Abstract
Recent work has vastly expanded the known viral genomic sequence space, but the seasonal dynamics of viral populations at the genome level remain unexplored. Here we followed the viral community in a freshwater lake for 1 year using genome-resolved viral metagenomics, combined with detailed analyses of the viral community structure, associated bacterial populations and environmental variables. We reconstructed 8950 complete and partial viral genomes, the majority of which were not persistent in the lake throughout the year, but instead continuously succeeded each other. Temporal analysis of 732 viral genus-level clusters demonstrated that one-fifth were undetectable at specific periods of the year. Based on host predictions for a subset of reconstructed viral genomes, we for the first time reveal three distinct patterns of host–pathogen dynamics, where the viruses may peak before, during or after the peak in their host’s abundance, providing new possibilities for modelling of their interactions. Time series metagenomics opens up a new dimension in viral profiling, which is essential to understand the full scale of viral diversity and evolution, and the ecological roles of these important factors in the global ecosystem.
Similar content being viewed by others
Introduction
One of the major challenges in studies of viral dynamics is the absence of a phylogenetically informative universal marker, analogous to the bacterial 16S or eukaryotic 18S ribosomal RNA (rRNA) genes. To analyse temporal changes of some viral subgroups (for example, marine T4-like myoviruses or freshwater cyanomyoviruses), recent studies have used sequencing of amplicons of viral conserved structural proteins, such as capsid proteins g23 or g20 (Chow and Fuhrman, 2012; Wang et al., 2015; Yeo and Gin, 2015). However, this approach does not allow assessment of the dynamics of the whole community. A shotgun metagenomics approach does not share this limitation and provides a means to study seasonal changes without any a priori assumptions about the structure of a viral community. Using shotgun metagenomics, some attempts have been made to study viral dynamics, for example, by tracking the temporal changes of 35 individual de novo assembled viral genomes (Emerson et al., 2012), or by binning sequencing reads into assemblages (possibly at a viral family level; Bolduc et al., 2015) to study their temporal stability and/or fluctuations (Emerson et al., 2013; Bolduc et al., 2015). Although these studies have provided much-needed insight into possible scenarios of viral dynamics, there is still no global picture available of seasonal changes of viral populations and their links to other factors in an ecosystem.
Owing to the mosaic nature of viral genome organisation, assessment of viral genetic similarity is a non-trivial task. To tackle this problem, Lima-Mendez et al. in 2008 proposed a method of reticulate classification of phage genetic relatedness. The method provides means to subdivide the whole sequence space of viral metagenomics data into groups approximately corresponding to genus level of taxonomical classification. At that time the approach has been successfully used in several studies to gain deeper insight into phage biology and to connect newly assembled genomes with already known sequences (Roux et al., 2015, 2016). At the same time, it is well known that sequence relatedness within characterised viral genera can vary substantially (King et al., 2011), but in natural environments the genetic variation of newly assembled viral genomes within ‘genera’ resulting from reticulate clustering has not yet been analysed.
Along with the gaps in knowledge of global viral sequence diversity, there is a lack of information about the possible variants of bacteria–phage dynamic interactions. To date, a range of models describing behaviour of some host–pathogen relationships have been developed. First and foremost, the Kill-the-Winner model (Thingstad, 2000), which assesses populations’ changes within the framework of the classic Lotka–Volterra model. Recently, Knowles et al. (2016) have noticed discrepancies between the predictions of the model and the experimentally measured virus and host abundances in natural environments, which poses a question about the possible existence of other dynamics of host–pathogen interactions in natural microbial communities.
Here we present a detailed exploration of the structure, seasonal dynamics and functional potential of the viral community in a temperate freshwater eutrophic lake (Lough Neagh, Northern Ireland). Our novel data include 12 viral shotgun metagenomes and 13 bacterial 16 S rRNA-amplicon data sets collected over a period of 1 year (Supplementary Table 1, sheet 1). This unique collection of data allowed us to explore the range of interaction dynamics of viruses and their hosts in a natural ecosystem. We also investigate the possibility of functional manipulations of bacteria by phages by analysing auxiliary metabolic genes (AMGs), revealing that their functions are clearly different in winter compared with summer.
Materials and methods
Data availability
Raw reads from the Illumina sequencing and sequences of bacterial 16S rRNA gene amplicons are available for download from the Short Reads Archive (BioProject PRJNA350258 and PRJNA292054). Annotated viral reads and assembled sequences are also available on MetaVir and MG-RAST databases (for accession numbers see Supplementary Table 1, sheet 1).
Sample collection, processing and sequencing
Lough Neagh is a large eutrophic polymictic shallow freshwater lake located in Northern Ireland (UK). Water samples were collected from the deepest site in the lake (54°37′06″N, 6°23′43″W) at 12 time points over the period of a year (Supplementary Table 1, sheet 1) as described previously (Skvortsov et al., 2016). Some environmental parameters, such as temperature and pH at 5 m depth, were recorded at the collection site and several extra water samples were taken for chemical analysis (Supplementary Table 5, sheet 2). Sample processing steps, DNA extraction, library preparation and sequencing procedures have been described in detail previously (Skvortsov et al., 2016). Briefly, water samples were filtered through 0.22 μm filters to obtain a ‘virus-like particle’ water fraction, which was concentrated using 100 kDa filters and treated with DNAse I. Extracted and purified DNA was used for library preparation with Nextera DNA Sample Preparation kit (Illumina, San Diego, CA, USA) and sequenced from both ends with the 600-cycle MiSeq Reagent Kit v3 on MiSeq (Illumina) at the University of Cambridge DNA Sequencing facility.
Total DNA (particle sizes >0.22 μm) was extracted from 500 ml of water using a PowerWater DNA Isolation kit (MO BIO, Carlsbad, CA, USA). Partial bacterial 16S rRNA gene sequences were amplified with 909- F/1492- R primers and sequenced on a 454 GS Junior (Roche, Basel, Switzerland) with Lib-L Shotgun chemistry.
Sequencing library processing, assembly and annotation
The Illumina reads were processed with BBMap v 33.54 (http://sourceforge.net/projects/bbmap/) software, and all reads with an average Q-score<15 or containing Ns were discarded. We applied a two-step assembly strategy. First all 12 libraries were assembled separately using the graph-based assembler IDBA-UD (Peng et al., 2012) (kmer range 20–250, step—10). Next, all the libraries were combined and assembled collectively (kmer range 20–1500, step—10). This allowed us to use all available reads in the assembly to reconstruct even low-abundance viral genomes, as well as to maximise assembly effectiveness for genomes appearing only in individual libraries. After that, an additional attempt to elongate the contigs obtained in the two previous steps was made using an overlap-layout-consensus assembler with very strict parameters (CAP3; Huang and Madan (1999), overlap>2000 bp, percentage of nucleotide identity—99%). This step also reduced drastically the number of duplicated sequences. To completely remove duplicates and leave only the longest assembled contigs, we used the cd-hit (Li and Godzik, 2006) program (-c 0.98 -n 11 -d 0). For subsequent analyses, only sequences longer than 7000 bp were retained. To estimate what part of the viral population this set of contigs represented, reads from all 12 libraries were mapped onto contigs using BBMap (70% of nucleotide identity).
Open reading frames (ORFs) in the assembled contigs were predicted with MetaGeneAnnotator (Noguchi et al., 2008). For functional annotation, the contigs assembled separately from 12 libraries were uploaded to the MG-RAST (Meyer et al., 2008) and MetaVir (Roux et al., 2014) servers (please see Supplementary Table 1, sheet 1 for the accession numbers). The resulting functional annotations with SEED subsystems were downloaded from MG-RAST, percentages of all categories were calculated for each individually annotated library and were used in correlation analysis.
Raw reads obtained from the sequencing of 16S rRNA gene amplicons were processed using the QIIME pipeline v 1.8.0 (Caporaso et al., 2010) with the settings described previously (Skvortsov et al., 2016). All sequences assigned to the non-bacterial ‘Unclassified’ category and having similarity to the rRNA genes of chloroplasts were excluded from the subsequent analysis.
Identification of complete genomes
To detect end overlaps in the assembled contigs, the first 2000 bp of each contig were aligned against the whole contig’s sequence. A contig was considered complete if a repeat of at least 150 bp at its end was detected.
Analysis of contig dynamics
To assess the dynamics of individual viral genomes, the sequencing reads from each library were mapped onto sequences from the representative data set using BBMap (percentage of nucleotide identity—99, randomly selected best mapping site). The obtained number of reads mapped on a contig was normalised to the contig length and was additionally divided by the number of reads in a given library and multiplied by the mean value of the number of reads in 12 libraries. The resulting values were used as proxies of the relative abundances of viral genomes. To reduce the amount of information on the abundance of 8950 contigs, peaks of abundance were determined. Relative abundances, which were higher than the mean value of abundance for a particular genome, were considered as belonging to a peak of abundance. A small fraction of the assembled contigs had two peaks of abundance—at the start of the period studied and at its end; these were considered as a single peak of abundance spanning the winter-early spring period. To visualise the seasonal succession of viral genomes, peaks of abundance were sorted and plotted using R (http://www.R-project.org/).
Analysis of the detectability of protein-based clusters in the environment by a metagenomics method
To assess the number of reads in each sequencing library, which could potentially belong to the assembled contigs, all reads from each library were mapped onto contigs with 95% nucleotide identity. A contig was considered undetectable in a given library if no reads mapped onto it (coverage=0.0). We then analysed the protein-based clusters (see below), and considered a protein-based viral cluster (P-VC) to be undetectable in a given library if all contigs comprising it were undetectable in this library.
Clustering
For the clustering of viral contigs, a method developed by Lima-Mendez et al. (2008) was implemented. Briefly, the predicted protein sequences of contigs were aligned against themselves (‘all-to-all’ protein blast search, threshold of 50 on bitscore) and protein families were determined with the application of Markov cluster algorithm software (Enright et al., 2002; inflation factor 1.2). Next, the pairwise comparison of shared gene content between contigs was made using a hypergeometric formula, and significance was calculated with correction for multiple comparisons (threshold of 0 on significance). After that, the next round of clustering (Markov cluster algorithm, inflation factor 1.1) generated groups of related genomes. The inflation factor controls granularity of final clusters and as we analysed community structure on two levels of similarity, for protein-based clusters (highest level of organisation) we adjusted this parameter to maximise sizes of clusters. To obtain the clusters of contigs sharing nucleotide homology, this method was adjusted and the protein blast search was replaced by a nucleotide one. Thresholds were also adjusted and more strict criteria were applied (a threshold value of 5 for significance and an inflation factor of 2 were used for the second round of clustering). We then combined the results of these two clustering procedures in a single structure.
The third clustering was performed with the combined seeded sequences of isolated viruses (viral RefSeq, version 9/06/2016), contigs assembled from the publicly available metagenomes and contigs assembled in this study with settings as for the first protein clustering. The clusters obtained, which included both types of contigs—long contigs of the Lough Neagh representative data set, and seeded sequences—were transformed in pairs of long contigs and similar seeded genomes and assigned to the structure of the viral community generated in previous clustering procedures.
Assembly of publicly available freshwater metagenomes
Nine freshwater metagenomes were downloaded (Supplementary Table 1, sheet 2). Metagenomes were assembled using IDBA-UD (kmer range 20–200, step—10). Sequences longer than 10 kb were combined and seeded to clustering.
Host–bacteriophage pairs prediction
The software metaCRT (Rho et al., 2012) was used to predict CRISPR arrays in bacterial genomes (bacterial NCBI RefSeq, version of 22/08/2016). The sequences of spacers were collected, aligned against the set of long contigs, and only complete matches of the full length of spacers to contigs were allowed for the host prediction. Manual curation of predicted hosts was performed and links, which included bacteria present among Lough Neagh operational taxonomic units were left (Supplementary Table 3, sheet 4).
AMG identification
AMGs were considered to be genes that colocalised with ORFs of known viral origin on the same contig. To that end, contigs from all 12 libraries whose ontological annotation (subsystems; Overbeek et al., 2005) comprised the words ‘phage’, ‘terminase’ or ‘capsid’ were selected. Next, all functional annotations assigned to contigs selected in the previous step were summarised. The category ‘phages, prophages, transposable elements, plasmids’ was removed from the final list of AMGs as it contains structural viral proteins and common viral enzymes (Supplementary Table 4, sheet 1).
To assess changes of gene content of reconstructed viral genomes in the environment throughout the year, we evaluated and weighted the presence of functional categories of the highest annotation level of SEED subsystems for these genomes at each sample collection time point. In order to do this, viral contigs were uploaded to MG-RAST server (Supplementary Table 1, sheet 1) for annotation, and annotations of the highest level were collected for each contig. In each of the sample collection points, each functional annotation was assigned a weight equal to the relative abundance of the contig that annotated feature belonged to. Weights of all annotations of each particular functional category were summed, normalised to the sum of all weights, and clustered with dist/hclust functions of R (Euclidean distance, Ward clustering method).
Experimental verification of contigs
Experimental validation of the existence of DNA sequences of six contigs was performed using PCR amplification of specific genome regions and subsequent partial resequencing of amplicons from forward and reverse primers. The primers were designed with Primer-BLAST (Ye et al., 2012) online software (Supplementary Table 5, sheet 3 and Supplementary Figure 1). For PCR amplification, the same viral DNA samples were used as for the library preparation for Illumina sequencing. The 25 μl of PCR mixture included 1 U of DreamTaq DNA polymerase and its buffer (1x) (Thermo Fisher Scientific, Waltham, MA, USA), 0.2 mmol of each dNTP, 0.3 μmol of each primer and 8–10 ng of DNA template. PCR cycling conditions were as follows: (1) initial denaturation at 95 °C for 4 min, (2) denaturation at 95 °C for 30 s, (3) annealing at 60 °C for 30 s, (3) elongation at 72 °C for 7 min, (4) repeat steps 2–4 45 times, (5) final extension at 72 °C for 4 min. The full volumes of PCR products were loaded on 0.8% agarose gel. The lengths of amplicons were determined using the GeneRuler 1 kbp DNA ladder (Thermo Scientific) and products of required size were excised from the gel under ultraviolet light. DNA amplicons from agarose gels were extracted with High Pure PCR Product Purification kit (Roche Diagnostics, Rotkreuz, Switzerland) and sequenced at the University of Dundee DNA Sequencing and Services Facility.
Visual data exploration
To visualise pairwise genomic homology and similarity we used Easyfig v.2.2.2 (Sullivan et al., 2011).
The software package Gephi (Bastian et al., 2009) was used to visualise the results of the viral population clustering. To this end, the list of graphs (filtered pairwise comparisons of contigs with an estimation of their gene shared content) produced during DNA-based clustering was filtered in accordance to generated DNA-based viral clusters (DNA-VCs) (during this step all weak connections between contigs were removed). To the list obtained, graphs of protein clusters without DNA-VCs within them were added. These graphs were obtained from the protein-based clustering experiment. After that, a single random contig from each DNA-VCs within a given P-VCs was additionally connected to an artificial node as well as to all contigs unclustered into DNA-VCs within the same P-VC. All unique contigs, which remained fully unclustered, were transformed into a form of self-connected graphs and added to the final list of graphs, which was loaded to Gephi. To generate the picture, the ForceAtlas2 algorithm was used.
Statistical analysis
Wilcoxon–Mann–Whitney test was used to compare highest abundances of two groups of contigs: with narrow form of peaks of abundances and with wide peaks (U=8269784.5, P<0.01). Spearman's rank correlation test was used to assess the strength and direction of correlations, with a value of rho>0.5or rho<–0.5 considered as meaningful. Statistical analysis was performed in R version 3.2.2 (http://www.Rproject.org/) and using Scipy (van der Walt et al., 2011) packages for Python.
Results and discussion
Succession of viral genotypes in Lough Neagh
To generate a representative data set of viral genomic contigs that contains sequences of less abundant viruses and viruses with pronounced seasonality, we applied a hybrid assembly approach combining both assembly of individual metagenomic libraries and cross-assembly (see Materials and methods section). The final data set comprised 8950 long contigs (⩾7 kb), which accounted for 59.2% of all reads. Among these contigs, 313 were considered to be complete genomes as they had end overlaps (Supplementary Table 3, sheet 2). The integrity of several assembled contigs was verified experimentally using PCR amplification and partial resequencing by Sanger’s method (Supplementary Figure 1). These contigs were chosen mostly at random, but included one complete small 7148 bp genome of a putative temperate phage (based on the similarity of one its ORF with integrases), whose circular form was verified using PCR. Another one was a contig encompassing a CRISPR array, the accurate assembly of which was proved with resequencing.
To draw a picture of the annual succession of viruses, we determined the temporal dynamics of all individual genomic contigs (Figure 1, Supplementary Table 2, sheet 1). For visual clarity in Figure 1, we have omitted some information and retained only data on abundances, which were higher than the mean value—peaks of abundance. Most viral contigs analysed (85.4%) had a single peak of abundance during the year, and it was possible to distinguish two main types—narrow (33% of all contigs) and wide (52%) peaks of abundance. Interestingly, viruses with narrow peaks of abundance also were among the most abundant genomes in the community (Wilcoxon-Mann-Whitney test, P<0.01, see Methods). The detection of peaks of the same genomes at the beginning and the end of the 12-month period studied (Figure 1) suggests that this cycle of succession of viral species is annually repeated. After analysis of dynamic changes in contigs, we assessed their presence in the environment during the year. This study demonstrated that only 39.1% of viruses (3502 partial genomes) persisted in the lake throughout the year, whereas most viruses were undetectable by metagenomics methods at one or more time points. The characteristics of the dynamic changes in Lough Neagh viral populations should not be considered specific only to this particular environment; on the contrary, it is likely to be an instance of a universal phenomenon, reflecting processes common to different ecosystems on the global scale. For example, in a previous study of marine myoviruses it was demonstrated that during 3 consecutive years a number of viral genomes appeared only once a year at specific seasons and that only 25% of myoviruses persisted in the environment (Chow and Fuhrman, 2012). A study of viral dynamics in the hypersaline Lake Tyrell also revealed the presence of two types of viruses—those considered persistent and those detectable only at specific time points (Emerson et al., 2012, 2013).
Succession of 8950 assembled contigs throughout a year. Each row in the left panel of the picture presents information about peaks of abundance for individual contigs. For each library, a dot was placed if the peak of abundance for a particular contig was attributed to this library; otherwise, an empty space was left. Jitter was applied to distribute dots belonging to different contigs within a single column. The right panel schematically depicts dynamic changes in individual contigs to provide illustrations of different observed cases. Contig identification numbers are specific to this Figure and do not correspond to contig IDs used elsewhere in the study.
Structure of the viral community
Next, we characterised the structure of the viral community. Reticulate classification of viral sequences allows estimation of the relatedness of genomes by assessing shared gene content (Lima-Mendez et al., 2008). This method uses comparisons of amino-acid sequences, allowing grouping of viral genomes, which do not necessarily have nucleotide homology (P-VCs) into clusters that approximately correspond to viral genera (Lima-Mendez et al., 2008; Roux et al., 2015, 2016; Paez-Espino et al., 2016). One of the goals of our analysis was to additionally divide assembled genomes within these clusters into subgroups of homologous sequences. To this end, we modified the method of reticulate classification and performed a second clustering using comparison of nucleotide sequences (DNA-VCs). As the result of this, the majority of contigs were organised into 732 P-VCs (Figure 2,Supplementary Table 3, sheet 1) consisting of 2 to 696 members, whereas 1198 contigs (13.4%) remained as singletons. Within the P-VCs, sequences were arranged into subclusters on the basis of sequence homology in DNA-VCs (1811 clusters in total, Supplementary Figure 2). The analysis of genome relatedness within this double-clustered structure showed that the similarity of viral genomes within P-VCs varied, which additionally characterises the community studied. For example, genomes within P-VC_2 (Figure 3) were very similar and retained some nucleotide homology across the whole cluster/viral genus. This could point to the possibility that these viruses underwent gene reshuffling more often than they accumulated point mutations. By contrast, genomes within P-VC_20 (Figure 3) are likely to have evolved under different constraints, as the genomes detected were more distantly related even in smaller groups (DNA-VCs), retaining only protein similarity between genomes from different DNA-based clusters.
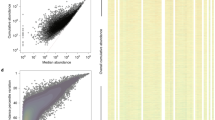
(a) An overview of the viral sequence space organisation in the community. Each dot represents an individual from 8950 assembled contigs. These contigs can be (i) genetically unique and fully unclustered, (ii) clustered into DNA-VCs (middle-size clustering level), (iii) clustered into P-VC being within DNA-VCs or being unclustered within DNA-VCs (large-size clustering level). The outer ring of light grey dots is constituted by unique individual genomes that are not members of P-VCs (variant i). Each separate group of dots within the inner circle represents an individual P-VC (variant iii). All DNA-VCs as well as all unclustered contigs within each P-VC were joined to an artificial central node. To avoid confusion with colours, larger P-VCs were arbitrary coloured to provide more information about their inner structure. Contigs comprising DNA-VCs within P-VCs are coloured either in orange (relatively bigger) or in dark grey (relatively smaller), whereas unclustered into DNA-VCs contigs are coloured in green. By default, a dark grey colour is used for all contigs within other P-VCs. Three distinct types of clusters are indicated: (b) P-VC, which includes sequences with nucleotide homology organised in DNA-VCs, as well as unclustered genomes (mixed type); (c) P-VC, which aggregates (mostly) unclustered in DNA-VCs genomes (contigs within these clusters have only similarity at the protein level); (d) P-VC, which comprises mostly DNA-VCs.
Examples of genome relatedness within the double-clustered structure of the viral sequence space organisation. Full-length nucleotide (a, b) and amino-acid (c) alignments of representatives from DNA-VCs of two P-VCs are shown on the left side of the Figure, the degree of homology between aligned fragments is colour coded. All genomes presented are complete, if not stated otherwise. The dynamic changes in abundance of these individual genomes are shown on the right (d). (a) Genomes preserving nucleotide homology between different DNA-VCs within a single P-VC. Two upper genomes were clustered into one DNA-VC, whereas the other nine genomes belonged to different DNA-VCs within one P-VC_2. (b) Genome alignments of three members of DNA-VC-20. (c) Genomic map of protein similarity (tblastx) between three representatives from different DNA-VCs (including DNA-VC_20 from panel b) within a single P-VC_20. These genomic sequences do not have nucleotide homology.
The temporal dynamics of the clusters adds a new dimension to our understanding of viral biodiversity. We explored how contigs, the majority of which had distinct seasonality, were distributed between clusters and found that large P-VCs (with >20 partial genomes) persisted during the year, although they could include DNA-VCs with specific seasonalities. Thus, although certain genetic variants could appear for short periods only, the group of viruses they belonged to could be detected throughout the whole year. At the same time, smaller P-VCs could be abundant only during particular periods of the year (382 of all P-VCs, that is, 52.2%; 51 of these included >4 contigs, Supplementary Table 3, sheet 1) and one-third of these (131 P-VCs) were undetectable by metagenomics technologies in other periods (Materials and methods section). Moreover, we found that about one-fifth of all P-VCs (164, 22.4%) were undetectable at specific time points.
Identification of related sequences among known phages
To identify how the de novo assembled contigs were related to known viruses, complete viral genomes from RefSeq were seeded to a standard reticulate classification (see Materials and methods section). We also included in the analysis 488 long contigs (>10 kb) assembled from nine published viral metagenomes originating from freshwater environments in different continents (Europe (Roux et al., 2012), North America (Green et al., 2015; Watkins et al., 2015), Africa (Fancello et al., 2013), Asia (Tseng et al., 2013), Supplementary Table 1, sheet 2). The fact that only 48 RefSeq viruses were assigned to reconstructed viral genomes (Supplementary Table 3, sheet 3) and 18 of them included in 19 DNA-VCs from our data set reveals just how limited exploration of freshwater viral diversity has been. Among these were eight species of Cellulophaga phages, eight Pseudomonas phages and seven cyanophages. We also identified one contig with similarity to an algal virus virophage—Phaeocystis globosa virus virophage (Supplementary Table 3, sheet 3). The seeding of long contigs assembled from other freshwater metagenomes allowed us to determine that 106 of them (21.7%) were related to the viruses in Lough Neagh (Supplementary Table 3, sheet 1). In total, 69 DNA-VCs (from 40 P-VCs) recruited contigs from other freshwater environments. One of the P-VCs (P-VC_19) seemed to represent a ‘core freshwater cluster’ of genomes, as it recruited viral sequences from five freshwater reservoirs from very distant sites: Lough Neagh (British Isles), Lake Michigan (North America), Lakes Pavin and Bourget (Continental Europe) and the Feitsui freshwater reservoir in Taiwan (Asia). Several sequences from this ‘core’ cluster were related to Cellulophaga phage 46:1 (Holmfeldt et al., 2013) (Supplementary Table 3, sheet 1).
Although, the method of co-clustering of viral genomes allows to detect more distant relatives among known sequences, we additionally explored results of read-mapping approach of MetaVir pipeline to identify eukaryotic viruses, which were less likely to assemble because of predominance of bacteriophages in the environment. The highest number of reads of eukaryotic viruses were assigned to the Phycodnaviridae family of algae viruses represented by all genera with Chlorovirus as the most abundant one. Among other viruses of eukaryotic organisms, sequences for several gigantic viruses of family Mimiviridae, such as amoebic Acanthamoeba polyphaga moumouvirus and flagellate Cafeteria roenbergensis virus, were found. Sequences related to viruses of vertebrate and invertebrate animals of families Iridoviridae, Herpesvirales and Poxviridae were detected as well.
Dynamic relationships of viruses and their predicted hosts
To gain insights into the biology of the reconstructed viruses, we predicted their bacterial hosts using a sequence-based bioinformatics method of CRISPR matching. In a recent benchmarking analysis, CRISPR matches yielded the highest accuracy (92%) of all tested bioinformatics approaches designed to link phages to their hosts (Edwards et al., 2016). Throughout the year, we generated structural profiles of the bacterial community using methods of amplicon-based metagenomics (Materials and methods section, Supplementary Figure 3,Supplementary Table 2, sheet 2). Among known caveats of this approach is that the resolution provided by 16S amplicons is not necessarily sufficient to distinguish ecotypes, which have identical 16S sequences, but different genomes and may demonstrate individually distinct dynamics in the ecosystem. To link reconstructed phage genomes to their potential hosts, we identified CRISPR arrays in the complete genome sequences of bacterial species that were closely related to the operational taxonomic units detected in Lough Neagh by using selected bacterial genomes from the database, and matched those spacers to our reconstructed viral genomes (see Materials and methods section). Hosts were predicted for 225 of the 8950 reconstructed viral genomes. For several contigs, up to three potential bacterial hosts of different operational taxonomic units were assigned (possible viral generalists), therefore in total we found 260 phage–host pairs (Supplementary Table 3, sheet 4). Although the database bacteria were isolated from different locations and have never been exposed to the Lough Neagh phages, we presumed that their recent ancestors were indeed infected by close relatives of these phages, as evidenced by the 100% identical CRISPR spacers. Viruses tend to be species or strain specific, and when they do change their host tropism, they mostly switch to taxonomically very closely related hosts (Popa et al., 2017). This is an indirect approach to predict phage–host pairs, but we believe that it provides accurate insights into phage–bacteria relationships for the minority of cases where hits were found.
The contigs with the hosts assigned belonged to 131 DNA-VCs of 97 P-VCs (13.4%). The analysis of P-VCs showed that, although viruses from a given cluster usually infect a single dominant bacterial taxonomic group, there were also clusters with predicted hosts from up to five different classes and two different phyla. This finding supports the idea that, although the majority of genetically related viruses have a narrow host repertoire, there are also generalist viruses and viral genera, which can prey on hosts across bacterial taxonomic borders (Malki et al., 2015, Peters et al., 2015; Knowles et al., 2016; Roux et al., 2016).
We studied dynamic changes in viral contigs and presumed hosts (operational taxonomic units) to identify possible patterns of their interactions in a natural environment. In order to do that, we plotted the distribution of highest abundance of reconstructed viral genomes in relation to the maximum of corresponding bacterial abundance (Figure 4). In accordance with the ‘Kill-the-Winner’ (KtW) model of host–pathogen relationships (Thingstad, 2000), the dynamics of bacteria and their viruses are co-dependent, and the peak of abundance of a virus should appear with some delay after the peak of abundance of its host. The correlational analysis (Spearman’s rank correlation, rho>0.5) of relationships of identified viral contig–bacterial operational taxonomic unit pairs demonstrated that 54 pairs behaved in accordance with the KtW model (20.8%, see Materials and methods section) and the dynamic changes of 28 other phage–host pairs coincided (31.5% in total). But this plot demonstrates that many viruses peaked before their hosts. We performed correlational analysis and found that in 43 pairs (16.5%), the increase of viral abundance was indeed followed by the increase of host density. To our knowledge, this is one of the few examples when viral abundance peaks occurring before the peaks of their cognate host have been observed in natural environment.
Positions of the peaks of abundance for 260 viral genomes in relation to maximum abundance of their predicted host bacteria. The numbers of viruses in groups, organised by the distance (in sample collection intervals) of their peaks of abundance from the peaks of abundance of host bacteria, are plotted as black dots (•), whereas host abundance maximum is used as a reference point and represented by a dashed line (- - -).
Next, we performed an investigation of the existing literature, looking for evidence where this counter-intuitive pattern may have been registered. Wilson et al. (1998) presented time series data of marine mesocosms where, after addition of phosphorus to the environment, and before the development of a peak of cyanobacterial abundance as a response, there was a distinguishable high peak of abundance of viral particles. Similarly, in a time-series study of marine Synechococcus and cyanophage populations, although this observation was outside the scope of the article, preceding peaks of viral abundances were noticeable and were repeated on several occasions over the period studied (McDaniel et al., 2002).
Moreover, this dynamic pattern was modelled for situation of effective defence of prey from low-offence predators (Cortez and Weitz, 2014). We offer several possible mechanisms of such defence that might explain the observed dynamics. First, they might be explained by the development of resistance of bacteria to the phage, for example, by acquisition of CRISPR spacers or modification of their receptor binding proteins, facilitating subsequent expansion of the bacterial population. However, mechanisms of resistance acquisition can also be due to super-infection exclusion caused by the switch of phages from the lytic to the temperate state. Recently, Knowles et al. (2016) proposed an extension to the KtW model—the Piggyback-the-Winner (PtW) model, in accordance with which ‘temperateness is favoured at high host densities as viruses exploit their hosts through lysogeny rather than killing them’. We might expect that the dynamic pattern identified could be a result of phage–host interactions in accordance with this PtW model. Moreover, it was recently revealed that some viruses can communicate with each other via short quorum-sensing peptides, where an increase of the peptide concentration causes switch of temperate phages from the lytic to the lysogenic state (Erez et al., 2017). It is possible that this mechanism could also explain the ‘early’ loss of viruses from the environment, as observed in our study.
Environmental parameters in Lough Neagh
To discover as many drivers of viral community changes as possible, we characterised bacterial community composition and environmental parameters in the lake’s ecosystem (Supplementary Table 2, sheet 2 and Supplementary Table 5, sheet 1). Predominance of cyanobacteria in eutrophic Lough Neagh was detected in summertime (Supplementary Figure 3). Comparative analysis of the dynamics of the bacterial populations and changes in physical and chemical parameters showed that temperature was likely to be the main driver of changes in the bacterial community under study (Spearman’s rank correlation, rho>0.5, Supplementary Table 5, sheet 1). We also found that, surprisingly, the bacterial community did not react to changes in phosphorus concentration—the main limiting factor for growth of microbial populations in freshwater environments (Doering et al., 1995; Correll, 1999). Apparently, in this eutrophic lacustrine ecosystem the main limiting factor is different, which is in accordance with previous findings that in Lough Neagh nitrogen loading can have a stronger long-term impact than phosphorus on lake eutrophication (Buntig et al., 2007).
Viral AMGs and their changes throughout the year
Viruses can carry AMGs that augment their fitness by affecting host metabolism (Breitbart et al., 2007). As it is not possible to exclude the occurrence of bacterial genes caused by generalised transduction events and the presence of gene transfer agents in phage metagenomes, we applied strict criteria for the detection of phage-associated AMGs. Metabolic genes were considered as AMGs only if they were colocalised on the same contig with ORFs having similarity to known phage genes (such as structural genes, see Materials and methods section). Contigs from all 12 assembled libraries were analysed and 189 phage-associated AMGs were identified (Supplementary Figure 4, Supplementary Table 4, sheet 1). The attribution of AMGs to SEED subsystems showed that freshwater viruses in Lough Neagh had acquired genes from a wide variety of metabolic pathways, related to almost all aspects of bacterial life, as genes from 25 out of 30 of subsystems were found in viral genomes.
To assess how the appearance of various genes (functional categories of SEED subsystems; Overbeek et al., 2005) in the viral population depended on the dynamics of the bacterial community and on environmental parameters, a correlational analysis was performed (Spearman’s rank correlation, rho>0.5, Supplementary Table 4, sheet 2). An increase in relative abundance of ‘Genes of temperate phages’ in the summer viral community was detected, supporting findings from previous studies obtained by using different methods (Laybourn-Parry et al., 2007; Palesse et al., 2014; Knowles et al., 2016). We also identified correlations between the appearance of genes of ‘Oxidative stress response regulation’ in the summertime and Cyanobacteria changes and alkalinity fluctuations (Figure 5b). Cyanobacteria undergo oxidative stress more often than heterotrophic bacteria because of their photosynthetic ability (Latifi et al., 2009), and it was shown that marine cyanophages can carry genes involved in photoprotection, such as those encoding high light inducible proteins (Ma et al., 2014). ‘Oxidative stress response regulation’ genes of freshwater viruses identified in this study included a wide range of molecules guarding living organisms from oxidative damage: iron and manganese superoxide dismutases, peroxidase, catalase, ferroxidase, rubrerythrin, etc. In contrast with high light inducible proteins of marine cyanophages, which are tightly connected with photosystem formation and functioning (Komenda and Sobotka, 2016), antioxidant defence genes of freshwaters are more general and include cytoplasmic, mitochondrial and chloroplast-associated molecules. Other notable correlations were detected between the abundance of the Bacteroidetes phylum, the Verrucomicrobiae class and genes of ‘Quorum sensing and biofilm formation’ (Figure 5c). The relative abundance of this functional category did not correlate with any other taxa or any environmental parameter, suggesting that this type of phage manipulation is specific to these clades.
(a) Clustering of 12 viromes based on functional annotations of 8950 reconstructed genomes, weighted by genome's abundance. (b) Dynamic changes of relative abundance of Cyanobacteria (black, rho=0.6) and ‘Regulation of oxidative stress response’ functional category genes (red). (c) Dynamic changes of relative abundance of the Bacteroidetes phylum (blue, rho=0.68), the Verrucomicrobiae class (black, rho=0.67) and genes of ‘Quorum sensing and biofilm formation’ (red) functional category.
To further investigate the seasonal dependence of viral functional potential we annotated separately reconstructed viral genomes on MG-RAST server and clustered functional annotations, weighted by contig relative abundance in the community (see Materials and methods section, Figure 5a). We identified that reconstructed viral genomes clearly differed in the winter-early spring and summer-autumn periods by functions, these two groups being largely subdivided in accordance with calendar seasons. These findings could additionally point to the specialisation of viruses to their hosts through acquisition of specific AMGs.
Conclusions
Overall, this study changes our understanding of viral diversity by demonstrating the transient nature of most viral groups of genomes in an ecosystem. This variation of the whole metagenomic content of the environment between different seasons/months should also be considered when assessing the criteria for the sampling completeness of an ecosystem. Visualisation of the genetic relationships between viruses further characterises the community as a whole and points to the diversity of evolutionary constraints in a natural environment. Besides providing much-needed insight into freshwater viral sequence diversity and ecosystem organisation, our research offers a basis for long-term studies on the stability of individual viral genomes, on the repeatability of seasonal cycles, and on their interplay with bacterial host communities. In our study, we analysed only enveloped DNA viruses existing as viroid particles in the environment. Previous studies have highlighted that viruses can also subsist inside their host cell for prolonged periods of time, so it will be interesting to analyse time series of combined free-viroid and induced viromes side-by-side (Maurice et al., 2011). Moreover, including time-series experiments of RNA viruses can also provide complementary insight into the dynamics of viral communities in the future.
References
Bastian M, Heymann S, Jacomy M . (2009). Gephi: an open source software for exploring and manipulating networks. International AAAI Conference on Weblogs and Social Media: San Jose, CA, USA.
Bolduc B, Wirth JF, Mazurie A, Young MJ . (2015). Viral assemblage composition in Yellowstone acidic hot springs assessed by network analysis. ISME J 9: 2162–2177.
Breitbart M, Thompson LR, Suttle CA, Sullivan MB . (2007). Exploring the vast diversity of marine viruses. Oceanography 20: 135–139.
Buntig L, Leavitt PR, Gibson CE, McGee EJ, Hall VA . (2007). Degradation of water quality in Lough Neagh, Northern Ireland, by diffuse nitrogen flux from a phosphorus-rich catchment. Limnol Oceanogr 52: 354–369.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK et al. (2010). QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7: 335–336.
Chow CE, Fuhrman JA . (2012). Seasonality and monthly dynamics of marine myovirus communities. Environ Microbiol 14: 2171–2183.
Correll DL . (1999). Phosphorus: a rate limiting nutrient in surface waters. Poult Sci 78: 674–682.
Cortez MH, Weitz JS . (2014). Coevolution can reverse predator-prey cycles. Proc Natl Acad Sci USA 111: 7486–7491.
Doering PH, Oviatt CA, Nowicki BL, Klos EG, Reed LW . (1995). Phosphorus and nitrogen limitation of primary production in a stimulated estuarine gradient. Mar Ecol Prog Ser 124: 271–287.
Edwards RA, McNair K, Faust K, Raes J, Dutilh BE . (2016). Computational approaches to predict bacteriophage-host relationships. FEMS Microbiol Rev 40: 258–272.
Emerson JB, Thomas BC, Andrade K, Allen EE, Heidelberg KB, Banfield JF . (2012). Dynamic viral populations in hypersaline systems as revealed by metagenomic assembly. Appl Environ Microbial 78: 6309–6320.
Emerson JB, Thomas BC, Andrade K, Heidelberg KB, Banfield JF . (2013). New approaches indicate constant viral diversity despite shifts in assemblage structure in an Australian hypersaline lake. Appl Environ Microbial 79: 6755–6764.
Enright AJ, Van Dongen S, Ouzounis CA . (2002). An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res 30: 1575–1584.
Erez Z, Steinberger-Levy I, Shamir M, Doron S, Stokar-Avihail A, Peleg Y et al. (2017). Communication between viruses guides lysis-lysogeny decisions. Nature 541: 488–493.
Fancello L, Trape S, Robert C, Boyer M, Popgeorgiev N, Raoult D et al. (2013). Viruses in the desert: a metagenomic survey of viral communities in four perennial ponds of the Mauritanian Sahara. ISME J 7: 359–369.
Green JC, Rahman F, Saxton MA, Williamson KE . (2015). Metagenomic assessment of viral diversity in Lake Matoaka, a temperate, eutrophic freshwater lake in southeastern Virginia, USA. Aquat Microb Ecol 75: 117–128.
Holmfeldt K, Solonenko N, Shah M, Corrier K, Riemann L, Verberkmoes NC et al. (2013). Twelve previously unknown phage genera are ubiquitous in global oceans. Proc Natl Acad Sci USA 110: 12798–12803.
Huang X, Madan A . (1999). CAP3: a DNA sequence assembly program. Genome Res. 9: 868–877.
King AMQ, Lefkowitz E, Adams MJ, Carstens EB . (2011) Virus Taxonomy: Ninth Report of the International Committee on Taxonomy of Viruses. Elsevier: Amsterdam, The Netherlands.
Knowles B, Silveira CB, Bailey BA, Barott K, Cantu VA, Cobián-Güemes AG et al. (2016). Lytic to temperate switching of viral communities. Nature 531: 466–470.
Komenda J, Sobotka R . (2016). Cyanobacterial high-light-inducible proteins – protectors of chlorophyll-protein synthesis and assembly. BBA-Bioenerget 1857: 288–295.
Latifi A, Ruiz M, Zhang CC . (2009). Oxidative stress in cyanobacteria. FEMS Microbiol Rev 33: 258–278.
Laybourn-Parry J, Marshall WA, Madan NJ . (2007). Viral dynamics and patterns of lysogeny in saline Antarctic lakes. Polar Biol 30: 351–358.
Li W, Godzik A . (2006). Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22: 1658–1659 2006.
Lima-Mendez G, Van Helden J, Toussaint A, Leplae R . (2008). Reticulate representation of evolutionary and functional relationships between phage genomes. Mol Biol Evol 25: 762–777.
Ma Y, Allen LZ, Palenik B . (2014). Diversity and genome dynamics of marine cyanophages using metagenomic analyses. Environ Microbiol Rep 6: 583–594.
Malki K, Kula A, Bruder K, Sible E, Hatzopoulos T, Steidel S et al. (2015). Bacteriophages isolated from Lake Michigan demonstrate broad host-range across several bacterial phyla. Virology J 12: 164.
Maurice CF, Mouillot D, Bettarel Y, Wit RD, Sarmento H, Bouvier T . (2011). Disentangling the relative influence of bacterioplankton phylogeny and metabolism on lysogeny in reservoirs and lagoons. ISME J 5: 831–843.
McDaniel L, Houchin LA, Williamson SJ, Paul JH . (2002). Lysogeny in marine Synechococcus. Nature 415: 2002.
Meyer F, Paarmann D, D’Souza M, Olson R, Glass EM, Kubal M et al. (2008). The metagenomics RAST server – a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinform 9: 386.
Noguchi H, Taniguchi T, Itoh T . (2008). MetaGeneAnnotator: detecting species-specific patterns of ribosomal binding site for precise gene prediction in anonymous prokaryotic and phage genomes. DNA Res 15: 387–396.
Overbeek R, Begley T, Butler RM, Choudhuri JV, Chuang HY, Cohoon M et al. (2005). The subsystems approach to genome annotation and its use in the project to annotate 1000 genomes. Nucleic Acids Res 33: 5691–5702.
Paez-Espino D, Eloe-Fadrosh EA, Pavlopoulos GA, Thomas AD, Huntemann M, Mikhailova N et al. (2016). Uncovering Earth’s virome. Nature 536: 425–430.
Palesse S, Colombet J, Pradeep Ram AS, Sime-Ngando T . (2014). Linking host prokaryotic physiology to viral lifestyle dynamics in a temperate freshwater lake (Lake Pavin, France). Microb Ecol 68: 740–750.
Peng Y, Leung HCM, Yiu SM, Chin FYL . (2012). IDBA-UD: a de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinformatics 28: 1420–1428.
Peters DL, Lynch KH, Stothard P, Dennis JJ . (2015). The isolation and characterisation of two Stenotrophomonas maltophilia bacteriophages capable of cross-taxonomic order infectivity. BMC Genomics 16: 664.
Rho M, Wu Y, Tang H, Doak TG, Ye Y . (2012). Diverse CRISPRs evolving in human microbiomes. PLoS Genet 8: e1002441.
Popa O, Landan G, Dagan T . (2017). Phylogenomic networks reveal limited phylogenetic range of lateral gene transfer by transduction. ISME J 11: 543–554.
Roux S, Brum JR, Dutilh BE, Sunagawa S, Duhaime MB, Loy A et al. (2016). Ecogenomics and potential biogeochemical impacts of globally abundant ocean viruses. Nature 537: 689–693.
Roux S, Enault F, Robin A, Ravet V, Personnic S, Theil S et al. (2012). Assessing the diversity and specificity of two freshwater viral communities through metagenomics. PLoS ONE 7: e33641.
Roux S, Hallam SJ, Woyke T, Sullivan MB . (2015). Viral dark matter and virus-host interactions resolved from publicly available microbial genomes. eLife 4: 1–20 (2015).
Roux S, Tournayre J, Mahul JA, Debroas D, Enault F . (2014). Metavir 2: new tools for viral metagenome comparison and assembled virome analysis. BMC Bioinform 15: 76.
Skvortsov T, de Leeuwe C, Quinn JP, McGrath JW, Allen CCR, McElarney Y et al. (2016). Metagenomic characterisation of the viral community of Lough Neagh, the largest freshwater lake in Ireland. PLoS ONE 11: e0150361.
Sullivan MJ, Petty NK, Beatson SA . (2011). Easyfig: a genome comparison visualizer. Bioinformatics 27: 1009–1010.
Thingstad TF . (2000). Elements of a theory for the mechanisms controlling abundance, diversity, and biogeochemical role of lytic bacterial viruses in aquatic systems. Limnol Oceanogr 45: 1320–1328.
Tseng CH, Chiang PW, Shiah FK, Chen YL, Liou JR, Hsu TC et al. (2013). Microbial and viral metagenomes of a subtropical freshwater reservoir subject to climatic disturbances. ISME J 7: 2374–2386.
van der Walt S, Colbert SC, Varoquaux G . (2011). The NumPy array: a structure for efficient numerical computation. Compu Sci Eng 13: 22–30.
Wang MN, Ge XY, Wu YQ, Yang XL, Tan B, Zhang YJ et al. (2015). Genetic diversity and temporal dynamics of phytoplankton viruses in East Lake. China Virol Sin 30: 290–300.
Watkins SC, Kuehnle N, Ruggeri CA, Malki K, Bruder K, Elayyan J et al. (2015). Assessment of a metaviromic dataset generated from nearshore Lake Michigan. Mar Freshwater Res 67: 1700–1708.
Wilson WH, Turner S, Mann NH . (1998). Population dynamics of phytoplankton and viruses in a phosphate-limited mesocosm and their effect on DMSP and DMS production. Estuar Coast Shelf Sci 46: 49–59.
Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden TL . (2012). Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinformatics 13: 134.
Yeo BH, Gin KY . (2015). Population dynamics of cyanomyovirus in a tropical eutrophic reservoir. Microbes Environ 30: 12–20.
Acknowledgements
We thank Dr Catherine Watson and Ms Hanna Cromie for her help with the collection of the samples. The project was funded by Leverhulme Trust Grant RPG-2013-040. BED and KA were supported by the Netherlands Organization for Scientific Research (NWO) Vidi grant 864.14.004. All data will be available from SRA, MetaVir and MG-RAST databases immediately after the publication date (for accession numbers see Supplementary Table 1, sheet 1).
Author contributions
LAK, TS and KA designed the study; TS processed water samples; KA and TS performed bioinformatics analyses of data; KA, TS, JPQ, JWG, CCRA, BED, YM, CW and LAK discussed results and wrote and edited the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Arkhipova, K., Skvortsov, T., Quinn, J. et al. Temporal dynamics of uncultured viruses: a new dimension in viral diversity. ISME J 12, 199–211 (2018). https://doi.org/10.1038/ismej.2017.157
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2017.157
This article is cited by
-
Interactions between bacterial and phage communities in natural environments
Nature Reviews Microbiology (2022)
-
Viral diversity is linked to bacterial community composition in alpine stream biofilms
ISME Communications (2022)
-
Host population diversity as a driver of viral infection cycle in wild populations of green sulfur bacteria with long standing virus-host interactions
The ISME Journal (2021)
-
Metaviromics coupled with phage-host identification to open the viral ‘black box’
Journal of Microbiology (2021)
-
Depth-related variability in viral communities in highly stratified sulfidic mine tailings
Microbiome (2020)