In total, CAP analysis explained 95% of the variation in the bacterial communities (choice of m=19, P=0.0001), with very high canonical correlation between the bacteria community composition and chl-a concentration (0.92, P=0.0001).
The bacterial communities differed significantly among the ice types (PERMANOVA: pseudo-F 12.124, P=0.0001; pairwise tests: PCI vs FYI, PCI vs ANOX, FYI vs ANOX, FYI vs MYI and MYI vs ANOX P: 0.0001, PCI vs SYI 0.0002, Figure 4). As the differences in dispersion (that is, assumption of the homogeneity of variance is violated, permutational multivariate analysis of dispersion; P<0.05) were detected in comparison with the PCI and other ice types, the differences were explained by both location (that is, ice type) and dispersion (that is, differences between group variances). The dispersion effect was visualised in the non-metric multidimensional scaling (Figure 4), because the PCI station formed a tighter group than did the other ice types. However, despite the dispersion effect, the development of the communities along with the age of the ice was evident according to non-metric multidimensional scaling (Figure 4), supporting the view that there were real differences in bacterial communities between the ice types.
Non-metric multidimensional scaling (NMDS) plot showing changes in the sea-ice bacterial communities, based on 16S ribosomal RNA (rRNA) gene sequences, in the different developmental stages of the ice. The two-dimensional (2D) stress indicates how well the rank order relationships are described in two-dimensional space (0.1=good ordination with no real prospect of a misleading interpretation; Clarke and Warwick, 2001).
Alphaproteobacteria, Gammaproteobacteria and Flavobacteriia dominated the bacterial communities in all ice types (PCI, FYI, SYI, ANOX; Figure 5). Alphaproteobacteria decreased along with the age of the ice (PCI: 48.8%; FYI: 40.0%; SYI: 31.2%) whereas Gammaproteobacteria and Flavobacteriia increased (PCI: 12.7 and 19.8%; FYI: 19.7 and 30.5%; SYI: 23.2 and 32.9%). The predominant bacterial groups in FYI and SYI were mostly common sea-ice bacteria such as Rhodobacteraceae (for example, genera Octadecabacter and Loktanella), Alteromonadaceae (for example, genera Glaciecola and Candidatus Endobugula) and the BD7-8 marine group (belonging to Gammaproteobacteria), as well as Flavobacteriaceae (for example, genus Polaribacter; Figure 5). However, the prevalence of the most abundant genera in PCI, FYI and SYI varied among the ice types, for example, showing a shift from predominance of common open-water SAR11 clade bacteria (Morris et al., 2002) in PCI to the more typical sea-ice bacteria, such as Polaribacter (Figures 5 and 6). The predominance of Gammaproteobacteria and Flavobacteriia has been reported especially in spring/summer during the sea-ice algal ‘bloom’ (Brown and Bowman, 2001; Brinkmeyer et al., 2003; Bowman et al., 2012; Torstensson et al., 2015). However, they can also be predominant at other times of the year, for example, the Austral autumn, if algal-derived substrate is available (Brinkmeyer et al., 2003; Eronen-Rasimus et al., 2014). Thus, it is likely that these copiotrophic bacteria are present in Antarctic sea ice throughout the winter and able to maintain their activity, if the chl-a concentrations are high enough and physical conditions favourable. This is in line with an earlier culture-based study in the Weddell Sea ice (Helmke and Weyland, 1995), which showed the proportion of psychrophiles in bacterial community increasing with the age of the ice. The results also show that the wintertime bacterial community dynamics in Antarctic sea ice differ from those of the oligotrophic Alphaproteobacteria (SAR11 clade)-dominated winter Arctic sea ice (Collins et al., 2010).
Class (on the left) and genus (on the right) level bacterial diversity of 16S ribosomal RNA (rRNA) gene sequences (~450 bp) representing >0.1% of all normalised operational taxonomic units (OTUs) derived from the Weddell Sea ice. T=top ice, U=upper intermediate ice, M=middle ice, L=lower intermediate ice and B=bottom ice. Plot created with ggplot2 (v. 2.1.0.; Wickham, 2009).
Ternary diagram (ggtern, v2.1.1; Hamilton, 2016) of 16S ribosomal RNA (rRNA) gene sequences (~450 bp) representing >0.1% of all normalised operational taxonomic units (OTUs) in each ice type (PCI, FYI and SYI) collected from the Weddell Sea, Antarctica.
In addition to these common sea-ice bacterial classes, the ANOX ice samples also showed a predominance of atypical sea-ice bacterial classes, such as Deltaproteobacteria (for example, genera Desulforhopalus and Desulfofrigus), Epsilonproteobacteria (for example, genera Sulfurospirillum and Arcobacter) and Bacteroidia (for example, genus Marinifilum; Figure 5). ANOX likely formed because of the entrapment of dense algal accumulations between rafted ice floes, where subsequent bacterial activity may have caused oxygen depletion and the establishment of anaerobic bacterial communities including potential sulphate-reducing bacteria. Based on the detected odour of H2S, the bacteria had been actively growing and reducing sulphur compounds in the ice. There is also previous evidence of transient anoxic conditions and anaerobic reactions such, as denitrification, in the ice (Kaartokallio, 2001; Rysgaard and Glud, 2004; Rysgaard et al., 2008). Based on our data, we cannot conclude how commonly anaerobic bacteria occur in the Antarctic winter-pack ice or deduce the origin of the anaerobic bacterial community. However, the presented data suggest that, under specific conditions, anaerobic bacteria may become locally predominant in sea ice.
Conclusions
Our study is the first of its kind to describe the wintertime bacterial community dynamics in Antarctic sea ice and show that under suitable conditions anaerobic bacteria may become predominant in the sea-ice bacterial community. The thick insulating snow cover warmed the ice sustaining high chl-a concentrations. In general, common sea-ice bacterial genera, for example, Octadecabacter (Alphaproteobacteria), Polaribacter (Flavobacteriia) and Glaciecola (Gammaproteobacteria), usually common in spring and summer sea ice, predominated in the communities. The bacterial community structure, abundance and activity were driven by chl-a, suggesting that permeable ice with associated high algal biomass sustain bacterial activity during the Antarctic winter. In addition, to the common sea-ice bacterial classes, predominance of sulphate-reducing bacteria (for example, Desulforhopalus, Desulfofrigus and Sulfurospirillum) was observed in the ANOX together with the odour of H2S suggesting that under suitable conditions sulphur compounds may be reduced in sea ice. In all, the results suggest that sea-ice bacterial communities can remain dynamic throughout the winter if physical conditions are favourable and the chl-a concentrations high enough. Thus, the possible future warming of sea ice and consequent increase in bacterial production may induce changes in major bacteria-mediated biogeochemical processes in the Antarctic sea-ice zone.
References
Anderson MJ . (2001). A new method for non-parametric multivariate analysis of variance. Austral Ecol 26: 32–46.
Anderson MJ . (2006). Distance-based test for homogeneity of multivariate dispersion. Biometrics 62: 245–253.
Anderson MJ, Willis TJ . (2003). Canonical analysis of principal coordinates: a useful method of constrained ordination for ecology. Ecology 84: 511–525.
Anderson MJ, Gorley RN, Clarke KR . (2008) PERMANOVA+ for PRIMER: Guide to Software and Statistical Methods. Primer-E Ltd: Plymouth, UK.
Arrigo KR . (2014). Sea ice ecosystems. Ann Rev Mar Sci 6: 439–467.
Arrigo KR, Mock T, Lizotte MP . (2010) Primary producers in sea ice. In: Thomas DN, Dieckmann GS (eds). Sea Ice2nd ednWiley-Blackwell Publishing: Oxford, UK, 283–325.
Bowman JP, McCammon SA, Brown MV, Nichols DS, McMeekin TA . (1997). Diversity and as- sociation of psychrophilic bacteria in Antarctic sea ice. Appl Environ Microb 63: 3068–3078.
Bowman JS, Rasmussen S, Blom N, Deming JW, Rysgaard S, Sicheritz-Ponten T . (2012). Microbial community structure of Arctic multiyear sea ice and surface seawater by 454 sequencing of the 16 S RNA gene. ISME J 6: 11–20.
Brinkmeyer R, Knittel K, Jurgens J, Weyland H, Amann R, Helmke E . (2003). Diversity and structure of bacterial communities in Arctic versus Antarctic pack ice. Appl Environ Microb 69: 6610–6619.
Brown MV, Bowman JP . (2001). A molecular phylogenetic survey of sea-ice microbial communities (SIMCO). FEMS Microbiol Ecol 35: 267–275.
Chung J, Ha ES, Park HR, Kim S . (2004). Isolation and characterization of Lactobacillus species inhibiting the formation of Streptococcus mutans biofilm. Oral Microbiol Immun 19: 214–216.
Clarke KR, Gorley RN . (2006) Primer v6: User Manual/Tutorial. Primer-E Ltd: Plymouth, UK.
Clarke KR, Warwick RM . (2001). Change in marine communities: an approach to statistical analysis and interpretation, 2nd edition. PRIMER-E: Plymouth, UK.
Collins RE, Carpenter SD, Deming JW . (2008). Spatial heterogeneity and temporal dynamics of particles, bacteria, and pEPS in Arctic winter sea ice. J Marine Syst 74: 902–917.
Collins RE, Rocap G, Deming JW . (2010). Persistence of bacterial and archaeal communities in sea ice through an Arctic winter. Environ Microbiol 12: 1828–1841.
Collins RE . (2015) Microbial Evolution under Extreme Conditions Bakermans C (Ed.). In: Microbial Evolution under Extreme Conditions Vol. 2. Walter de Gruyter GmbH & Co KG: Göttingen, Germany.
Comiso J . (2010) Variability and trends of the global sea ice cover. In: Thomas DN, Dieckmann GS (eds). Sea Ice, 2nd edn. Wiley-Blackwell Publishing: Oxford, UK, 205–246.
Cowie RO, Williams GJ, Maas EW, Voyles KM, Ryan KG . (2014). Antarctic sea-ice microbial communities show distinct patterns of zonation in response to algal-derived substrates. Aquat Microb Ecol 73: 123–134.
Delille D, Fiala M, Kuparinen J, Kuosa H, Plessis C . (2002). Seasonal changes in microbial biomass in the first-year ice of the Terre Adélie area (Antarctica). Aquat Microb Ecol 28: 257–265.
Deming JW . (2010) Sea ice bacteria and viruses. In: Thomas DN, Dieckmann GS (eds). Sea Ice. 2nd edn, Wiley-Blackwell Publishing: Oxford, UK, 247–282.
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R . (2011). UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27: 2194–2200.
Edgar RC . (2013). UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10: 996–998.
Edwards U, Rogall T, Blocker H, Emde M, Bottger E . (1989). Isolation and direct complete nucleotide determination of entire genes. Characterization of a gene coding for 16 S ribosomal RNA. Nucleic Acids Res 17: 7843–7853.
Eicken H, Lange MA, Wadhams P . (1994). Characteristics and distribution patterns of snow and meteoric ice in the Weddell Sea and their contribution to the mass balance of sea ice. Ann Geophysicae 12: 80–93.
Eronen-Rasimus E, Kaartokallio H, Lyra C, Autio R, Kuosa H, Dieckmann G et al. (2014). Bacterial community dynamics and activity in relation to dis- solved organic matter availability during sea-ice formation in a mesocosm experiment. MicrobiologyOpen 3: 139–156.
Eronen-Rasimus E, Lyra C, Rintala JM, Jurgens K, Ikonen V, Kaartokallio H . (2015). Ice formation and growth shape bacterial community structure in Baltic Sea drift ice. FEMS Microbiol Ecol 9: 1–9.
Eronen-Rasimus E, Piiparinen J, Karkman A, Lyra C, Gerland S, Kaartokallio H . (2016). Bacterial communities in Arctic first-year drift ice during the winter/spring transition. Environ Microbiol Rep 8: 527–535.
Fritsen CH, Lytle VI, Ackley SF, Sullivan CW . (1994). Autumn bloom of Antarctic pack-ice algae. Science 266: 782–784.
Fuhrman JA, Azam F . (1980). Bacterioplankton secondary production estimates for coastal waters of British Columbia, Antarctica, and California. Appl Environ Microbiol 39: 1085–1095.
Fuhrman JA, Azam F . (1982). Thymidine incorporation as a measure of heterotrophic bacterioplankton production in marine surface waters: evaluation and field results. Mar Biol 66: 109–120.
Garrison DL, Ackley SF, Buck KR . (1983). A physical mechanism for establishing algal populations in frazil ice. Nature 306: 363–365.
Gasol JM, Del Giorgio PA . (2000). Using flow cytometry for counting natural planktonic bacteria and understanding the structure of planktonic bacterial communities. Sci Mar 64: 197–224.
Golden KM, Ackley SF, Lytle VI . (1998). The percolation phase transition in sea ice. Science 282: 2238–2241.
Grossmann S, Dieckmann GS . (1994). Bacterial standing stock, activity, and carbon production during formation and growth of sea ice in Weddell Sea, Antarctica. Appl Environ Microb 60: 2746–2753.
Haas C . (2010) Dynamics versus Thermodynamics: the sea ice thickness distribution. In: Thomas DN, Dieckmann GS (eds). Sea Ice, 2nd edn. Wiley-Blackwell Publishing: Oxford, UK, 113–151.
Hamilton N . (2016). ggtern: An Extension to 'ggplot2', for the Creation of Ternary Diagrams. R package version 2.1.1 https://CRAN.R-project.org/package=ggtern.
Hatam I, Charchuk R, Lange B, Beckers J, Haas C, Lanoil B . (2014). Distinct bacterial assemblages reside at different depths in Arctic multiyear sea ice. FEMS Microbiol Ecol 90: 115–125.
Hatam I, Lange B, Beckers J, Haas C, Lanoil B . (2016). Bacterial communities from Arctic seasonal sea ice are more compositionally variable than those from multi-year sea ice. ISME J 10: 1–10.
Helmke E, Weyland H . (1995). Bacteria in sea ice and underlying water of the eastern Weddel sea in midwinter. Mar Ecol Prog Ser 117: 269–287.
IPCC. (2013). Summary for Policymakers. In Climate Change 2013: The Physical Science Basis. Contribution of Working Group I to the Fifth Assessment Report of the Intergovernmental Panel on Climate Change Stocker, Qin TF, Plattner D, Tignor G-K, Allen M, Boschung SK, Nauels J, Xia A, Bex VY, Midgley PM (eds) Cambridge University Press: Cambridge, UK, pp 1–30.
Junge K, Imhoff F, Staley T, Deming JW . (2002). Phylogenetic diversity of numerically important Arctic sea-ice bacteria cultured at sub- zero temperature. Microb Ecol 43: 315–328.
Kaartokallio H . (2001). Evidence for active microbial nitrogen transformations in sea ice (Gulf of Bothnia, Baltic Sea) in midwinter. Polar Biol 24: 21–28.
Kaartokallio H . (2004). Food web components, and physical and chemical properties of Baltic Sea ice. Mar Ecol Prog Ser 273: 49–63.
Kaartokallio H, Tuomainen J, Kuosa H, Kuparinen J, Martikainen PJ, Servomaa K . (2008). Succession of sea-ice bacterial communities in the Baltic Sea fast ice. Polar Biol 31: 783–793.
Kottmeier ST, Grossi SM, Sullivan CW . (1987). Sea ice microbial communities. VIII. Bacterial production in annual sea ice of McMurdo sound, Antarctica. Mar Ecol Prog Ser 35: 175–186.
Kuosa H, Kaartokallio H . (2006). Experimental evidence on nutrient and substrate limitation of Baltic sea sea-ice algae and bacteria. Hydrobiologia 554: 1–10.
Lange MA, Ackley SF, Wadhams P, Dieckmann GS, Eicken H . (1989). Development of sea ice in the Weddell Sea. Ann Glaciol 12: 92–96.
Lange MA, Schlosser P, Ackley SF, Wadhams P, Dieckmann GS . (1990). 18O concentrations in sea ice of the Weddell Sea, Antarctica. J Glaciol 36: 315–323.
Lannuzel D, De Jong J, Schoemann V, Trevena A, Tison JL, Chou L . (2006). Development of a sampling and flow injection analysis technique for iron determination in the sea ice environment. Anal Chim Acta 556: 476–483.
Lannuzel D, Schoemann V, De Jong J, Tison JL, Chou L . (2007). Distribution and biogeochemical behaviour of iron in the East Antarctic sea ice. Mar Chem 106: 18–32.
Laurion I, Demers S, Vezina AF . (1995). The microbial food web associated with the ice algal assemblage: biomass and bacterivory of nanoflagellate protozoans in Resolute Passage (High Canadian Arctic). Mar Ecol Prog Ser 120: 77–87.
Manly BFJ . (1997) Randomization, Bootstrap and Monte Carlo Methods in Biology2nd editionChapman and Hall: London.
Martin M . (2011). Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J 17: 10–12.
McArdle BH, Anderson MJ . (2001). Fitting multivariate models to community data: a comment on distance-based redundancy analysis. Ecology 82: 290–297.
Meiners KM, Vancoppenolle M, Thanassekos S, Dieckmann GS, Thomas DN, Tison JL et al. (2012). Chlorophyll a in Antarctic sea ice from historical ice core data. Geophys Res Lett 39: L21602.
Melnikov IA . (1998). Winter production of sea ice algae in the western Weddell Sea. J Marine Syst 17: 195–205.
Mock T, Thomas DN . (2005). Recent advances in sea‐ice microbiology. Environ Microbiol 7: 605–619.
Morris RM, Rappe MS, Connon SA, Vergin KL, Siebold WA, Carlson CA et al. (2002). SAR11 clade dominates ocean surface bacterioplankton communities. Nature 420: 806–810.
NOAA National Centers for Environmental Information, State of the Climate: Global Snow and Ice for February 2017, published online March 2017, retrieved on 21 April 2017 from https://www.ncdc.noaa.gov/sotc/global-snow/201702.
Paulson JN, Stine OC, Bravo HC, Pop M . (2013). Differential abundance analysis for microbial marker-gene surveys. Nat Methods 10: 1200–1202.
Perovich D, Gerland S, Hendricks S, Meier W, Nicolaus M, Tschudi M . (2014), Sea ice. In Arctic Report Card [WWW document]. URL http://www.arctic.noaa.gov/reportcard.
Petrich CP, Eicken H . (2010) Growth, structure and properties of sea ice. In: Thomas DN, Dieckmann GS (eds). Sea Ice, 2nd edn. Wiley-Blackwell Publishing: Oxford, UK, 23–77.
Piiparinen J, Kuosa H . (2011). Impact of UVA radiation on algae and bacteria in Baltic Sea ice. Aquat Microb Ecol 63: 75–87.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P et al. (2013). The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41: D590–D596.
R Development Core Team. (2011) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing: Vienna, Austria, ISBN 3-900051-07-0. Available online at http://www.R-project.org/.
Riedel A, Michel C, Gosselin M . (2007). Grazing of large-sized bacteria by sea-ice heterotrophic protists on the Mackenzie shelf during the winter-spring transition. Aquat Microb Ecol 74: 3–4.
Rysgaard S, Glud RN . (2004). Anaerobic N2 production in Arctic sea ice. Limnol Oceanogr 49: 86–94.
Rysgaard S, Glud RN, Sejr MK, Blicher ME, Stahl HJ . (2008). Denitrification activity and oxygen dynamics in Arctic sea ice. Polar Biol 31: 527–537.
Schlitzer R . (2016), Ocean Data View https://odv.awi.de.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann ME, Hollister B et al. (2009). Introducing Mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75: 7537–7541.
Smith RE, Clement P . (1990). Heterotrophic activity and bacterial productivity in assemblages of microbes from sea ice in the high Arctic. Polar Biol 10: 351–357.
Stewart FJ, Fritsen CH . (2004). Bacteria-algae relationships in Antarctic sea ice. Antarct Sci 16: 143–156.
Stammerjohn SE, Martinson DG, Smith RC, Yuan X, Rind D . (2008). Trends in Antarctic annual sea ice retreat and advance and their relation to El Niño–Southern Oscillation and Southern Annular Mode variability. J Geophys Res-Oceans 113: C3.
Sullivan CW, Palmisano AC . (1984). Sea ice microbial communities: distribution, abundance, and diversity of ice bacteria in McMurdo sound, Antarctica, in 1980. Appl Environ Microbiol 47: 788–795.
Tison J-L, Delille B, de Jong J, Janssens J, Dieckmann G, Gussone N et al. (2014). Sea ice tBiogeochemistry. In: Lemke P (ed). The Expedition of the Research Vessel ‘Polarstern’ to the Antarctic in 2013 (ANT-XXIX/6). Reports on Polar and Marine Research 679, ISSN 1866-3192 http://hdl.handle.net/10013/epic.44124.
Torstensson A, Dinasquet J, Chierici M, Fransson A, Riemann L, Wulff A . (2015). Physicochemical control of bacterial and protist community composition and diversity in Antarctic sea ice. Environ Microbiol 17: 3869–3881.
UNESCO. (1981). Tenth report of the joint panel on oceanographic tables and standards. United Nations Educational, Scientific and Cultural Organization (UNESCO) technical papers in marine science 36.
Wickham H . (2009) ggplot2: Elegant Graphics for Data Analysis. Springer-Verlag: New York.
Zhang J, Kobert K, Flouri T, Stamatakis A . (2014). PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30: 614–620.
Acknowledgements
The work described here was supported by the Walter and Andrée de Nottbeck Foundation (EER, AML, JMR), Academy of Finland (EER) and the Belgian Science Policy (BD, JLT, Bigsouth project, SD/CA/05). BD is a research associate at the F.R.S-FNRS. We thank Finnish Antarctic Research Program (FINNARP) for the logistics and financial support with cargo and travel expenses as well as Professor Katarina Abrahamsson (University of Gothenburg) and her research group for providing samples from the sibling, ANOX ice core, station 515.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Eronen-Rasimus, E., Luhtanen, AM., Rintala, JM. et al. An active bacterial community linked to high chl-a concentrations in Antarctic winter-pack ice and evidence for the development of an anaerobic sea-ice bacterial community. ISME J 11, 2345–2355 (2017). https://doi.org/10.1038/ismej.2017.96
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2017.96
This article is cited by
-
Selection processes of Arctic seasonal glacier snowpack bacterial communities
Microbiome (2023)
-
Nitrogen and sulfur metabolisms encoded in prokaryotic communities associated with sea ice algae
ISME Communications (2023)
-
Monitoring a changing Arctic: Recent advancements in the study of sea ice microbial communities
Ambio (2022)
-
Autumn to spring microbial community in the northern Baltic Sea: temporal variability in bacterial, viral and nanoflagellate abundance during the cold-water season
Polar Biology (2020)
-
The future of Arctic sea-ice biogeochemistry and ice-associated ecosystems
Nature Climate Change (2020)