Abstract
Decline-diseases are complex and becoming increasingly problematic to tree health globally. Acute Oak Decline (AOD) is characterized by necrotic stem lesions and galleries of the bark-boring beetle, Agrilus biguttatus, and represents a serious threat to oak. Although multiple novel bacterial species and Agrilus galleries are associated with AOD lesions, the causative agent(s) are unknown. The AOD pathosystem therefore provides an ideal model for a systems-based research approach to address our hypothesis that AOD lesions are caused by a polymicrobial complex. Here we show that three bacterial species, Brenneria goodwinii, Gibbsiella quercinecans and Rahnella victoriana, are consistently abundant in the lesion microbiome and possess virulence genes used by canonical phytopathogens that are expressed in AOD lesions. Individual and polyspecies inoculations on oak logs and trees demonstrated that B. goodwinii and G. quercinecans cause tissue necrosis and, in combination with A. biguttatus, produce the diagnostic symptoms of AOD. We have proved a polybacterial cause of AOD lesions, providing new insights into polymicrobial interactions and tree disease. This work presents a novel conceptual and methodological template for adapting Koch’s postulates to address the role of microbial communities in disease.
Similar content being viewed by others
Introduction
Trees are essential to landscape function and aesthetics, supporting diverse ecologies (Rackham, 2008) and providing key ecosystem services (Boyd et al., 2013). However, significant areas of forest have been lost due to increasing outbreaks of disease and pest attack, and tree health is a current global concern (Cohen et al., 2016). Tree diseases, including decline-diseases, are rising in profile due to an increased risk of introduction and spread through international plant trade and amplification effects of current and future climate change (McDowell et al., 2011; Oliva et al., 2014; Millar and Stephenson, 2015). Emerging evidence suggests that complex biotic interactions, including polymicrobial and insect activity, affect disease occurrence and severity (Adams et al., 2013; Buonaurio et al., 2015; Lamichhane and Venturi, 2015), yet little progress has been made in applying the latest advances in sequencing and culture-based methodologies to characterize pathosystems in trees. The prevailing paradigm of infection biology contends that one organism causes one disease (proved using Koch’s postulates). In contrast, there is increasing recognition of the importance of polymicrobial interactions in human disease, following developments in sequencing technologies that allow microbiome-wide association studies to identify the role of microbial communities in disease (Gilbert et al., 2016). In the medical field, this is leading to adaptations of Koch’s postulates to include complex interactions between the environment, host and microbial communities (Hill, 1965; Falkow, 1988; Fredricks and Relman, 1996). However, progress in characterizing polyspecies interactions in plant disease has been limited, although there is clearly a need for contemporary approaches to investigating complex tree diseases (Lamichhane and Venturi, 2015). Analysis of such complex biotic interactions requires an integrated, systems approach, particularly in the case of decline-diseases where both complex abiotic and biotic interactions underpin disease development.
Decline-diseases, first formally described as a specific disease in the United States (Sinclair, 1965; Manion, 1981) but well documented elsewhere (Delatour, 1983; Sinclair and Lyon, 2005; Thomas, 2008) are of global concern (Pautasso et al., 2015). Unlike most common tree diseases, decline-diseases are not caused by single primary pests or pathogens; instead they are complex syndromes, involving the sequential, combined and cumulative effects of (often secondary) biotic and abiotic agents (Manion, 1981; Thomas, 2008; Sallé et al., 2014; Brown et al., 2016). Currently, the United Kingdom is facing an episode of Acute Oak Decline (AOD), which occurs widely in southern and midland England, extending into Wales (Brown et al., 2016; Denman et al., 2016), and represents a significant threat to oak, particularly native species Quercus robur and Quercus petraea (Denman et al., 2014). First recognized in Britain in the 1980s (Denman and Webber, 2009), similar declines have occurred in continental Europe (Hartmann et al., 1989; Gibbs and Greig, 1997; Biosca et al., 2003; Vansteenkiste et al., 2004). AOD-affected trees show distinctive, weeping stem patches (stem bleeds), signifying areas of necrosis and fluid-filled cavities in the underlying inner bark (Denman et al., 2014) (see Figures 1a–c), which disrupt vascular flow of nutrients and water essential to tree survival (Vansteenkiste et al., 2004). Larval galleries of the buprestid beetle Agrilus biguttatus are found in conjunction with lesions (Brown et al., 2015, 2017) (Figure 1d) and can also impact tree condition by girdling the tree when colonization is intense, leading to tree death (Sallé et al., 2014). AOD typically affects mature oaks but has also been reported in young trees (Brown et al., 2016).
Symptoms of AOD observed in the field (a–d), reproducing the symptoms of AOD through log inoculations with combinations of bacteria and Agrilus larvae associated with AOD (e–l), and statistical analysis of lesion formation in log inoculations (m–n). (a) Stem bleeds symptomatic of AOD on a mature Quercus robur in the field. (b) Close up view of stem bleeds and bark cracks characteristic of AOD. (c) Cross-section through AOD stem bleeds, showing degradation of vascular tissue (red arrow). (d) Longitudinal section through an AOD stem bleed showing the association between bacterial lesions (blue arrow) and Agrilus biguttatus galleries (red arrow). (e) Wound response in the innerbark of the control treatment (inoculation of a wound with sterile water) of log inoculation trials. (f) Lesion formed in the innerbark of oak logs inoculated with Brenneria goodwinii in log inoculation trials. (g) Lesion formed in the innerbark of oak logs inoculated with Gibbsiella quercinecans in log inoculation trials. (h) Lesion formed in the innerbark of oak logs inoculated with a combination of B. goodwinii and G. quercinecans in log inoculation trials. (i) Wound response in the innerbark of the A. biguttatus treatment (inoculation of a wound with live eggs of A. biguttatus). Note the wound response (blue arrow) and clean gallery created by a larva (red arrow). (j) Lesion formed in the innerbark of oak logs inoculated with B. goodwinii and eggs of A. biguttatus. Note the lesion developing from the inoculation point (blue arrow) and from the galleries (red arrow). (k) Lesion formed in the innerbark of oak logs inoculated with G. quercinecans plus eggs of A. biguttatus. Note the lesion developing from the inoculation point (blue arrow) and from the galleries (red arrow). (l) Lesion formed in the innerbark of oak logs inoculated with B. goodwinii, G. quercinecans plus eggs of A. biguttatus. Note the lesion developing from the inoculation point (blue arrow) and from the galleries (red arrow). (m) Mean lesion area formed in the inner bark of logs inoculated with bacteria. Colour indicates statistical significance (see key). The bacterial species inoculated were back-isolated in fulfilment of Koch’s fourth postulate. Error bars=−s.e. (n) Mean lesion area formed in the inner bark of logs inoculated with bacteria and A. biguttatus eggs. Colour indicates statistical significance (see key in panel (m)). The bacterial species inoculated were back-isolated in fulfilment of Koch’s fourth postulate. Eb—Erwinia billingiae, Bg—Brenneria goodwinii, Gq—Gibbsiella quercinecans. Error bars=−s.e.
First steps to determine the causes of stem bleeds led to the isolation of several novel bacterial species from AOD lesions, with three novel species, Gibbsiella quercinecans (Enterobacteriaceae) (Brady et al., 2010), Brenneria goodwinii (Pectobacteriaceae) (Denman et al., 2012; Adeolu et al., 2016), Rahnella victoriana (Yersiniaceae) (Brady et al., 2014; Adeolu et al., 2016), and an unnamed Pseudomonas (Denman et al., 2016; Sapp et al., 2016) consistently isolated. Some of these species, for example, G. quercinecans as Serratia sp. (Biosca et al., 2003; Poza-Carrión et al., 2008) on Quercus pyrenacia and Quercus ilex are implicated as causative agents of stem bleeding on other oak species in Europe (Biosca et al., 2003; Poza-Carrión et al., 2008; Denman et al., 2016). There was less consistent isolation of various other bacterial species, for example, Lonsdalea quercina ssp. britannica (Brady et al., 2012), which is closely related to Lonsdalea quercina ssp. quercina, the causative agent of acorn gummosis on Q. agrifolia and Q. wislizenii in America (Hildebrand and Schroth, 1967), and several other novel species that are in the process of being formally described. Although correlation of certain bacterial species with AOD symptomology has been observed, empirical evidence on the causative agent(s) of AOD lesions is lacking and remains a barrier to developing informed management strategies for this disease. In the absence of a single putative primary pathogen as the causal agent of AOD lesions, we hypothesized that a polymicrobial complex is responsible for AOD lesion formation. However, demonstrating causation by a polymicrobial complex with associated insect activity on mature oak trees or logs is challenging in the context of fulfilling Koch’s postulates and requires a multi-faceted methodological and conceptual approach.
Here we applied a systems-level approach to determine cause(s) of necrosis in AOD using microbial isolation and culture, phenotypic tests, genomic analyses of G. quercinecans, B. goodwinii and R. victoriana, metagenome and metatranscriptome analysis of healthy and diseased trees and inoculation tests comprising both polybacterial inoculations and the addition of live A. biguttatus eggs to recreate the symptoms of AOD. This combined sequencing and cultivation-based approach provides a contemporary adaptation of Koch’s postulates to address the biotic components of a complex decline-disease and represents a conceptual model for future analyses of polymicrobial infections in trees and other systems.
Methods
Isolation of bacteria from healthy and diseased oak trees
In the search for putative causal agents of stem lesions, we wanted to determine the veracity of differences in occurrence and composition of bacterial communities in lesions and visually healthy trees, as the bacterial microbiome has previously been identified as the likely causal agent of stem lesions (Denman et al., 2016). Conventional isolation and culture and microbiome analyses were used. Samples were acquired through citizen science reports (CSR), as well as structured studies (Denman et al., 2016; Sapp et al., 2016) ensuring the broadest possible coverage of AOD sites (Supplementary Table S1 and S2). Eighteen CSR sites were sampled (Supplementary Figure S1 and Supplementary Table S1), together with those from structured studies (Denman et al., 2016; Sapp et al., 2016), as well as five sites that had no history of AOD (N-AOD). Trees were sampled by forest pathologists who made site visits following the enquiry. In total, isolations from 66 trees were analysed. Destructive sampling of trees, by removing panels of diseased oak tissue, as well as healthy oak tissue from apparently healthy trees on the same sites, was carried out where permitted, and isolations were made using PYGA medium as described in Denman et al. (2014, 2016) and Sapp et al. (2016). Owing to the nature of the CSR studies, the number of tissue pieces plated out was variable, dependent upon the sample. Bacterial colonies emerging from chips of tissue were purified using standard streak-plating techniques; single visually representative colonies were selected, cultivated in Luria Bertani broth (LB) and identified with PCR and DNA amplicon sequencing as described in Denman et al. (2016).
Statistical analysis of isolation study data sets
Bacterial yield between healthy and diseased trees was tested by fitting a generalized linear mixed-effects model with logit link function and binomial error distribution. Fixed effects were fitted for tree health and tissue position and random effects fitted for sites and trees within sites. Overdispersion in the model was taken into account by including an additional dispersion parameter.
Differences in bacterial communities were analysed by detrended correspondence analysis, down weighting bacterial species occurring in <5% of tissue combinations. Monte Carlo permutation tests were used to test for significant differences in bacterial communities between healthy and diseased trees and to test the effect of tissue position within the tree. Finally, Jaccard’s similarity index was used to identify any significant associations between bacteria across the 66 trees of the study.
Genome sequencing of bacterial strains isolated from AOD-affected trees
Maintenance of bacterial strains used in genome analyses
G. quercinecans FRB97 (T) (Brady et al., 2010), B. goodwinii FRB141 (T) (Denman et al., 2012) and R. victoriana BRK18a (T) (Brady et al., 2014) were isolated by Forest Research in the CSR studies from oak trees affected by AOD. Strains were stored in glycerol stocks at −80 °C and maintained on nutrient agar (Oxoid, ThermoFisher Scientific, Loughborough, UK) at 20 °C.
DNA preparation and genome sequencing on Pacific Biosciences RSII sequencing platform
The whole genomes of G. quercinecans FRB97, B. goodwinii FRB141 and R. victoriana BRK18a were sequenced using the Single Molecule Real-Time (SMRT) technology of the Pacific Biosciences RSII platform. A single colony of each isolate was sampled from nutrient agar (Oxoid) and used as inoculum for liquid culture, which was grown overnight in nutrient broth (Oxoid) at 28 °C, and shaken at 150 r.p.m. Total genomic DNA was isolated using the Gentra Puregene Yeast/Bact. Kit (Qiagen, Manchester, UK) and quantified using a Qubit fluorometer (Life Technologies, Paisley, UK). DNA integrity was assessed using 1% agarose gel electrophoresis. G. quercinecans and B. goodwinii DNA libraries were prepared using 10 μg of genomic DNA and sequenced by DUGSIM at Duke University, NC, USA, using P4/C2 chemistry and six SMRT cells per isolate. The R. victoriana DNA library was prepared and sequenced by the Centre for Genomic Research, University of Liverpool, UK using P6/C4 chemistry and one SMRT cell. All sequence data described in this study are available under BioProject PRJNA323828, (Supplementary Table S3). Genome assembly and annotation are described in Supplementary Methods.
Metagenome sequencing of diseased and healthy oak trees
Collection of samples for metagenome sequencing
Tissue samples were collected from Runs Wood, Ross-on-Wye and two sites in Attingham park (Supplementary Table S2). More than half of the trees sampled for metagenome analysis were also sampled in the isolation study. A panel comprising all layers of stem tissue (outerbark, innerbark, sapwood and heartwood) was removed from the visible bleed area on each diseased tree or from stem areas at similar height on healthy trees according to previously described methods (Denman et al., 2014, 2016; Sapp et al., 2016). Samples were immediately flash frozen on dry ice and stored at −80 °C prior to processing.
Metagenome assembly, annotation and mapping to the genomes of G. quercinecans FRB97 (T), B. goodwinii FRB141 (T) and R. victoriana BRK18a (T)
Metagenomic reads were assembled using RAY-meta v2.3.1 (Boisvert et al., 2012) using default parameters. Assemblies were annotated using Prokka v1.11 (Seemann, 2014). Translated annotations were aligned against the translated protein sequences of G. quercinecans FRB97 (T), B. goodwinii FRB141 (T), R. victoriana BRK18a (T) and two control genomes Pectobacterium carotovorum ssp. carotovorum PC1 and Paenibacillus polymyxa SC2 using BLASTx v2.2.26 (Altschul et al., 1990). Metagenome sequences with >97% homology for at least 50 amino acids to proteins identified in G. quercinecans, B. goodwinii and R. victoriana were considered a match (Supplementary Table S4). Control genomes were selected to measure the stringency of alignment. P. carotovorum ssp. carotovorum is a canonical plant pathogen and a member of the soft-rot Enterobacteriaceae (SRE) (N.B. many members have recently been reclassified into novel families (Charkowski et al., 2012; Adeolu et al., 2016)), which had been identified sporadically and at low relative abundance from metagenomic taxonomic surveys within this study (Supplementary Table S5). However, P. carotovorum ssp. carotovorum had not previously been isolated from necrotic lesions on AOD-affected trees. Therefore, it was proposed that coding domain alignments to P. carotovorum ssp. carotovorum align to conserved genes, which are present in many Enterobacteriaceae, and their low relative abundance does not signify their presence but is an artefact of the alignment process. Resultant Circos plots agree with this proposal as conserved genes are frequently found in the metagenomic alignment against the P. carotovorum ssp. carotovorum PC1 genome, which are likely to align to other Enterobacteriaceae. Within G. quercinecans, B. goodwinii and R. victoriana, there is strong homology to conserved and variant genes. P. polymyxa SC2 was identified at low relative abundance in healthy and diseased metagenomic samples (Supplementary Table S5), therefore it was selected to test stringency of the metagenomic alignment and to measure its relative abundance in healthy and diseased microbiomes using an alternative method. Homologous bacterial protein identities and the workflow used for metagenomic analysis is available from GitHub: (https://github.com/clydeandforth/multi_omics_study.git).
Taxonomic classification of metagenome sequences
To compare the taxonomic composition of the oak microbiome, raw sequence reads were taxonomically labelled using Kraken v0.10.5 beta (Wood and Salzberg, 2014) and One Codex (Minot et al., 2015) (Figure 2a). Taxonomic labelling using Kraken was performed on the standard RefSeq genome database supported by Kraken, with the addition of the genomes of B. goodwinii FRB141 (T), R. victoriana BRK18a (T), G. quercinecans FRB97 (T) and Lonsdalea quercina ssp. quercina ATCC 29281 (Figure 2d and Supplementary Table S5). Taxonomic analysis using One Codex was conducted against the One Codex March ’16 Preview database with the addition of the genomes of G. quercinecans, B. goodwinii and R. victoriana.
Comparative metagenomic analysis of the taxonomic composition and function of the oak microbiome in healthy tissue (from trees without AOD) and diseased tissue (from trees with AOD). (a) The plot depicts the genera represented in all metagenomes based on One Codex binning of raw reads, demonstrating a clear shift in the microbial community between healthy and diseased trees. (b) Brenneria and Gibbsiella are statistically correlated to diseased tissue, as shown by a principal coordinate ordination analysis based on statistics calculated by Primer v7 and PERMANOVA+ using One Codex binning data. The first two axes depict the plotted community composition. Correlation vectors in the graph are significant using an R2>0.05. A Welch’s t-test was performed to test significance of differences between key taxa (identified above) and between healthy and diseased trees (pooled abundances for each factor). Resultant P-values from Welch’s t-test are overlaid on the correlation biplot. (c) Gene groups involved in bacterial phytopathogenic activity are significantly increased in AOD diseased trees, as shown by comparative functional analysis of SEED subsystem categories as annotated by MG-RAST on assembled metagenome contigs. The analysis of SEED subsystem metagenome data was performed using Stamp. Statistically significant functional differences between diseased and healthy communities were calculated using G-test with Yates’ correction. The Newcombe–Wilson test was performed to calculate confidence intervals between two binomial population proportions. (d) The genomes of 17 species were found to be common across all AOD metagenomes. Visualization of Kraken metagenome analysis of stem samples demonstrates the shifts in bacterial microbiome compositions. Bubble sizes are categorized based on the relative percentage frequencies of Kraken species-level alignment of raw metagenome sequencing reads and are depicted in the figure as 1 (read frequency <0.01), 2 (read frequency 0.01–0.1), 3 (read frequency 0.1–1), 4 (read frequency 1–10) and 5 (read frequency >10). Red bubbles signify samples from diseased trees, while blue bubbles signify samples from healthy trees. The 17 common species were determined based on common occurrence across all diseased tissue samples. Additionally, six species more abundant among the healthy trees were included to provide a contrasting shift.
Functional annotation of metagenome sequences
Metagenome datasets derived from samples AT2, AT3, AT4, AT5, AT6, ROW1, ROW2 and ROW3 were analysed via MG-RAST (Meyer et al., 2008) using Hierarchical Classification against the Subsystems database with an E-value cutoff of 1e-5, a minimum percentage identity cutoff of 80% and a minimum alignment length of 50. Descriptions of the taxonomic and functional composition of the metagenomes derived from MG-RAST were comparable with those derived from analysis of the same data sets using One Codex and Kraken and further validated by mapping of metagenome reads against the finished genomes of B. goodwinii, G. quercinecans and R. victoriana.
Statistical analysis of metagenome data sets
Statistical analyses were performed using Primer v7 (Clarke and Gorley, 2015) with PERMANOVA+ add on to explore relationships between community changes (Figure 2b). One Codex metagenome data was log (N+1) transformed to downweight the most abundant genera. Next, dissimilarities were calculated with the S17 Bray–Curtis similarity coefficient. A principal coordinate ordination analysis was performed by plotting the interpoint dissimilarity values for each factor (site and disease status), and the variation in community composition was plotted as the first two axes (preserving actual dissimilarities) (Gower, 1966). A correlation was performed between each taxon and each community coordinate. Correlations with each component were deemed significant (R2>0.5) and a vector biplot was overlaid to visualize the strength of the correlation. A Welch’s t-test was performed to test significance of differences between key taxa (identified above) and between healthy and diseased trees (pooled abundances for each factor). Resultant P-values from Welch’s t-test are overlaid on correlation biplot with significance at >95% (P<0.05) deemed significant. Comparative functional analysis of MG-RAST (Meyer et al., 2008) annotated, metagenome data was performed using Stamp v2.1.3 (Parks et al., 2014). Statistically significant functional differences between diseased and healthy communities were calculated using G-test with Yates’ correction. The Newcombe–Wilson test was performed to calculate confidence intervals between two binomial population proportions (Brown and Li, 2005).
Metatranscriptome sequencing of AOD diseased oak trees
Collection of samples for metatranscriptome sequencing and RNA extraction
For RNA sampling, two separate lesions (samples AT11 and AT12) from a single tree were analysed in June 2013. Swabs of the lesion fluid were collected in addition to tissue from the active margins of the lesion and immediately frozen in liquid nitrogen and transported back to the laboratory in a vessel containing liquid nitrogen. Samples were stored at −80 °C prior to processing. Before RNA extraction, samples in liquid nitrogen were ground with a pestle in a mortar to homogenize the tissue. RNA was extracted from 2 g of tissue using the PowerSoil Total RNA Isolation Kit (MoBio, Cambridge, UK) according to the manufacturer’s instructions. RNA was quantified using a Qubit fluorometer (Thermo Fisher, Warrington, UK) and quality assessed using the Bioanalyser 2100 (Agilent, Manchester, UK).
Metatranscriptome sequencing
Sequencing libraries were prepared from samples of total RNA using the strand-specific ScriptSeq Preparation Kit (Illumina, Cambridge, UK) and sequenced using 2 × 100 bp paired-end sequencing on the Illumina HiSeq platform. Reads were trimmed using first Cutadapt 1.2.1 (Martin, 2011) and additionally Sickle 1.2.00 (Joshi and Fass, 2011). Owing to low RNA yields from the lesion samples, total RNA was sequenced and rRNA sequence reads were subsequently depleted in silico prior to mRNA transcript analysis.
Metatranscriptome assembly and functional annotation
Two in silico rRNA-depleted metatranscriptome libraries were aligned to the G. quercinecans FRB97 (T), B. goodwinii FRB141 (T), R. victoriana BRK18a (T) and control genomes (P. carotovorum ssp. carotovorum PC1 and P. polymyxa SC2) (Supplementary Figures S2 and S3) with Bowtie2 v2.2.4 (Langmead and Salzberg, 2012), using local mode to maximize alignment score. Aligned reads were converted from Sequence Alignment/Map (SAM) to Binary Sequence Alignment/Map (BAM) format and indexed using SAMtools v1.2 (Li et al., 2009). To avoid false positives in the detection of gene expression, a gene was considered as being expressed if ⩾3 transcripts were aligned and the combined coverage from both libraries represented >20% of the gene, (adapted from (Versluis et al., 2015) (Figures 2a–c and Supplementary Figure S4). A custom Perl script was designed to extract transcript alignments and is available from GitHub: (https://github.com/clydeandforth/multi_omics_study.git). Aligned transcripts were visualized in Artemis (Carver et al., 2012).
Log inoculations
Pathogenicity tests were set up to reproduce lesions characteristic of AOD under controlled conditions. The following hypotheses were tested: (1) Key species consistently isolated from AOD symptomatic oak can cause necrosis of oak stem tissue, (2) Combinations of key bacterial species cause more severe tissue necrosis (reflected in larger lesions), than individual species alone, and (3) The interaction between A. biguttatus larvae (derived from eggs) and bacteria leads to the development of AOD symptoms.
Four experiments were carried out over 3 consecutive years as testing could only be carried out annually when beetle eggs were available. Three trials were carried out on oak logs in growth chambers, and the fourth trial was set out in the field, where stems of young plantation-oak (25 years old), were used instead of logs.
Growth chamber log trials
Logs used in the trials were obtained from freshly felled Q. robur trees, with diameter at 1.3 m (DBH)=16–20 cm. The trees were located in the Straits Enclosure of the Alice Holt Forest, Hampshire, England, UK, and logs were transported to the laboratory after felling where they were prepared for inoculation. Logs measuring 40 cm (mini-logs) or 130 cm (long logs) in length had the uppermost cut surface sealed with isoflex liquid rubber (Ronseal Ltd, Chapeltown, Sheffield) to prevent desiccation. Logs were placed, lower cut surface down, in saucers containing water. Logs were inoculated in August or September and incubated at 25 °C in the growth chamber with a 12 h photoperiod. Mini-logs were used for all treatments involving A. biguttatus eggs; in each case, only one bacterial+eggs treatment type per mini-log was tested to guard against cross-contamination through larval spread (Supplementary Table S2). All the bacterial treatments without eggs (either single species or combinations of species) were placed on the same long log, with inoculation points marked out along its length. Eggs of A. biguttatus were produced in Forest Research’s laboratories at Alice Holt (Reed, 2016). In a single experiment (2014 NW; Supplementary Table S2), non-wound inoculations were carried out; the remaining two growth chamber trials and the field trial were inoculated using shallow wounds to the outer surface of the innerbark, made by a 10 mm cork-borer. Half a loop of inoculum scooped from 24-h-old inoculum plates using disposable plastic loops was inserted into the wound and rubbed to dislodge the bacterial cells around the wound surface. The outer bark plug was replaced on the inoculation point, wounds covered with parafilm and damp cotton wool and sealed with duct tape. After 4 months of incubation, experiments were terminated, outer bark stripped from inoculation points to expose lesions, which were hand traced onto tracing paper, and back isolations were cultured onto peptone yeast glucose agar. Mass bacterial colonies that developed were tested for the presence of B. goodwinii, G. quercinecans, Rahnella and Lonsdalea using a multiplex Taqman qPCR assay (Roche, Brighton, UK). Lesion areas were calculated using ASSESS V2 (APS, Minneapolis, MN, USA).
Statistical analyses
Log lesion areas
The four pathogenicity trials were used to establish the impact of different bacterial species tested individually, in combination (polybacterial inoculations) and with or without the addition of A. biguttatus eggs (Supplementary Table S2) on tissue necrosis (assessed by the size of the lesion area associated with each inoculation point).
Lesion area data were refined in a hierarchical manner, such that the following data sets were used for lesion area analysis:
-
1
Non-contaminated samples fulfilling Koch’s postulates, with galleries present (for A. biguttatus samples indicating that eggs had hatched);
-
2
Non-contaminated samples fulfilling Koch’s postulates, with or without galleries present (for A. biguttatus samples); and
-
3
Non-contaminated samples.
Further information on statistical analyses of log lesion areas are described in Supplementary Methods.
Results
Isolation study
In the isolation study, analysis of 38 diseased trees from 23 sites, plus 13 healthy trees in 11 of these sites, and 15 healthy trees from five sites with no history of AOD identified 159 bacterial taxa. Higher yields of bacteria were obtained from lesion margin tissue of symptomatic trees compared with healthy tree tissue (F1,28, P<0.001), and the lesion margin bacterial profile was significantly different to healthy tissues (Monte-Carlo permutation test of 1000 permutations, P<0.001) (Supplementary Figure S5 and Supplementary Table S6). Key genera isolated included Pseudomonas (comprising multiple taxa and occurring in healthy as well as diseased trees); but Gibbsiella, Brenneria and Rahnella dominated lesion margins (Supplementary Table S6). G. quercinecans occurred on all disease sites and was isolated from 83% of diseased and 4% of healthy trees sampled, comprising 17% of total diseased samples and <0.1% of total healthy samples. B. goodwinii was more difficult to isolate but was obtained from 15 sites, accounting for 16% of total diseased and <0.1% of total healthy samples. Members of the genus Rahnella were obtained from 65% of diseased sites, but R. victoriana was isolated only from diseased trees (37%), on nine of the sites. L. quercina ssp. britannica (Brady et al., 2012) was isolated sporadically on four sites, and a Pseudomonas species (P. fulva-like) not yet formally identified, was isolated at eight sites from diseased trees only (Supplementary Table S6). There was a significant co-occurrence of G. quercinecans and B. goodwinii in diseased tissue (J=0.56, P<0.001), but neither was isolated from trees on sites with no history of AOD.
Oak microbiome analysis
Taxonomic analysis of the metagenome using unassembled metagenome sequence reads against the One Codex (Minot et al., 2015) March ’16 Preview database (with the addition of the genomes of G. quercinecans, B. goodwinii and R. victoriana) also revealed a shift in microbiome composition between healthy stem tissue and AOD lesions (Figures 2a and b). Periglandula, Burkholderia, Streptomyces, Bacillus and Auriemonas were the most abundant genera in healthy trees, whereas Brenneria dominated diseased tissue (mean read abundance, 37.5%) (Figure 2a). The mean read abundance of Gibbsiella (0.9%) and Rahnella (3.7%) was also greater in diseased tissue when compared with healthy tissue (both 0.1%). Pseudomonas, a diverse genus comprising both endophytic and phytopathogenic bacteria (Vinatzer et al., 2014), had similar mean abundances in both diseased (4.8%) and healthy trees (3.3%). Correlation coefficients and Welch’s unequal variances t-tests revealed that Streptomyces (tWelch’s=49.7, P=0.004) and Periglandula (tWelch’s=821.8, P<0.001) were significantly associated with healthy trees, whereas Brenneria (tWelch’s=12.4, P=0.006) and Gibbsiella (tWelch’s=4.7, P=0.05) were strongly correlated with the lesions of diseased trees (Figure 2b and Supplementary Table S7). Pseudomonas and Rahnella were not strongly correlated with either health state (Supplementary Table S7).
To identify the most abundant species in the lesion microbiome, raw metagenome reads were aligned using Kraken (Wood and Salzberg, 2014) against reference genome databases (see Methods section), revealing 17 genomes that were commonly detected in diseased tissue (Figure 2d and Supplementary Table S7). B. goodwinii, G. quercinecans and R. victoriana were detected in the lesion metagenome of all trees with AOD. Overall, B. goodwinii was the most abundant genome in the lesion microbiome (range, 0.3–49% of metagenome sequence reads; mean, 12%) but was also detected in much lower proportions in healthy trees (0.01% in all samples). R. victoriana (range, 0.01–15%; mean, 2.1%) and G. quercinecans (range, 0.02–0.8%; mean, 0.3%) were the second and fourth most abundant genomes in the lesions of diseased trees, respectively. Functional metagenome analysis (Figure 2c and Supplementary Figure S6) revealed that genes associated with carbohydrate metabolism, membrane transport and virulence, defence and disease are key features of the lesion microbiome, suggesting that many of these functions are encoded in the genome of B. goodwinii, G. quercinecans and R. victoriana.
Genome analysis of AOD-associated bacteria
Whole-genome sequencing of B. goodwinii FRB141, G. quercinecans FRB97 and R. victoriana BRK18a (Supplementary Table S8) revealed that they are phylogenetically related to opportunistic phytopathogens belonging to the SRE (Charkowski et al., 2012; Adeolu et al., 2016) and possess a catalogue of virulence genes associated with canonical phytopathogens (Supplementary Table S9). The SRE alternate their lifestyle from benign commensals to brute force necrotrophic pathogens, which macerate cell wall polysaccharides by releasing plant cell wall-degrading enzymes (PCWDEs) (Toth et al., 2006). A genome-wide search for PCWDEs in G. quercinecans, B. goodwinii and R. victoriana revealed the presence of pectinases, cellulases and tannases (Figure 3b, Supplementary Table S9) whose activity has been confirmed phenotypically (Supplementary Figure S7). Furthermore, G. quercinecans and R. victoriana possess a type II secretion system, an operon which releases most PCWDEs and is therefore the central virulence facilitator of necrotrophic plant pathogens. B. goodwinii encodes a type III secretion system, the principal virulence factor of established hemibiotrophic pathogens such as Pseudomonas syringae, which use the operon to evade immune surveillance, allowing bacteria to increase their numbers before releasing necrotic enzymes as nutrients are depleted (Tampakaki et al., 2010).
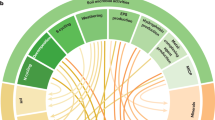
Functional genome analysis of Gibbsiella quercinecans FRB97 (T), Brenneria goodwinii FRB141 (T) and Rahnella victoriana BRK18a. (a) Circular representations of B. goodwinii FRB141, G. quercinecans FRB97 and R. victoriana BRK18a genomes. From outside to inside, circles represent: (1) Assembled bacterial genomes, outermost (orange) circle, with encoded secretion system annotated at their genomic loci. (2) Metatranscriptome heatmap. Alignment of two in silico combined metatranscriptomes recovered from two necrotic lesions of AOD-affected trees against the bacterial genomes. Blue saturation represents increasing transcript alignments. (3–9) Seven metagenomes from necrotic lesions on AOD-affected trees and one healthy tree (metagenomes were extracted from two sites, Attingham and Runs Wood) were aligned through their coding domains to homologous regions in the bacterial genomes. (3) Attingham healthy (AT1) aligned metagenome-coding domains (light purple). (4) Attingham diseased (AT7) aligned metagenome-coding domains (aqua). (5) Attingham diseased (AT8) aligned metagenome-coding domains (blue). (6) Attingham diseased (AT9) aligned metagenome-coding domains (orange). (7) Runs Wood diseased (RW1) aligned metagenome-coding domains (green). (8) Runs Wood diseased (RW2) aligned metagenome-coding domains (pink). (9) Runs Wood diseased (RW3) aligned metagenome-coding domains (grey). (10) G+C content across the bacterial genome. (11) G+C skew across the bacterial genome. (b) Schematic diagram of transporters and transported proteins recovered from lesion metatranscriptomes and aligned against each bacterial genome. The number of significantly expressed (>3 aligned transcripts, covering >20% of the gene) genes is shown in parentheses. (c) Number of transcripts aligned against selected virulence genes encoded within B. goodwinii FRB141, G. quercinecans FRB97 and R. victoriana BRK18a. Gene categories are represented by the following colours: red—PCWDE, purple—general secretory pathway (GSP), yellow—type II secretion system (T2SS), blue—type III secretion system (T3SS), pink—type III secretion system effectors (T3SS effectors), and green—global regulators (GR).
To address the role of B. goodwinii, G. quercinecans and R. victoriana in the aetiology of AOD, we aligned metagenome sequences and transcripts recovered from necrotic lesions of AOD-affected trees against their genomes (Figure 3a). Alignment of six AOD lesion metagenomes revealed an average of 2225 homologous proteins in B. goodwinii FRB141, 858 in G. quercinecans FRB97 and 396 in R. victoriana BRK18a (Supplementary Table S4). Furthermore, annotated genes from the assembled metagenome of a healthy oak revealed only two homologous proteins in B. goodwinii FRB141, one in G. quercinecans FRB97 and two in R. victoriana BRK18a. Lesion transcripts were aligned to coding regions within B. goodwinii FRB141, G. quercinecans FRB97 and R. victoriana BRK18a; this revealed that the transcripts aligned significantly to virulence genes, including PCWDEs, secretion system machinery and regulators of PCWDEs (Figure 3c and Supplementary Figure S4). G. quercinecans FRB97 expressed the pectic enzymes, polygalacturonase and rhamnogalacturonate lyase, and β-glucosidase (cellulase), pectate exo-lyase and oligogalacturonide lyase (which cleaves polygalacturonic acid, the by-product of pectin lyase) (Moran et al., 1968). B. goodwinii FRB141 expressed phosphocellosbiose, β-galactosidase and several type III secretion system effectors, R. victoriana BRK18a expressed many general secretory pathway genes, a β-glucosidase, a tannase and carbohydrate esterase enzyme (Yao et al., 2013). Key regulators of pectinolysis were expressed in all three bacteria, including kdgR, phoP, pecT, rsmA/rsmB and gacA (Figure 3a).
Log lesion areas
Figures 1e–l show lesions caused by bacteria with or without A. biguttatus inoculated into logs in pathogenicity tests. Notable results for lesion area analysis are presented in Figures 1m and n (non-contaminated samples fulfilling Koch’s postulates, with galleries present for A. biguttatus samples). In bacteria-only inoculations, lesions significantly bigger than wound controls (Figure 1m) were made by the combination of bacterial species B. goodwinii+G. quercinecans (tdunnetx=2.97, P=0.044); the bacterial species G. quercinecans had larger lesion areas versus the controls, although this was not significant and the P<0.05 level (tdunnetx=2.79, P=0.068); B. goodwinii lesions were smaller and not significantly different from the controls in terms of mean lesion area (tdunnetx=1.30, P=0.796), but the necrosis was clearly different to the control wound response (see Figure 1e vs f). Inoculation with Erwinia billingiae, a known ubiquitous non-pathogenic bacterium, served as a negative control demonstrating lesion area no different to the control wound response. The pattern strengthened when bacteria plus A. biguttatus (applied as eggs) were co-inoculated (Figure 1n). Lesions created by B. goodwinii and A. biguttatus eggs (larvae) were significantly bigger than wound controls (tdunnetx=3.40, P=0.0125), as were G. quercinecans and A. biguttatus (tdunnetx=4.65, P=0.0002) and the combination of B. goodwinii+G. quercinecans+A. biguttatus (tdunnetx=3.92, P=0.0027) (Figure 1n). Bacteria were re-isolated from lesion margins and at intervals along the larval galleries, which were becoming necrotic-forming part of the lesion. Apart from demonstrating the necrogenic ability of the bacteria, these results showed that the larvae and larval galleries have an important part in increasing lesion area, implicating them in the spread of bacteria within infected trees.
Bacterial-positive back-isolation and A. biguttatus contamination
There was a high success of back-isolation for both B. goodwinii and G. quercinecans when these species were included as treatments (70% and 80%, respectively); both species were also identified as contaminants (that is, back-isolated when not part of the treatment inoculation; B. goodwinii=9%; G. quercinecans=16%), although the contamination rate was significantly lower than treatment back-isolation in both cases (B. goodwinii: z=7.98, P<0.001; G. quercinecans: z=6.65, P<0.001). For G. quercinecans, the rate of contamination was affected by the presence of A. biguttatus eggs: when eggs were absent G. quercinecans contamination was 3%; when eggs were present this rose to 38% contamination, a significant increase (z=3.59, P<0.001). There was no significant effect of A. biguttatus eggs on B. goodwinii contamination (eggs absent, B. goodwinii contamination=2%; eggs present, B. goodwinii contamination=7%, z=1.60, P=0.11).
Discussion
Fulfilling Koch’s postulates represents the traditional paradigm for proving disease causation. However, contemporary molecular approaches have transformed our appreciation of the role of microbial communities and polymicrobial interactions in disease. There are situations where the one pathogen causes one disease model is not suitable to prove causality and must be adapted to accommodate polymicrobial infections. For example, in decline-diseases of trees, application of Koch’s postulates in the strictest sense cannot fully address proving causality of lesion formation as more than one necrogenic agent is involved in the disease syndrome. Here causation of AOD lesions was demonstrated using a combined cultivation-based and sequencing approach in a dynamic adaptation of Koch’s postulates analogous to that described by Byrd and Segre (2016). Koch’s first postulate is fulfilled by the consistent isolation and metagenomic detection of G. quercinecans, B. goodwinii and R. victoriana in trees affected by AOD.
This combined approach delivered an improved understanding of lesion microbiome composition, as some species were difficult to isolate and culture. For example, B. goodwinii and Lonsdalea quercina ssp. britannica were under-represented, whereas G. quercinecans, which is more amenable to isolation and cultivation, was overrepresented, with the opposite trend evident in metagenomic studies. Furthermore, G. quercinecans, B. goodwinii and R. victoriana were absent on sites with no history of AOD, were rarely isolated from healthy oak on diseased sites and had negligible abundance in healthy metagenome samples at AOD sites, complying with Koch’s second postulate. Ultra-low levels of G. quercinecans, B. goodwinii and R. victoriana were detected in some of the healthy trees but only from sites where AOD is present, raising interesting questions about their existence in the wider environment. Possible explanations for the presence of G. quercinecans, B. goodwinii and R. victoriana in healthy trees on AOD sites may include; (1) the tree is in early stages of lesion formation, where visible symptoms have not yet developed, or that (2) G. quercinecans, B. goodwinii and R. victoriana are endophytes or epiphytes that opportunistically multiply after tissue necrosis is initiated by another organism. In the first situation, the chances of detecting asymptomatic lesion formation seem fairly remote, especially as crown condition is not a reliable indicator of tree predisposition status in the early stages of decline development. In the second case, several pieces of evidence counteract the possibility that these organisms multiply opportunistically after tissue necrosis occurs, as (1) we observed the expression of putative necrogenic enzymes and virulence factors of these bacteria in AOD lesions in planta, implicating them in having an active role in tissue degradation, and (2) G. quercinecans and B. goodwinii caused significant lesion formation in log inoculation trials, demonstrating actual lesion-forming capabilities. It is important to note that the oak logs and trees used in our trials were selected from areas with no history of the disease.
Furthermore, all three bacterial species possess the genomic capability to cause tissue necrosis, as determined through genomics, in situ functional metagenomics and metatranscriptomics. Inoculation onto live oak logs confirmed significant necrogenic ability of B. goodwinii and G. quercinecans, but further work with R. victoriana inoculation is required. Thus the pathogenic phenotype of B. goodwinii and G. quercinecans has been confirmed, broadly fulfilling Koch’s third postulate that inoculation tests can cause the disease anew. However, bacterial species combinations caused even greater necrosis indicating cumulative effect and possible synergism, this is a contemporary adaptation of Koch’s postulates sensu stricto. Finally, once inoculated onto live oak panels, B. goodwinii and G. quercinecans were re-isolated to fulfil Koch’s fourth postulate (although back-isolation of some species was difficult, and there is a need for developing rapid, cost-effective tools to detect pathogens that are difficult to isolate). Consequently, we propose that the biotic component of the AOD lesions is a polybacterial complex comprised primarily of G. quercinecans and B. goodwinii, which are now established as key causal agents of tissue necrosis, the primary symptom of AOD. Our studies also indicated that other members of the microbiome may contribute to the pathology of AOD. Microbiome analysis suggested that R. victoriana is abundant and also important in AOD lesions; however, back-isolation of R. victoriana was variable, and further tests to characterize its role and interactions with B. goodwinii and G. quercinecans are required. In addition, Lonsdalea quercina ssp. britannica in particular demonstrated variable but at times virulent pathogenicity in log inoculations, although it was not consistently present in AOD lesions. Our results indicate that Agrilus larvae potentiate the spread of these necrogenic bacterial species within tree tissue, generating new break-out points of tissue necrosis and replicating the observed aetiology of AOD. Ultimately, vascular degradation arising from a combination of bacterial tissue necrosis and inner bark damage from larval galleries exacerbates and accelerates the decline by interrupting carbon resource allocation, preventing accumulation in roots and reducing water availability.
It is clear that, where possible, microbiome analysis methods together with inoculation assays should become the accepted standard for profiling disease complexes, particularly when considering complex interactions between microorganisms, insects, the environment and the host. This phenomenon has never previously been addressed in arboreal systems using the approaches described here. Our work therefore highlights the importance of a systems-level approach for characterizing the pathology of complex diseases and represents a conceptual and methodological template for adapting Koch’s postulates to address the role of microbial communities in disease. In recent years, microbiome studies have transformed our understanding of the role of human gut microbiota in a variety of conditions, including bowel and cardiovascular disease (Frank et al., 2007; Koeth et al., 2013), and consequently, microbiome-wide association studies that link microbial consortia to disease will have an important role in the development of future diagnostics and therapies for disease (Gilbert et al., 2016). For decline-diseases in particular, further adaptation may be required to include the role of host predisposition.
Although AOD has been formally described as a major threat to UK oak, similar decline-diseases have been reported in mainland Europe (Delatour, 1983; Hartmann et al., 1989; Biosca et al., 2003; Vansteenkiste et al., 2004; Poza-Carrión et al., 2008), the Middle East and the Americas (Lynch et al., 2014), indicating that AOD is a global concern that has likely evaded attention due to the complexity of its causative agents. The polybacterial nature of AOD exhibits similarities to other economically important tree diseases such as olive knot disease (Buonaurio et al., 2015) but may also be applied to other complex diseases. Generally, our findings highlight the importance of understanding polymicrobial interactions in the context of future-proofing plant health to protect important but increasingly disease-prone forests and crops that are a fundamental part of our global landscape.
References
Adams AS, Aylward FO, Adams SM, Erbilgin N, Aukema BH, Currie CR et al . (2013). Mountain pine beetles colonizing historical and naive host trees are associated with a bacterial community highly enriched in genes contributing to terpene metabolism. Appl Environ Microbiol 79: 3468–3475.
Adeolu M, Alnajar S, Naushad S, Gupta R . (2016). Genome based phylogeny and taxonomy of the ‘Enterobacteriales’: proposal for Enterobacterales ord. nov. divided into the families Enterobacteriaceae Erwiniaceae fam. nov., Pectobacteriaceae fam. nov., Yersiniaceae fam. nov., Hafniaceae fam. nov., Morganellaceae fam. nov., and Budviciaceae fam. nov. Int J Syst Evol Microbiol 66: 5575–5599.
Altschul SF, Gish W, Miller W, Myers E, Lipman DJ . (1990). Basic Local Alignment Search Tool. J Mol Biol 215: 403–410.
Biosca EG, González R, López-López MJ, Soria S, Montón C, Pérez-Laorga E et al . (2003). Isolation and characterization of Brenneria quercina, causal agent for bark canker and drippy nut of Quercus spp. in Spain. Phytopathology 93: 485–492.
Boisvert S, Raymond F, Godzaridis E, Laviolette F, Corbeil J . (2012). Ray Meta: scalable de novo metagenome assembly and profiling. Genome Biol 13: R122.
Boyd IL, Freer-Smith PH, Gilligan CA, Godfray HCJ . (2013). The consequence of tree pests and diseases for ecosystem services. Science 342: 823–834.
Brady C, Denman S, Kirk S, Venter S, Rodríguez-Palenzuela P, Coutinho T . (2010). Description of Gibbsiella quercinecans gen. nov., sp. nov., associated with Acute Oak Decline. Syst Appl Microbiol 33: 444–450.
Brady C, Hunter G, Kirk S, Arnold D, Denman S . (2014). Rahnella victoriana sp. nov., Rahnella bruchi sp. nov., Rahnella woolbedingensis sp. nov., classification of Rahnella genomospecies 2 and 3 as Rahnella variigena sp. nov. and Rahnella inusitata sp. nov., respectively and emended description of the genus Rahnella. Syst Appl Microbiol 37: 545–552.
Brady CL, Cleenwerck I, Denman S, Venter SN, Rodríguez-Palenzuela P, Coutinho TA et al . (2012). Proposal to reclassify Brenneria quercina (Hildebrand and Schroth 1967) Hauben et al. 1999 into a new genus, Lonsdalea gen. nov., as Lonsdalea quercina comb. nov., descriptions of Lonsdalea quercina subsp. quercina comb. nov., Lonsdalea quercina subsp. iberica subsp. nov. and Lonsdalea quercina subsp. britannica subsp. nov., emendation of the description of the genus Brenneria, reclassification of Dickeya dieffenbachiae as Dickeya dadantii subsp. dieffenbachiae comb. nov., and emendation of the description of Dickeya dadantii. Int J Syst Evol Microbiol 62: 1592–1602.
Brown L, Li X . (2005). Confidence intervals for two sample binomial distribution. J Stat Plan Inference 130: 359–375.
Brown N, Inward DJG, Jeger M, Denman S . (2015). A review of Agrilus biguttatus in UK forests and its relationship with acute oak decline. Forestry 88: 53–63.
Brown N, Jeger M, Kirk S, Xu X, Denman S . (2016). Spatial and temporal patterns in symptom expression within eight woodlands affected by Acute Oak Decline. For Ecol Manage 360: 97–109.
Brown N, Jeger MJ, Kirk S, Williams DT, Xu X, Pautasso M et al . (2017). Acute Oak Decline and Agrilus biguttatus: the co-occurrence of stem bleeding and D-shaped emergence holes in Great Britain. Forests 8: 87.
Buonaurio R, Moretti C, da Silva DP, Cortese C, Ramos C, Venturi V . (2015). The olive knot disease as a model to study the role of interspecies bacterial communities in plant disease. Front Plant Sci 6: 434.
Byrd AL, Segre JA . (2016). Adapting Koch’s postulates. Science 351: 224–226.
Carver T, Harris SR, Berriman M, Parkhill J, McQuillan JA . (2012). Artemis: An integrated platform for visualization and analysis of high-throughput sequence-based experimental data. Bioinformatics 28: 464–469.
Charkowski A, Blanco C, Condemine G, Expert D, Franza T, Hayes C et al . (2012). The role of secretion systems and small molecules in soft-rot Enterobacteriaceae pathogenicity. Annu Rev Phytopathol 50: 425–449.
Clarke K, Gorley R . (2015) PRIMER v7: User Manual/Tutorial. PRIMER-E: Plymouth, UK.
Cohen W, Yang Z, Stehman SV, Shroeder TA, Bell DM, Masek JG et al . (2016). Forest disturbance across the conterminous United States from 1985-2012: The emerging dominance of forest decline. For Ecol Manage 360: 242–252.
Delatour C . (1983). Die-back of oak in Europe. Biol For 35: 265–282.
Denman S, Brady C, Kirk S, Cleenwerck I, Venter S, Coutinho T et al . (2012). Brenneria goodwinii sp. nov., associated with acute oak decline in the UK. Int J Syst Evol Microbiol 62: 2451–2456.
Denman S, Brown N, Kirk S, Jeger M, Webber J . (2014). A description of the symptoms of Acute Oak Decline in Britain and a comparative review on causes of similar disorders on oak in Europe. Forestry 87: 535–551.
Denman S, Plummer S, Kirk S, Peace A, McDonald JE . (2016). Isolation studies reveal a shift in the cultivable microbiome of oak affected with Acute Oak Decline. Syst Appl Microbiol 39: 18–20.
Denman S, Webber J . (2009). Oak declines: new definitions and new episodes in Britain. Q J For 103: 285–290.
Falkow S . (1988). Molecular Koch’s postulates applied to microbial pathogenicity. Rev Infect Dis 10: 7–10.
Frank DN St, Amand AL, Feldman RA, Boedeker EC, Harpaz N, Pace NR . (2007). Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc Natl Acad Sci USA 104: 13780–13785.
Fredricks DN, Relman DA . (1996). Sequence-based identification of microbial pathogens: a reconsideration of Koch’s postulates. Clin Microbiol Rev 9: 18–33.
Gibbs JN, Greig BJW . (1997). Biotic and abiotic factors affecting the dying back of pedunculate oak Quercus robur L. Forestry 70: 399–406.
Gilbert JA, Quinn RA, Debelius J, Xu ZZ, Morton J, Garg N et al . (2016). Microbiome-wide association studies link dynamic microbial consortia to disease. Nature 535: 94–103.
Gower JC . (1966). Some distance properties of latent root and vector methods used in multivariate analysis. Biometrika 53: 325–338.
Hartmann G, Blank R, Lewark S . (1989). Oak Decline in Northern Germany: distribution, symptoms, possible causes. Forst Holz 44: 475–487.
Hildebrand D, Schroth M . (1967). A new species of Erwinia causing the drippy nut disese of live oaks. Phytopathology 57: 250–253.
Hill AB . (1965). The environment and disease: association or causation? Proc R Soc Med 58: 295–300.
Joshi N, Fass J . (2011), Sickle. A sliding-window, adaptive, quality-based trimming tool for FastQ files https://github.com/najoshi/sickle.
Koeth RA, Wang Z, Levison BS, Buffa JA, Org E, Sheehy BT et al . (2013). Intestinal microbiota metabolism of L-carnitine, a nutrient in red meat, promotes atherosclerosis. Nat Med 19: 576–585.
Lamichhane JR, Venturi V . (2015). Synergisms between microbial pathogens in plant disease complexes: a growing trend. Front Plant Sci 6: 385.
Langmead B, Salzberg SL . (2012). Fast gapped-read alignment with Bowtie 2. Nat Methods 9: 357–359.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N et al . (2009). The sequence alignment/map format and SAMtools. Bioinformatics 25: 2078–2079.
Lynch SC, Zambino PJ, Scott TA, Eskalen A . (2014). Occurrence, incidence and associations among fungal pathogens and Agrilus auroguttatus, and their roles in Quercus agrifolia decline in California. For Pathol 44: 62–74.
Manion P . (1981) Tree Disease Concepts. Prentice Hall: Englewood Cliffs, New Jersey, USA.
Martin M . (2011). Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.Journal 17: 10.
McDowell NG, Beerling DJ, Breshears DD, Fisher RA, Raffa KF, Stitt M . (2011). The interdependence of mechanisms underlying climate-driven vegetation mortality. Trends Ecol Evol 26: 523–532.
Meyer F, Paarmann D, D’Souza M, Olson R, Glass E, Kubal M et al . (2008). The metagenomics RAST server – a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinformatics 9: 386.
Millar CI, Stephenson NL . (2015). Temperate forest health in an era of emerging megadisturbance. Science 349: 823–826.
Minot SS, Krumm N, Greenfield NB . (2015). One Codex: a sensitive and accurate data platform for genomic microbial identification. bioRxivhttps://doi.org/10.1101/027607.
Moran F, Seiichi N, Starr M . (1968). Oligogalacturonide trans-eliminase of Erwinia carotovora. Arch Biochem Biophys 125: 734–741.
Oliva J, Stenlid J, Martínez-Vilalta J . (2014). The effect of fungal pathogens on the water and carbon economy of trees: Implications for drought-induced mortality. New Phytol 203: 1028–1035.
Parks DH, Tyson GW, Hugenholtz P, Beiko RG . (2014). STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30: 3123–3124.
Pautasso M, Schlegel M, Holdenrieder O . (2015). Forest health in a changing world. Microb Ecol 69: 826–842.
Poza-Carrión C, Aguilar I, Gallego FJ, Nuñez-Moreno Y, Biosca EG, González R et al . (2008). Brenneria quercina and Serratia spp. isolated from Spanish oak trees: molecular characterization and development of PCR primers. Plant Pathol 57: 308–319.
Rackham O . (2008). Ancient woodlands: modern threats. New Phytol 180: 571–586.
Reed K . (2016). The role of the two spotted oak buprestid in Acute Oak Decline. Pract Bull Chart Inst Ecol Environ Manag 91: 33–36.
Sallé A, Nageleisen LM, Lieutier F . (2014). Bark and wood boring insects involved in oak declines in Europe: current knowledge and future prospects in a context of climate change. For Ecol Manage 328: 79–93.
Sapp M, Lewis E, Moss S, Barrett B, Kirk S, Elphinstone J et al . (2016). Metabarcoding of bacteria associated with the Acute Oak Decline syndrome in England. Forests 7: 95.
Seemann T . (2014). Prokka: rapid prokaryotic genome annotation. Bioinformatics 30: 2068–2069.
Sinclair WA . (1965). Comparisons of recent declines of white ash, oaks, and sugar maple in northeastern woodlands. Cornell Plant 20: 62–67.
Sinclair W, Lyon H . (2005) Diseases of Trees and Shrubs. Comstock Publishing Associates: Ithaca, New York, USA.
Tampakaki AP, Skandalis N, Gazi AD, Bastaki MN, Sarris PF, Charova SN et al . (2010). Playing the Harp: evolution of our understanding of hrp/hrc genes. Annu Rev Phytopathol 48: 347–370.
Thomas F . (2008). Recent advances in cause-effect research on oak decline in Europe. CAB Rev Perspect Agric Vet Sci Nutr Nat Resour 3: 1–12.
Toth IK, Pritchard L, Birch PRJ . (2006). Comparative genomics reveals what makes an enterobacterial plant pathogen. Annu Rev Phytopathol 44: 305–336.
Vansteenkiste D, Tirry L, Van Acker J, Stevens M . (2004). Predispositions and symptoms of Agrilus borer attack in declining oak trees. Annu For Sci 61: 815–823.
Versluis D, Leimena MM, Garcia JR, Andrea MMD . (2015). Mining microbial metatranscriptomes for expression of antibiotic resistance genes under natural conditions. Sci Rep 5: 11981.
Vinatzer BA, Monteil CL, Clarke CR . (2014). Harnessing population genomics to understand how bacterial pathogens emerge, adapt to crop hosts, and disseminate. Annu Rev Phytopathol 52: 19–43.
Wood DE, Salzberg SL . (2014). Kraken: ultrafast metagenomic sequence classification using exact alignments. Genome Biol 15: R46.
Yao J, Chen QL, Shen AX, Cao W, Liu YH . (2013). A novel feruloyl esterase from a soil metagenomic library with tannase activity. J Mol Catal B Enzym 95: 55–61.
Acknowledgements
This work was supported by Woodland Heritage, the Rufford Foundation, the Monument Trust, the JPG fund, HPC Wales, the Forestry Commission and DEFRA (TH0108). We thank the private land owners, public forest estate, the National Trust and the Woodland Trust for site access and allowing tissue sampling of oaks. We also thank the internal reviewers for their helpful suggestions on the manuscript. Sequence data generated in this study are available under NCBI BioProject PRJNA323828 and PRJNA321868. We thank Katy Reed and Daegan Inward for supplying Agrilus egg and the Forest Research Technical Service Unit for supplying logs and assistance with field and log trials. We also thank Lucille Rainbow and Margaret Hughes of the Centre of Genomic Research for library preparation and sequencing.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Denman, S., Doonan, J., Ransom-Jones, E. et al. Microbiome and infectivity studies reveal complex polyspecies tree disease in Acute Oak Decline. ISME J 12, 386–399 (2018). https://doi.org/10.1038/ismej.2017.170
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2017.170
This article is cited by
-
First report of bacteria causing Acute Oak Decline on Quercus robur in Slovakia
European Journal of Plant Pathology (2024)
-
Association of Dryocola boscaweniae, Gibbsiella greigii and Gibbsiella quercinecans with oak decline in Iran
European Journal of Forest Research (2024)
-
Population genomic analysis of an emerging pathogen Lonsdalea quercina affecting various species of oaks in western North America
Scientific Reports (2023)
-
Description of a novel species of Leclercia, Leclercia tamurae sp. nov. and proposal of a novel genus Silvania gen. nov. containing two novel species Silvania hatchlandensis sp. nov. and Silvania confinis sp. nov. isolated from the rhizosphere of oak
BMC Microbiology (2022)
-
Global challenges facing plant pathology: multidisciplinary approaches to meet the food security and environmental challenges in the mid-twenty-first century
CABI Agriculture and Bioscience (2021)