Abstract
Bacteria face tough competition in polymicrobial communities. To persist in a specific niche, many species produce toxic extracellular effectors to interfere with the growth of nearby microbes. These effectors include the recently reported non-canonical D-amino acids (NCDAAs). In Vibrio cholerae, the causative agent of cholera, NCDAAs control cell wall integrity in stationary phase. Here, an analysis of the composition of the extracellular medium of V. cholerae revealed the unprecedented presence of D-Arg. Compared with other D-amino acids, D-Arg displayed higher potency and broader toxicity in terms of the number of bacterial species affected. Tolerance to D-Arg was associated with mutations in the phosphate transport and chaperone systems, whereas D-Met lethality was suppressed by mutations in cell wall determinants. These observations suggest that NCDAAs target different cellular processes. Finally, even though virtually all Vibrio species are tolerant to D-Arg, only a few can produce this D-amino acid. Indeed, we demonstrate that D-Arg may function as part of a cooperative strategy in vibrio communities to protect non-producing members from competing bacteria. Because NCDAA production is widespread in bacteria, we anticipate that D-Arg is a relevant modulator of microbial subpopulations in diverse ecosystems.
Similar content being viewed by others
Introduction
Bacteria live in polymicrobial communities characterized by a great diversity of co-existing species and competition for available resources. Unearthing the molecular mechanisms that shape microbial communities and their dynamics is a major challenge of the post-genomic era. One bacterial strategy for persisting in a specific niche is the production of toxic extracellular factors that interfere with the growth and/or viability of nearby microbes. These ‘bacteriocins’ disrupt the equilibrium among bacterial populations in a phenomenon known as dysbiosis, which can ultimately alter the homeostasis of different host ecosystems, such as humans and plants (Riley and Wertz, 2002). In the 1940s and 1950s, naturally occurring D-amino acids were first identified. Studies revealed that the addition of high concentrations of D-amino acids to bacterial cultures had a powerful effect on morphogenesis and ultimately caused lysis (Fox et al., 1944; Kobayashi et al., 1948; Yaw and Kakavas, 1952; Lark and Lark, 1959; Grula, 1960; Tuttle and Gest, 1960; Bopp, 1965). The effect of D-amino acids on bacterial shape was fundamental in elucidating their mode of action in the peptidoglycan (PG) polymer (also called murein), a specific bacterial structure with a crucial role in preserving bacterial morphology and cell integrity (Caparros et al., 1992). However, these studies recently gained new physiological meaning when it was reported that many taxonomically unrelated bacteria release millimolar concentrations of non-canonical D-amino acids (NCDAAs) into the environment (Lam et al., 2009). In Vibrio cholerae, the causative agent of the diarrheal disease cholera, the periplasmic broad-spectrum racemase BsrV produces mainly D-Met and D-Leu in stationary phase (Lam et al., 2009; Cava et al., 2011). Remarkably, these NCDAAs are incorporated into the PG by replacing the 4th and/or 5th residue of the disaccharide subunit N-acetylglucosamine-(β1-4)-N-acetylmuramic acid-L-Ala-D-Glu-(γ)-di-amino acid-D-Ala-D-Ala, depending on the bacterial species. Cell wall editing via the incorporation of NCDAAs downregulates PG synthesis, thereby permitting synchronization of cell wall metabolism and growth arrest in stationary phase, which preserves cell wall integrity (Lam et al., 2009; Cava et al., 2011). However, incorporation of NCDAAs is also observed in bacteria that do not produce these molecules. ‘Paracrine’ incorporation of NCDAAs causes morphological aberrations and impairs growth in multiple species.
Although virtually all D-amino acids can potentially modify the cell wall, it is unclear why some bacteria produce distinct sets of NCDAAs (Lam et al., 2009; Cava et al., 2011). Here, we report the extracellular production of D-Arg by V. cholerae. Like D-Met and D-Leu, D-Arg is produced by BsrV. However, compared with other D-amino acids, D-Arg exhibits significantly higher inhibitory activity against a wide diversity of bacterial species, suggesting a role of D-Arg as a fitness modulator of bacterial communities in the natural environment. Based on biochemical analyses of PG, microscopy and transposon sequencing, we demonstrate that in contrast to D-Met, which has a major modulatory role in cell wall biosynthesis, the inhibition of growth by D-Arg appears to be cell wall-independent. D-Arginine toxicity is suppressed by mutations in the DnaJ chaperone system and in the Pst phosphate uptake machinery, strongly supporting different roles for NCDAAs in bacterial physiology.
Finally, we analyze the distribution of BsrVs among the Vibrionaceae family and confirm that although highly conserved, production of D-amino acids is not a hallmark of all vibrios. Synteny and racemase activity-based studies confirmed that BsrV orthologues are absent in a number of Vibrionaceae species. In co-cultivation experiments, V. cholerae outcompeted Caulobacter crescentus by D-Arg production. Thus, NCDAAs production by multi-specific racemases might be a strategy employed by vibrios and, potentially, other species to prevail in competitive environments.
Materials and methods
Bacterial strains and culture conditions
Strains are listed in Supplementary Table S1. All strains were grown under optimal conditions (media and temperature) recommended by the DSMZ, ATCC and CECT bacterial collections as indicated in Supplementary Table S1. V. cholerae strains were grown in LB (Luria Bertani broth) media at 37 °C (Dziejman et al., 2002). The Vibrionaceae species were grown in diluted Marine Broth (Difco) at 22 ºC. Defined V. cholerae transposon mutants were taken from an ordered transposon insertion library (Cameron et al., 2008). For stalk length analyses, C. crescentus strains were grown in Hutner base-imidazole-buffered-glucose-glutamate minimal medium supplemented with 10 or 0.03 mm phosphate (Gonin et al., 2000). Agar 1.5% (w/v) was used in solid plates. When required, antibiotics were added to liquid or solid media: Streptomycin (V. cholerae: 200 μg ml−1), Kanamycin (Escherichia coli and V. cholerae: 50 μg ml−1; Agrobacterium tumefaciens: 300 μg ml−1) and Ampicillin (E. coli: 100 μg ml−1). Media was supplemented with L- or D-amino acid at the indicated concentration when needed.
For growth curves, at least three replicates per strain and condition tested were grown in two independent experiments in 200 μl medium in a 96-well plate inoculated 1:1000 from exponentially growing precultures. Optical density at 600 nm (OD600) was monitored using an Eon Biotek plate reader (Biotek, Winooski, VT, USA) at 10 min intervals and the optimal growth temperature.
Viability assays
Overnight cultures were normalized to 1 unit of optical density at 600 nm (OD600) and subjected to serial 10-fold dilution. Five-microliter drops of the 100 through 10−7 dilutions were then spotted onto the indicated agar plates and incubated at the appropriate temperature for 24–48 h prior to image acquisition.
Bacterial competition assay
A NCDAA producer (V. cholerae) and a NCDAA-sensitive bacteria (C. crescentus) were inoculated in compartments A and B respectively of a Stericup system separated by a 0.22 μm membrane. Both compartments were filled with peptone yeast extract (PYE) media, for optimal growth of C. crescentus, and supplemented or not (control) with 5 mm L-Arg. Compartment A was inoculated with V. cholerae wild-type (lacZ+), ΔbsrV mutant (lacZ−), a 1:1 mix of both or none (-). Compartment B was inoculated with C. crescentus. Devices were incubated at 28 °C with mild-agitation (100 rpm). The 0.22 μm filter avoids bacterial contact while allowing free diffusion of small molecules such as amino acids. Both viability and D-amino acid concentration were measured at different time points (0, 24, 48 and 72 h). Viability was assessed by CFU ml−1 counting on LB (V. cholerae) or PYE (C. crescentus) agar plates. The ratio WT/ΔbsrV in V. cholerae mix was assessed by plating on LB plates containing X-gal 40 μg ml−1, to distinguish between WT (lacZ+, blue) and ΔbsrV (lacZ−, white) colonies. D-amino acid concentration was determined using a D-amino acid oxidase assay (see below). The competition assay was performed twice in triplicates.
PG analysis
PG was isolated and analyzed following previously described methods (Desmarais et al., 2013; Alvarez et al., 2016). In brief, PG sacculi were prepared by boiling bacterial cells in SDS 5%. After SDS removal by ultracentrifugation, the insoluble material was further digested with muramidase (Cellosyl). Soluble muropeptides were separated by liquid chromatography (high-performance liquid chromatography and/or ultra high-pressure liquid chromatography) and identified by MALDI-TOF and electrospray-ion trap MS. A detailed protocol is described in Extended Experimental Procedures.
Total PG amount was determined by quantification of total intensities of the PG profile from three biological replicas normalized to the same OD600 ml−1. Quantification of relative abundances of muropeptides was calculated as the relative area of the corresponding peak compared to the total area of the normalized chromatogram.
D-amino acid analysis and quantification
D-amino acids in the supernatant were derivatized with Marfey’s reagent (Bhushan and Bruckner, 2004) and analyzed by high-performance liquid chromatography as described previously (Espaillat et al., 2014). Total D-amino acid concentration was determined by D-amino acid oxidase assay coupled to peroxidase and 2,3 diaminophenazine (Espaillat et al., 2014). Detailed protocols are described in Extended Experimental Procedures.
Microscopy
Bacteria were immobilized on LB or PYE pads containing 1% agarose. Phase contrast microscopy was performed using a Zeiss Axio Imager.Z2 microscope (Zeiss, Oberkochen, Germany) equipped with a Plan-Apochromat × 63 phase contrast objective lens and an ORCA-Flash 4.0 LT digital CMOS camera (Hamamatsu Photonics, Shizuoka, Japan), using the Zeiss Zen Blue software. Image analysis and processing were performed using Fiji and the MicrobeJ plugin (Schindelin et al., 2012; Ducret et al., 2016). Sphere-inducing activity of D-amino acids was analyzed by cell shape analysis of 380–480 cells per condition.
Transposon insertion sequencing
For the identification of essential genes, transposon insertion sequencing (TnSeq) was performed as described previously (Chao et al., 2013). In brief, 3 × 105–4 × 105 transposon mutants were generated for each library by conjugation of V. cholerae with E. coli SM10λPIR carrying pSC189 (transposon donor plasmid) (Chiang and Rubin, 2002). Mutant libraries were selected under three conditions (control, 5 mm D-Arg and 5 mm D-Met) and pooled genomic DNA fragments were analyzed using a MiSeq sequencer (Illumina, San Diego, CA, USA). Insertion sites were identified and significance was determined using ConArtist simulation-based normalization as described (Chao et al., 2013; Pritchard et al., 2014).
Suppressor mutants
To obtain suppressor mutants, C. crescentus was grown in 2 ml PYE supplemented with 0.5 mm either D-Arg or D-Met. Cultures were grown for 24 h at 28 °C and reinoculated in fresh media containing the corresponding D-amino acid during 12 days (80 generations). After this preadaptation process, 0.1 ml aliquots (~1 × 108 cells) were inoculated onto PYE agar plates containing 1–10 mm D-amino acid. Plates were incubated at 28 °C until suppressor mutant colonies arose. For confirmation of the resistance, the selected derivatives were grown in presence and absence of the corresponding D-amino acid during several generations as described above, prior to viability testing in presence of D-amino acid.
Suppressor mutants of A. tumefaciens were obtained likewise by preadaptation in LB 0.5 mm D-Arg media during 12 days (80 generations) and selection of resistant derivatives on 10 mm D-Arg containing plates.
Whole-genome sequencing and single-nucleotide polymorphism analysis
Genomic DNA samples from suppressor mutants and parental strain were prepared. Indexed paired-end libraries were constructed and sequenced in a MiSeq sequencer (Illumina), following the manufacturer’s instructions. The sequences were analyzed using the Galaxy server tools (https://usegalaxy.org/, (Afgan et al., 2016)) for identification of variants (single-nucleotide polymorphisms). A detailed protocol is described in Extended Experimental Procedures.
Reconstruction of suppressor mutant pstBM108T in A. tumefaciens
For reconstruction of the point mutation pstBM108T in A. tumefaciens, a 1000 bp fragment containing the mutation site was amplified from purified genomic DNA with primers FCP2041 (5'-AAAAAAGCTTTCCGACAACGCCGATGGA-3') and FCP2044 (5'-AAAAAAGCTTGAGGCACCAGTCTTGTTC-3') and cloned into plasmid pNPTS139 (Fischer et al., 2002). E. coli DH5α λPIR was used in the cloning step and the resulting plasmid pNPTS139pstBM108T was verified by sequencing.
Nucleotide substitution in A. tumefaciens pstB gene (Atu0423) was performed following an established allelic-replacement protocol (Morton and Fuqua, 2012). In short, exconjugants obtained by conjugation with E. coli S17-1 λPIR cells as a donor of pNPTS139pstBM108T were selected on ATGN plates containing Kanamycin 300 μg/ml. Exconjugants were grown in ATGN medium overnight and then plated on ATSN plates containing 5% (w/v) sucrose. Colonies sensitive to kanamycin were streak-purified twice on ATSN plates and checked by sequencing.
Results
V. cholerae releases high concentrations of D-arginine to the extracellular medium
An exhaustive chemical analysis of the stationary-phase extracellular medium of V. cholerae revealed that this bacterium releases a greater diversity of D-amino acids than previously reported (Figure 1a). In addition to D-Met and D-Leu (Lam et al., 2009), we detected D-Gln, D-Cys, D-Lys and D-Arg, whose identities were confirmed by supplementing the samples with pure D-amino acid standards (for example, D-Arg, Supplementary Figure S1a). Using a D-amino acid oxidase activity-based assay, we determined that V. cholerae releases approximately 2.5 mm of D-amino acids (Figure 1b), including ~0.7 mm D-Arg.
D-arginine is released to the extracellular medium. (a) HPLC profile of Marfey’s derivatized preconditioned medium (PCM) from stationary-phase V. cholerae cultures for quantification of chiral amino acids. L- and D-forms were identified by comparison to standards. (b) Total D-amino acid content in wild-type and ΔbsrV mutant V. cholerae cultures, determined by D-amino acid oxidase assay. P-value <0.0001 (***). (c, d) HPLC analysis of Marfey’s derivatized PCM from wild-type (c) or ΔbsrV mutant (d) V. cholerae stationary-phase cultures. (e) Cell wall analysis of stationary-phase E. coli cultures grown in LB or the PCM from V. cholerae (Vc-PCM). Released D-Arg in Vc-PCM is incorporated into E.coli’s peptidoglycan pentapeptides (M5R). (f) Accumulation of M5R in wild-type V. cholerae, ΔbsrV mutant, E. coli and B. subtilis grown in LB and supplemented with 5 mm L-Arg or D-Arg. P-value <0.0001 (***).
D-Met production by V. cholerae relies on the broad-spectrum racemase BsrV (Lam et al., 2009; Espaillat et al., 2014). To determine whether this enzyme is also responsible for D-Arg production in vivo, we analyzed the D-amino acid composition of the extracellular medium of ΔbsrV (Figures 1c and d,Supplementary Figure S1b). D-Arg was not detected in the ΔbsrV supernatant, thus confirming the role of BsrV in extracellular D-Arg accumulation. As expected, deletion of bsrV impaired production of not only D-Arg but also other D-amino acids (for example, D-Leu) (Figure 1b, Supplementary Figure S1b).
Incorporation of D-arginine in the PG of V. cholerae
We previously reported that incorporation of NCDAAs into the muropeptides can modulate PG biogenesis (Lam et al., 2009; Cava et al., 2011). Indeed, high-performance liquid chromatography analysis of the PG of E. coli grown in V. cholerae stationary-phase preconditioned medium (Figure 1e) revealed an additional peak identified by MALDI-TOF as a disaccharide-pentapeptide ending in D-Arg (M5R: N-acetylglucosamine-(β1-4)-N-acetylmuramic acid-L-Ala-D-Glu-(γ)-mesoDAP-D-Ala-D-Arg) (Supplementary Figure S2a, b). To further confirm the incorporation of D-Arg into the cell wall, we spiked bacterial cultures with exogenous L- or D-Arg (5 mm) and monitored M5R levels. As shown in Figure 1f and Supplementary Figure 2c, supplementation with either enantiomer increased M5R 3.5-fold in wild-type V. cholerae. However, in non-producers (that is, V. cholerae ΔbsrV, E. coli or B. subtilis), M5R was only detected upon supplementation with D-Arg (Figure 1f, Supplementary Figure S2c) confirming that only the D-form of arginine can be used as substrate to edit the cell wall.
D-arginine does not regulate cell wall synthesis in V. cholerae
To determine whether, similar to D-Met, D-Arg downregulates cell wall biosynthesis in V. cholerae (Lam et al., 2009), we compared the effects of these D-amino acids on the growth and PG density of V. cholerae. Supra-physiological concentrations of D-Met but not D-Arg affected growth (Figures 2a and b), and this effect was fully specific for the D-enantiomer (Supplementary Figure S3a). The genes involved in the tolerance to sublethal concentrations of D-Arg and/or D-Met in V. cholerae were then identified by a transposon sequencing-based strategy. Although single mutations in the cell wall-associated loci mrcA, lpoA and csiV were lethal in the presence of D-Met, no mutations conferring sensitivity to D-Arg were observed (Figures 2c–e). These data support our previous findings (Dorr et al., 2014) and also add the DD-carboxypeptidase dacA1 and the aspartate kinase vc0547, the first enzyme in the branched biosynthetic pathway of methionine, lysine and threonine, as genetic determinants of D-Met tolerance in V. cholerae. In agreement with these data, D-Met treated cultures contained less PG than non-treated samples, whereas the PG density of D-Arg treated cells did not differ from that of the control (Figure 2f). Consistent with these observations, D-Met but not D-Arg triggered a morphological transition from curved rod-to-sphere in the mrcA strain, a PG synthesis mutant with reduced PG density compared with wild-type (Lam et al., 2009) (Supplementary Figure S3b, c). Indeed, the higher cell wall inhibitory activity of D-Met compared with D-Arg in V. cholerae correlated with its higher incorporation into the PG (Figure 2g,Supplementary Figure S4). Collectively, these data suggest that the inability of D-Arg to affect cell wall synthesis is owing to its reduced prevalence in the PG.
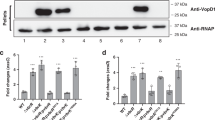
D-arginine does not regulate cell wall synthesis in V. cholerae. (a) Growth curves of wild-type V. cholerae in LB in the absence (control) and presence of 20 mm D-Arg or D-Met. (b) Growth of wild-type V. cholerae on D-amino acid containing plates. Serial dilutions were spotted onto LB plates containing 5–30 mm D-Arg or D-Met. (c, d) Volcano plots depicting the ratio of read counts mapped to individual genes in transposon libraries of V. cholerae plated onto 5 mm D-Arg (c) or D-Met (d) plates compared with control libraries plated onto LB agar without D-amino acid. Genes shown in red (cell wall-associated proteins) and green (amino acid biosynthesis protein) are considered significantly underrepresented. (e) Validation of plating defects of selected V. cholerae mutants identified in the transposon screen. Serial dilutions were spotted onto LB plates supplemented with 5 mm D-Arg or D-Met. (f) Peptidoglycan quantification in D-Arg and D-Met supplemented V. cholerae cultures. Peptidoglycan relative amount (%) is normalized to the wild-type in LB control media. P-value <0.05 (*). P-value <0.01 (**). (g) D-amino acid incorporation into the cell wall of V. cholerae upon addition of different concentrations of D-Arg or D-Met. Relative abundance (%) of D-Arg or D-Met modified muropeptides is represented.
D-arginine inhibits the growth of a wide diversity of bacteria
Our findings demonstrate that V. cholerae releases a cocktail of diverse D-amino acids whose specific functions remain to be elucidated. To determine whether D-amino acids have distinct effects, we compared the effects of D-Arg and other D-amino acids on the growth of a number of representative species belonging to the Firmicutes and Proteobacteria phyla. Final concentrations of 20, 10 and 5 mm provided a good range of responses, even in closely related species. Interestingly, distinct sensitivity profiles were observed, as shown in Figure 3. In general, Alphaproteobacteria were the most sensitive Proteobacteria species: Caulobacterales, Rhizobiales and Rhodobacterales were sensitive to all D-amino acids tested. Conversely, Burkholderia species were fairly resistant even at the highest concentrations used. Although growth inhibition was clearly dependent on the species and the type of D-amino acid, D-Arg exhibited the greatest inhibition in terms of specific activity and range of species affected (Figure 3, Supplementary Figure S5). Remarkably, even bacteria highly resistant to antibiotics, such as Burkholderia, were sensitive to D-Arg. These results suggest that different D-amino acids either have different molecular targets or significantly different potencies on the same target.
Sensitivity of Firmicutes and Proteobacteria to D-amino acids. Bacteria were grown in presence of 20 mm D-amino acids produced by V. cholerae and other bacteria. Relative OD600 is represented in gray shades. Bacteria were assigned to different categories according to their sensitivities (yellow, broadly sensitive bacteria; light blue, D-Arg sensitive; dark blue, resistant bacteria). A: alanine, R: arginine, M: methionine, S: serine, H: histidine, Q: glutamine, F: phenylalanine.
Mutations in the phosphate uptake and DnaJ chaperone systems suppress D-Arg lethality
To identify and compare the molecular targets of D-Arg and D-Met, we screened suppressor mutants of C. crescentus, a bacterium sensitive to these D-amino acids (Figures 4a and b). Seven derivatives resistant to each D-amino acid were selected for whole-genome sequencing and single-nucleotide polymorphism analyses (Figure 4c). The mutations that suppressed lethality differed for each D-amino acid (Figures 4d and e, Supplementary Table S2, S3). As expected, D-Met-resistant derivatives exhibited mutations in cell wall enzymes such as PG lytic transglycosylases, endopeptidases and a putative beta-lactamase (Figure 4d, Supplementary Table S2). Notably, mutations in the slt gene (CCNA_01393) were observed in four of the seven derivatives analyzed, whereas mutations in two putative membrane lytic transglycosylases (MltA-like and MltG-like) were detected in two other derivatives, underscoring the importance of PG turnover in D-Met tolerance (Figure 4d). Conversely, D-Arg-resistant derivatives exhibited mutations in either the dnaJ chaperone or the phosphate uptake system (pstCAB operon) (Figure 4e, Supplementary Table S3). D-Arg promoted filamentation in C. crescentus (Supplementary Figure S6a), but the D-Arg-resistant mutants retained the filamentous phenotype while recovering fitness (Figure 4c), further supporting that D-Arg targets pathways other than cell wall metabolism. Consistently, the D-Arg-resistant mutants and the control contained similar amounts of modified muropeptides, whereas the D-Met-resistant mutants contained significantly fewer modified muropeptides (Supplementary Figure S6b, c). Finally, as no suppressor mutations conferring resistance to D-Arg were detected in cell wall-related genes, these results collectively demonstrate that different D-amino acids target distinct cellular processes.
Mutations in the Pi uptake and protein folding systems suppress D-arginine lethality in sensitive bacteria. (a, b) Growth curves of wild-type C. crescentus in PYE in the absence (control) and presence of 2.5 or 5 mm D-Met (a) or D-Arg (b). (c) Relative growth of C. crescentus suppressor mutants isolated from D-Met containing plates (left, M1–M7) and from D-Arg containing plates (right, R1-R7). Bacteria were grown in PYE media without D-amino acid (control, white bars) or supplemented with 5 mm D-Met (dark gray bars) or 3 mm D-Arg (light gray bars). (d) COG classification of the mutations found in the genomes of D-Met-resistant derivatives of C. crescentus. Mutations conferring resistance to D-Met map to the lytic transglycosylases slt (CCNA_01393), mltA-like (CCNA_01181) and mltG-like (CCNA_01751) genes. (e) COG classification of the mutations found in the genomes of D-Arg-resistant derivatives of C. crescentus. Mutations conferring resistance to D-Arg mapped to the dnaJ gene (CCNA_00011) or the pstCAB operon (CCNA_00292-00294. (f) Suppressor mutation of pstB gene (atu0423) in a D-Arg-resistant A. tumefaciens derivative. (g) A. tumefaciens growth defect in the presence of 10 mm D-Arg is alleviated by a point mutation in PstB (pstBM108T) both in liquid and solid media. COG groups: P: inorganic ion transport and metabolism; O: post-translational modification, protein turnover, chaperone functions; H: coenzyme metabolism and transport; J: translation; E: amino acid metabolism and transport; M: cell wall/membrane/envelope biogenesis; S: unknown function; K: transcription; G: carbohydrate metabolism and transport; C: energy production and conversion; L: replication and repair; N: cell motility.
To explore whether these mechanisms of resistance to D-Arg are specific to C. crescentus, we performed similar experiments with Agrobacterium tumefaciens, which is also sensitive to D-Arg. Remarkably, the phosphate transport system was also mutated in A. tumefaciens derivatives resistant to D-Arg. We reconstructed the pstB M108T point mutation and confirmed that it was sufficient to restore the growth of A. tumefaciens on both liquid and solid media containing 10 mm D-Arg (Figures 4f and g). These data confirm the role of phosphate uptake in D-Arg sensitivity.
Extracellular D-arginine promotes interspecies altruistic cooperation in vibrio communities
The broad toxicity of D-Arg suggests that this D-amino acid might function as an effector to allow vibrios to outcompete other bacteria in the environment. To determine whether resistance to high concentrations of D-Arg is a specific trait of V. cholerae or common to all vibrios, we monitored the relative growth of representative species belonging to different clades of the Vibrionaceae family in the presence of 10 mm D-Arg, D-Met or D-Ala (as a canonical D-amino acid control). All species tested were able to grow normally in the presence of these D-amino acids (Figure 5a). However, resistance to D-amino acids was not linked to D-amino acid production. Comparative genome analysis confirmed that a number of vibrios do not encode BsrV homologs, which was correlated with an absence of detectable concentrations of D-amino acids (Figure 5a).
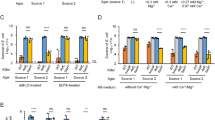
D-arginine mediates interspecies altruistic cooperation within Vibrio communities. (a) Analysis of the Vibrionaceae family growth in MB medium supplemented with 10 mm D-Ala, D-Arg or D-Met. Relative OD600 is represented in gray scale. bsr: comparative genomic analysis of the presence (+) or absence (−) of BsrV homologs. nd: not determined, genome not available. Total D-amino acid content in the supernatant of overnight cultures supplemented with 5 mm L-Arg, determined by D-amino acid oxidase assay. (b) Scheme of the co-cultivation chambers used for bacterial competition assays. Compartment A is inoculated with wild-type V. cholerae, ΔbsrV mutant, a mixture or none. Compartment B is inoculated with C. crescentus. Media is supplemented or not (control) with 5 mm L-Arg. (c) Effect of D-Arg in competition assays. Upper panel: C. crescentus growth, expressed as CFU ml−1. Lower panel: total D-amino acid concentration in the supernatant, determined by D-amino acid oxidase assay.
Because: (i) D-Arg is highly toxic to a wide variety of species; (ii) vibrios are tolerant to D-Arg; (iii) only a few species can produce this D-amino acid; and (iv) diverse vibrio species normally co-exist in the same niche (Tanaka et al., 2004), we hypothesized that D-Arg functions as a protective molecule in vibrio communities. To assess the potential fitness benefit of D-Arg production (by BsrV+ vibrios such as V. cholerae) for BsrV− vibrios, we performed competition experiments in which C. crescentus was co-cultured with either wild-type V. cholerae or the ΔbsrV mutant (Figure 5b). After 48 h, wild-type V. cholerae dramatically interfered with C. crescentus growth (Figure 5c). This result was dependent on the activity of BsrV as no growth inhibition was observed when C. crescentus was co-cultured with the ΔbsrV mutant (Figure 5c). To determine whether BsrV− species could benefit from paracrine D-Arg production, we performed another co-culture experiment in which a mixed population of wild-type and ΔbsrV V. cholerae cells competed with C. crescentus as above (Figure 5c). The subpopulation of BsrV+ vibrios again interfered with the growth of C. crescentus, and the relative proportions of the wild-type and ΔbsrV subpopulations remained fairly constant throughout the experiment (Supplementary Figure S7).
Collectively, these data highlight the potential of D-Arg to shape the biodiversity of an ecological niche by affecting the fitness and survival of co-existing microbial subpopulations (Figure 6).
Model illustrating the impact of extracellular D-arginine in shaping the environmental microbial biodiversity. NCDAAs production (that is, D-Arg), by Bsr+ bacteria works as a powerful bactericidal effector against competing microbes inhabiting the same niche. The inhibitory effect of D-Arg on bacterial growth bias the microbial biodiversity towards the proliferation of tolerant species, even when these organisms do not encode broad-spectrum racemases (Bsr−), establishing new relationships within the community.
Discussion
Microbes interact with the surrounding environment by releasing metabolites. The identification of these important bioactive effectors often relies on functional screenings. In 2009, we determined that V. cholerae produces diverse D-amino acids (mainly D-Met and D-Leu) that modulate cell wall biosynthesis via incorporation into the PG (Lam et al., 2009). These NCDAAs were identified in stationary-phase supernatant active fractions that induced a rod-to-sphere morphological transition of a cell wall-sensitive mutant (that is, mrcA) (Lam et al., 2009). Here, using a non-biased chemical analysis we detected the accumulation of D-Arg at high concentrations in the extracellular medium of V. cholerae. This D-amino acid was not detected in sphere-inducing fractions previously, raising a fundamental question: do all D-amino acids have the same biological role? To address this question, we performed transposon-sequencing experiments to uncover mutants sensitive to D-Met or D-Arg. The experiments with D-Met confirmed our previous data (Dorr et al., 2014) and identified two additional genetic determinants of the tolerance of V. cholerae tolerance to this D-amino acid: dacA1 and aspartate kinase vc0547. DacA1 is a DD-carboxypeptidase responsible for removing the fifth amino-acid residue of the muropeptide. In a dacA1 mutant, pentapeptides accumulate in the PG, resulting in severe growth impairment and altered morphology (Moll et al., 2015). Therefore, the toxicity of D-Met in the dacA1 mutant might be explained by a high accumulation of non-canonical D-Met-pentapeptides that inhibit PG-transpeptidase activities (that is, PBPs), thus further damaging the already weak cell wall of the dacA1 mutant. Accordingly, the innocuous effect of D-Arg on the cell wall may be owing to its lower prevalence in the murein sacculus compared with D-Met (Figure 2). However, we cannot discard the possibility that D-Arg muropeptides are better substrates (less inhibitory) for V. cholerae PBPs than muropeptides containing D-Met, even if these are present at similar levels. The sensitivity of the aspartate kinase mutant to D-Met is somewhat expected because this enzyme catalyzes the first step in the biosynthetic pathway for several amino acids in which diaminopimelic acid (mesoDAP), a constituent of the PG in most gram-negative bacteria, is an intermediate (Viola, 2001).
No conditional lethality was observed in the presence of D-Arg in V. cholerae, indicating that this D-amino acid has no inhibitory effect on the cell wall and, consequently, that different D-amino acids (that is, D-Arg and D-Met) can have distinct functions. In fact, D-Arg acts as a potent bactericidal molecule in many bacterial species (Figure 3). We compared suppressor mutants of C. crescentus that were resistant to either D-Met or D-Arg to obtain deeper insights on the different mechanisms of action of these D-amino acids. Suppression of D-Met inhibition mainly targeted cell wall lytic transglycosylases (Figure 4d, Supplementary Table S2), consistent with the regulatory role of D-Met in cell wall biogenesis. Cell wall growth and integrity rely on carefully coordinated PG synthesis and turnover (Johnson et al., 2013). Alteration of this equilibrium ultimately causes lysis and death. As incorporation of D-Met probably interferes with the synthetic activities of PBPs, it is likely that mutations inactivating the lytic transglycosylases alleviate this effect by decreasing the turnover rate. Conversely, the suppressor mutations against D-Arg lethality in C. crescentus targeted the phosphate uptake and protein folding systems (Figure 4e, Supplementary Table S3). Phosphorus is an essential element for all living organisms, and inorganic phosphate (Pi) is its preferential source. It is unclear why mutations in the Pi transport machinery confer resistance to D-Arg because Pi is central in diverse metabolic and regulatory networks. Inactivating mutations in the pst genes induce a state of continuous Pi starvation. Under these conditions, C. crescentus elongates its stalks to optimize the surface/volume ratio and in turn facilitate Pi uptake (Gonin et al., 2000). Indeed, the Caulobacter suppressors with mutations in the pst genes exhibited longer stalks than the wild-type strain in high-phosphate media, suggesting that these mutations impair Pi transport (Supplementary Figure S6d, e). Unexpectedly, wild-type cells in low-phosphate media did not develop resistance to D-Arg. Therefore, we hypothesize that Pi starvation signaling is not responsible for the prevention of D-Arg toxicity. Although more research is needed to answer this question, the identification of suppressors in this pathway in both C. crescentus and A. tumefaciens strongly supports the role of Pi transport in D-Arg resistance.
The presence of suppressor mutations against D-Arg in dnaJ suggests that D-Arg might target protein synthesis. In proteins, D-amino acids can cause misfolding, aggregation and, ultimately, loss of function (Leiman et al., 2013, 2015). Chaperone systems might help refold or degrade toxic misfolded proteins. Many bacteria possess two main chaperone systems with distinct roles in stress responses: GroES/GroEL and DnaK/DnaJ (Susin et al., 2006). When one system is missing, the accumulation of partially denatured proteins induces an increase in the expression of the other system (Susin et al., 2006). Thus, dnaJ suppressor mutants might lead to an increase in GroES/GroEL to specifically deal with the potential accumulation of misfolded D-Arg-containing proteins. Future research should directly assess the presence of D-Arg in proteins, its effects on protein stability and activity, and whether resistant bacteria (for example, V. cholerae) are less permissive toward the incorporation of D-amino acids in proteins.
The broad-spectrum toxicity of D-Arg makes this amino acid a powerful chemical weapon against microbes inhabiting the same niche. Accordingly, we demonstrated that D-Arg can be used by certain vibrios to outcompete other bacteria. Surprisingly, although all members of the Vibrionaceae family are highly resistant to NCDAAs, broad-spectrum racemases are not widely conserved in this family. Our data suggest that D-Arg could be a public good shared in vibrio communities to promote interspecies altruistic cooperation in challenging polymicrobial environments (Figure 6). This type of alliance, in which certain individuals (cheaters) benefit from the specialization of a few group members, is reminiscent of the phenomenon of bistability (Hibbing et al., 2010; Norman et al., 2015). However, in contrast to bistability, in which a genetically identical subpopulation of the same species exhibits distinct gene expression, interspecies cooperation is based on stable relationships between partner species that are well adapted to the same niche. Vibrio species co-inhabit diverse marine and fresh water niches (Urakawa and Rivera, 2006) and thus can potentially profit from the production of D-Arg. Moreover, production of D-Arg is feasible even when a small proportion of producers is present, and BsrV’s are highly resilient proteins (Espaillat et al., 2014) that retain activity even when added to the extracellular medium after cellular lysis. High concentrations of D-Arg can potentially be produced in the natural environment as L-Arg is abundant in a number of animal and plant sources (Li et al., 2011). Furthermore, although millimolar amounts of D-Arg are readily toxic, micromolar concentrations can influence the growth rate. The effect of D-Arg on fitness/survival can strongly shape the microbial biodiversity and increase the resistance of established communities against invaders. As there are diverse Bsr-producing bacterial species other than vibrios, we anticipate that D-Arg could govern additional microbial social interactions in other environments. The ability of some bacteria to efficiently generate suppressor mutations to circumvent the deleterious effects of D-Arg might explain the production of distinct sets of NCDAAs to target different cellular processes and minimize the emergence of competing microbes. Whether the bactericidal activities of certain NCDAAs can be applied in combinatory antimicrobial therapies remains to be determined.
Finally, the effects of D-Arg suggest that the cell wall might act as bait for environmental chemical prospecting to reveal novel D-amino acid-like metabolites in microbial communities. Further research into the role of NCDAAs in other processes, such as signaling, development and metabolic interference, using controlled multi-species microcosm communities would provide valuable mechanistic insights on the evolution of microbial ecosystems.
References
Afgan E, Baker D, van den Beek M, Blankenberg D, Bouvier D, Cech M et al. (2016). The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2016 update. Nucleic Acids Res 44: W3–W10.
Alvarez L, Hernandez SB, de Pedro MA, Cava F . (2016). Ultra-sensitive, high-resolution liquid chromatography methods for the high-throughput quantitative analysis of bacterial cell wall chemistry and structure. Methods Mol Biol 1440: 11–27.
Bhushan R, Bruckner H . (2004). Marfey's reagent for chiral amino acid analysis: a review. Amino acids 27: 231–247.
Bopp M . (1965). [Inhibition of Agrobacterium tumefaciens by D-amino acids]. Z Naturforsch B 20: 899–905.
Cameron DE, Urbach JM, Mekalanos JJ . (2008). A defined transposon mutant library and its use in identifying motility genes in Vibrio cholerae. Proc Natl Acad Sci USA 105: 8736–8741.
Caparros M, Pisabarro AG, de Pedro MA . (1992). Effect of D-amino acids on structure and synthesis of peptidoglycan in Escherichia coli. J Bacteriol 174: 5549–5559.
Cava F, de Pedro MA, Lam H, Davis BM, Waldor MK . (2011). Distinct pathways for modification of the bacterial cell wall by non-canonical D-amino acids. EMBO J 30: 3442–3453.
Chao MC, Pritchard JR, Zhang YJ, Rubin EJ, Livny J, Davis BM et al. (2013). High-resolution definition of the Vibrio cholerae essential gene set with hidden Markov model-based analyses of transposon-insertion sequencing data. Nucleic Acids Res 41: 9033–9048.
Chiang SL, Rubin EJ . (2002). Construction of a mariner-based transposon for epitope-tagging and genomic targeting. Gene 296: 179–185.
Desmarais SM, De Pedro MA, Cava F, Huang KC . (2013). Peptidoglycan at its peaks: how chromatographic analyses can reveal bacterial cell wall structure and assembly. Mol Microbiol 89: 1–13.
Dorr T, Lam H, Alvarez L, Cava F, Davis BM, Waldor MK . (2014). A novel peptidoglycan binding protein crucial for PBP1A-mediated cell wall biogenesis in Vibrio cholerae. PLoS Genet 10: e1004433.
Ducret A, Quardokus EM, Brun YV . (2016). MicrobeJ, a tool for high throughput bacterial cell detection and quantitative analysis. Nat Microbiol 1: 16077.
Dziejman M, Balon E, Boyd D, Fraser CM, Heidelberg JF, Mekalanos JJ . (2002). Comparative genomic analysis of Vibrio cholerae: genes that correlate with cholera endemic and pandemic disease. Proc Natl Acad Sci USA 99: 1556–1561.
Espaillat A, Carrasco-Lopez C, Bernardo-Garcia N, Pietrosemoli N, Otero LH, Alvarez L et al. (2014). Structural basis for the broad specificity of a new family of amino-acid racemases. Acta Crystallogr D Biol Crystallogr 70: 79–90.
Fischer B, Rummel G, Aldridge P, Jenal U . (2002). The FtsH protease is involved in development, stress response and heat shock control in Caulobacter crescentus. Mol Microbiol 44: 461–478.
Fox S, Fling M, Bollenback N . (1944). Inhibition of bacterial growth by D-leucine. J Biol Chem 155: 465–468.
Gonin M, Quardokus EM, O'Donnol D, Maddock J, Brun YV . (2000). Regulation of stalk elongation by phosphate in Caulobacter crescentus. J Bacteriol 182: 337–347.
Grula EA . (1960). Cell division in a species of Erwinia. I. Inhibition of division by D-amino acids. J Bacteriol 80: 375–385.
Hibbing ME, Fuqua C, Parsek MR, Peterson SB . (2010). Bacterial competition: Surviving and thriving in the microbial jungle. Nat Rev Microbiol 8: 15–25.
Johnson JW, Fisher JF, Mobashery S . (2013). Bacterial cell-wall recycling. Ann N Y Acad Sci 1277: 54–75.
Kobayashi Y, Fling M, Fox SW . (1948). Antipodal specificity in the inhibition of growth of Escherichia coli by amino acids. J Biol Chem 174: 391–398.
Lam H, Oh DC, Cava F, Takacs CN, Clardy J, de Pedro MA et al. (2009). D-amino acids govern stationary phase cell wall remodeling in bacteria. Science 325: 1552–1555.
Lark C, Lark KG . (1959). The effects of D-amino acids on Alcaligenes fecalis. Can J Microbiol 5: 369–379.
Leiman SA, May JM, Lebar MD, Kahne D, Kolter R, Losick R . (2013). D-amino acids indirectly inhibit biofilm formation in Bacillus subtilis by interfering with protein synthesis. J Bacteriol 195: 5391–5395.
Leiman SA, Richardson C, Foulston L, Elsholz AK, First EA, Losick R . (2015). Identification and characterization of mutations conferring resistance to D-amino acids in Bacillus subtilis. J Bacteriol 197: 1632–1639.
Li X, Rezaei R, Li P, Wu G . (2011). Composition of amino acids in feed ingredients for animal diets. Amino Acids 40: 1159–1168.
Moll A, Dorr T, Alvarez L, Davis BM, Cava F, Waldor MK . (2015). A D, D-carboxypeptidase is required for Vibrio cholerae halotolerance. Environ Microbiol 17: 527–540.
Morton ER, Fuqua C . (2012). Genetic manipulation of agrobacterium. Curr Protoc Microb Chapter 3: Unit 3D 2.
Norman TM, Lord ND, Paulsson J, Losick R . (2015). Stochastic switching of cell fate in microbes. Annu Rev Microbiol 69: 381–403.
Pritchard JR, Chao MC, Abel S, Davis BM, Baranowski C, Zhang YJ et al. (2014). ARTIST: high-resolution genome-wide assessment of fitness using transposon-insertion sequencing. PLoS Genet 10: e1004782.
Riley MA, Wertz JE . (2002). Bacteriocins: evolution, ecology, and application. Annu Rev Microbiol 56: 117–137.
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T et al. (2012). Fiji: an open-source platform for biological-image analysis. Nat Methods 9: 676–682.
Susin MF, Baldini RL, Gueiros-Filho F, Gomes SL . (2006). GroES/GroEL and DnaK/DnaJ have distinct roles in stress responses and during cell cycle progression in Caulobacter crescentus. J Bacteriol 188: 8044–8053.
Tanaka R, Ootsubo M, Sawabe T, Ezura Y, Tajima K . (2004). Biodiversity and in situ abundance of gut microflora of abalone (Haliotis discus hannai) determined by culture-independent techniques. Aquaculture 241: 453–463.
Tuttle AL, Gest H . (1960). Induction of morphological aberrations in Rhodospirillum rubrum by D-amino acids. J Bacteriol 79: 213–216.
Urakawa H, Rivera I . (2006) Aquatic Environment. In The Biology of Vibrios. Thompson F, Austin B, Swings J (eds). 12: pp 175–189, ASM Press: Washington, DC.
Viola RE . (2001). The central enzymes of the aspartate family of amino acid biosynthesis. Acc Chem Res 34: 339–349.
Yaw KE, Kakavas JC . (1952). Studies on the effects of D-amino acids on Brucella abortus. J Bacteriol 63: 263–268.
Acknowledgements
We thank Pilar Horcajo, Manon Bruyant and Akbar Espaillat for helping with data acquisition and all Cava lab members for their support and feedback. We thank Frédérique Le Roux and Bruno Gomez-Gil for kindly providing Vibrionaceae strains, and other collaborators for providing different strains. We also thank NBIS (National Bioinformatics Infrastructure Sweden), especially Jeanette Tångrot, for helping in the sequence analysis of suppressor mutants. Research in the Cava lab is funded by The Knut and Alice Wallenberg Foundation (KAW), The Laboratory of Molecular Infection Medicine Sweden (MIMS), the Swedish Research Council and the Kempe Foundation. LA and AA are supported by the Kempe Foundation and KAW.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Alvarez, L., Aliashkevich, A., de Pedro, M. et al. Bacterial secretion of D-arginine controls environmental microbial biodiversity. ISME J 12, 438–450 (2018). https://doi.org/10.1038/ismej.2017.176
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2017.176
This article is cited by
-
d-amino acids signal a stress-dependent run-away response in Vibrio cholerae
Nature Microbiology (2023)
-
d-amino acids involved in stress-induced response in Vibrio cholerae
Nature Microbiology (2023)
-
Shared features and reciprocal complementation of the Chlamydomonas and Arabidopsis microbiota
Nature Communications (2022)
-
Stimulation of root growth and enhanced nitrogenous metabolite content in habanero pepper (Capsicum chinense Jacq.) treated with a d-amino acid mixture
Theoretical and Experimental Plant Physiology (2020)
-
Influence of Phragmites communis and Zizania aquatica on rhizosphere soil enzyme activity and bacterial community structure in a surface flow constructed wetland treating secondary domestic effluent in China
Environmental Science and Pollution Research (2020)