Abstract
Resource availability influences marine microbial community structure, suggesting that population-specific resource partitioning defines discrete niches. Identifying how resources are partitioned among populations, thereby characterizing functional guilds within the communities, remains a challenge for microbial ecologists. We used proteomic stable isotope probing (SIP) and NanoSIMS analysis of phylogenetic microarrays (Chip-SIP) along with 16S rRNA gene amplicon and metagenomic sequencing to characterize the assimilation of six 13C-labeled common metabolic substrates and changes in the microbial community structure within surface water collected from Monterey Bay, CA. Both sequencing approaches indicated distinct substrate-specific community shifts. However, observed changes in relative abundance for individual populations did not correlate well with directly measured substrate assimilation. The complementary SIP techniques identified assimilation of all six substrates by diverse taxa, but also revealed differential assimilation of substrates into protein and ribonucleotide biomass between taxa. Substrate assimilation trends indicated significantly conserved resource partitioning among populations within the Flavobacteriia, Alphaproteobacteria and Gammaproteobacteria classes, suggesting that functional guilds within marine microbial communities are phylogenetically cohesive. However, populations within these classes exhibited heterogeneity in biosynthetic activity, which distinguished high-activity copiotrophs from low-activity oligotrophs. These results indicate distinct growth responses between populations that is not apparent by genome sequencing alone.
Similar content being viewed by others
Introduction
It is well established that heterotrophic marine microbes have a critical role in global carbon cycling, remineralizing carbon via heterotrophic respiration and transferring phytoplankton-derived organic carbon to higher trophic levels via secondary microbial production (Whitman et al., 1998; Azam and Malfatti, 2007). However, models of marine microbial systems have yet to progress substantially beyond ‘black box’ approaches that quantify whole-community processes to more complex models of ecosystem processes, which incorporate community structure and functional understandings of individual microbial taxa (Ducklow, 2000; Yokokawa and Nagata, 2010). This progress is currently limited by our understanding of how diverse populations within marine microbial communities interact and drive ecological processes.
Parallel lines of inquiry have examined microbial diversity across environmental gradients (that is, spatial, temporal and resource gradients) by quantifying community and population-level changes in 16S rRNA gene, metagenomic, metatranscriptomic and metaproteomic data. Taxonomic surveys have revealed structural shifts in microbial communities that correlate with physical and biogeochemical gradients (Morris et al., 2012; Dinasquet et al., 2013), as well as annually recurring resource gradients defined by biotic and abiotic factors, including vertical stratification, upwelling events and phytoplankton blooms (Fuhrman et al., 2006; Treusch et al., 2009; Gilbert et al., 2012). Structural changes in the functional repertoire of marine microbial communities have also been identified across spatial scales, including transitions in carbon metabolism along depth profiles (DeLong et al., 2006) and along a transect from higher-nutrient coastal waters to a more oligotrophic oceanic gyre (Morris et al., 2010). Placing these results within the framework of niche theory (Whittaker, 1972), these studies suggest that specific adaptations or ecological strategies of distinct microbial populations enable successful exploitation of environmental niches.
The application of transcriptomic and proteomic approaches to characterize in situ gene expression has enabled the development of hypotheses concerning the ecological strategies of these diverse populations. Analysis of microbial transcriptomes taken from southeastern US coastal waters identified gene expression patterns among coexisting populations, suggesting that ecological strategies fall along a continuum between slower growth coupled with metabolic specialization to faster growth and higher metabolic diversity (Gifford et al., 2013).
A study of North Sea waters (Teeling et al., 2012) examined the role of phytoplankton blooms, and, in turn, the temporal release of different resources, in shaping microbial community structure with combined taxonomic, functional and environmental observations. Principal findings included early bloom abundance and carbohydrate utilization enzyme expression by Flavobacteriia, mid and late bloom increases in Gammaproteobacteria populations, and the post or pre-phytoplankton bloom dominance of Alphaproteobacteria. Several mesocosm studies of induced phytoplankton blooms have explicitly examined the turnover of bloom-derived dissolved organic carbon (DOC), which includes substantial amounts of carbohydrates exuded during rapid bloom growth, as well as protein, amino acids and lipids that are released as cellular debris as blooms collapse (Norrman et al., 1995; Biddanda and Benner, 1997; Myklestad, 2000; Meon and Kirchman, 2001). Taken together, the results of these studies indicate that microbial community structure is influenced by both the concentration and type of substrates that are available. Therefore, defining how substrates are partitioned among discrete populations is a prerequisite for relating community structure to biogeochemical cycling.
The use of stable isotope probing has emerged as a powerful technique for linking biochemical functions to specific populations within complex microbial communities. Traditionally, this technique has utilized density-based separation of light, unlabeled and heavy, labeled, RNA and DNA to identify substrate-assimilating populations (Dumont and Murrell, 2005). Recently developed techniques utilize mass spectrometry to quantify stable isotope incorporation into rRNA or newly synthesized proteins, offering advantages in throughput and sensitivity. Chip-stable isotope probing (SIP) quantifies isotopic labeling of rRNA hybridized to a microarray (Mayali et al., 2012, 2013, 2014). Proteomic SIP utilizes exhaustive database searching to identify peptides and quantify their isotopic enrichment (Pan et al., 2011; Hyatt and Pan, 2012; Wang et al., 2013; Bryson et al., 2016). In this study we utilized Chip-SIP and proteomic SIP to describe the assimilation of six 13C-labeled substrates (glucose, starch, acetate, lipids, amino acids and protein) of importance to heterotrophic marine microbes (Geider and Roche, 2002; Kirchman, 2003; Nagata, 2008) by planktonic microbial populations in surface sea water sampled in Monterey Bay, CA. By combining this substrate assimilation data with metagenomics and 16S rRNA gene amplicon sequencing, we were able to examine the accompanying shifts in taxonomic composition of this coastal marine microbial community during these microcosm incubations.
Although previous studies have examined changes in both microbial community structure (Gómez-Consarnau et al., 2012) and function (McCarren et al., 2010) following the addition of high- and low-molecular-weight DOC substrates, the combination of SIP and sequencing approaches in this study enabled direct comparisons between substrate assimilation and relative abundance shifts, and allowed us to address several questions concerning how best to define functional guilds within these communities. In other words, do related populations assimilate the same substrates, and are taxonomic assignments indicative of functional guild membership? Do relative abundance shifts determined by DNA-sequencing approaches consistently reflect direct substrate assimilation measurements? And finally, are discrepancies between substrate assimilation into ribonucleotide and protein biomass indicative of underlying metabolic mechanisms that differentiate the environmental growth strategies of populations?
Materials and methods
Sample collection, processing and incubation
Surface water was collected from Monterey Bay (36°N 53.387′–121°W 57.257′, ~10 km from shore; Supplementary Figure S1) on 14 October 2013 while on board the R/V Rachel Carson as previously described (Mueller et al., 2015; Bryson et al., 2016). The ⩽1.5 μm fraction of cells was incubated in acid-washed and rinsed 10 l polycarbonate carboys amended with ~1 μm final concentration of 13C-labeled substrates (>98%13C) in triplicate: amino acids, protein, glucose, starch, acetate and lipids (Sigma-Aldrich, St Louis, MO, USA, 426199, 642878, 389374, 605336, 282014 and 426202). To determine the necessary time for quantifying substrate assimilation while also minimizing potential cross-feeding, incubations were carried out for 15 and 32 h. Because 13C label was detected after 15 h (see methods below), this study presents data primarily from the t=0 sample and 15 h incubations. Incubations were held at constant temperature close to the ambient water column (19 °C). Cells from replicate microcosms were concentrated in parallel on 0.2 μm polyethersulfone membrane filters, which were transferred to sterile 15 ml tubes, immediately frozen on dry ice and subsequently transferred to a −80 °C freezer for long-term storage. Formaldehyde-fixed and SYBR Green-stained cells were counted with a Guava Technologies (Hayward, CA, USA) flow cytometer as previously described (Tripp et al., 2008; Tripp, 2008).
16S rRNA gene amplicon analysis
PCR amplicon libraries of the v4 rRNA gene region were amplified from DNA extracts of all microcosms (Mueller et al., 2015) using dual-indexed 16S primers (Kozich et al., 2013). Resulting amplicons were sequenced on the Illumina (San Diego, CA, USA) MiSeq platform at Oregon State University’s Center for Genome Research and Biocomputing as previously described (Bryson et al., 2016). A more detailed protocol is available in the Supplementary Methods.
Metagenome sequencing, assembly and annotation
DNA extraction, metagenome library construction and sequencing were previously described (Mueller et al., 2015; Supplementary Methods). Metagenome abundances for predicted coding sequences (CDS) were determined by mapping non-redundant sequencing reads back to the contigs using Bowtie 2 (Langmead and Salzberg, 2012) with default settings and calculating the average coverage for each locus using the bedtools coverage tool (Barnett et al., 2011). Relative abundances of operational taxonomic units (OTUs) detected in the 16S rRNA amplicon libraries were highly correlated with 16S rRNA gene abundances that were assembled using EMIRGE (Miller et al., 2011) from the metagenomes (data not shown).
Peptide mass spectrometry, identification and database construction
Protein extraction, digestion and mass spectrometry analysis followed previously reported protocols (Bryson et al., 2016; Supplementary Methods). This procedure utilized SDS lysis and trichloroacetic acid precipitation of proteins, followed by cleanup and trypsin digestion on centrifugal filters, and finally Multidimensional Protein Identification Technology was used for the liquid chromatography tandem mass spectrometry measurements using an LTQ Orbitrap Elite mass spectrometer (Thermo Scientific, Waltham, MA, USA) (Washburn et al., 2001). Peptide identifications are available under the PRIDE archive PXD002641.
The final database used for SIP searches (60 006 CDS) was constructed from all CDS with at least one peptide identification in regular (that is, unlabeled search mode) Sipros searches of each metaproteome against the 391 847 CDS from the full metagenome assembly and a reduced database consisting of 181 704 CDS from only high metagenome coverage contigs. To ensure support for our reduced database selection approach, test Sipros searches in labeled mode were performed comparing subsets of the spectra data against the non-reduced metagenome database. Under these conditions all labeled proteins identified were also identified in unlabeled searches, suggesting a low rate of missed protein detections by using a reduced database (data not shown).
Proteomic SIP analysis
Label frequency and average enrichment values were calculated for each set of proteins assigned to a taxon at a specific rank, as previously described (Bryson et al., 2016). We considered substrate incorporation to be detected if a peptide spectral match (PSM) was labeled, that is, if it was identified with ⩾2% 13C content. Label frequency was defined as the relative frequency of labeled PSM to total PSM for a set of protein identifications assigned to a specific population, that is, the proportion of identified mass spectra that were labeled. Accordingly, this metric quantifies the extent of de novo protein synthesis for the population. Average enrichment was defined as the average % 13C content of all labeled PSM that were detected for a set of proteins assigned to a specific population. As such, this metric quantifies substrate specialization. Relative substrate assimilation values for taxonomic bins were calculated as the proportion of labeled PSM multiplied by the average % 13C enrichment for PSM of a bin divided by the sum of this value for all bins in a sample, that is, the proportion of total detected substrate assimilation within a replicate. To enable comparisons across substrates, it was necessary to standardize to the mean and s.d. of the observed average enrichment values for sets of proteins assigned to different genera (or higher taxonomic levels if genus level assignments were not available) within each treatment. Hierarchical clustering and heatmaps of standardized average enrichment and label frequency metrics (data in Supplementary Tables S1 and S2) were performed in MeV (Howe et al., 2010). Significant levels of average enrichment were determined by comparing the observed average enrichment for a population to the values obtained from 1000 random samples of labeled spectra from the entire set of labeled PSM obtained for a metaproteome. Values were determined to be significant if they were equal or more extreme than 95% of values obtained from the randomization test. Significant average enrichment values are identified with ‘*’ in Supplementary Table S1. Other statistical tests were performed with the scipy stats and skbio python libraries.
Chip-SIP analysis
Probes were designed for 16S rRNA gene OTUs commonly found in coastal Pacific Ocean samples (Supplementary Table S3) using ARB (Ludwig et al., 2004), as previously described (Mayali et al., 2012). Duplicate RNA samples were extracted, fluorescently labeled, fragmented and hybridized as previously described (Mayali et al., 2014), and secondary ion mass spectrometry analysis of microarrays hybridized with 13C rRNA was performed at LLNL with a Cameca NanoSIMS 50 (Cameca, Gennevilliers, France). A more detailed protocol is available in the Supplementary Methods. OTUs were considered significantly isotopically enriched if the slope minus two calculated s.e.’s was >0 and if the slope was significant based on a t-score statistic (t=slope/s.e.) with a P-value of <0.05. Data are presented for the average slope from two biological replicates. The phylogenetic tree for representative 16S rRNA genes was generated by aligning sequences with Muscle (Edgar, 2004), selecting conserved sites with Gblocks (Talavera and Castresana, 2007) and constructing the tree in FastTree (Price et al., 2010) with default settings.
Results
Substrate-specific shifts in microbial community structure
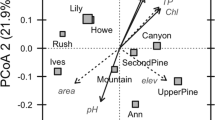
High-throughput sequencing of 16S rRNA gene amplicon libraries and whole-community DNA facilitated comparisons between the initial microbial community in the sampled surface water (t=0) and end-point communities (t=15 h), allowing us to determine how the six substrate amendments influenced community structure. Total cell counts increased by an average of 2.6-fold across all treatments, with the highest increases in the starch and glucose treatments, 3.9- and 2.9-fold increases, respectively (Supplementary Table S4). Principal coordinate analysis of pairwise community distances between rarefied sample OTU counts from 16S amplicon data (Figure 1a) indicated reproducible population shifts for each substrate addition; within treatment group distances were significantly different from random (permutational analysis of variance, pseudo-F=14.0, P-value ⩽0.001). Replicates from both the acetate and lipids treatments were the most structurally similar to the original community, whereas the remaining treatments were more distant from the t=0 community.
(a) Principal coordinate analysis plot of pairwise distances between rarefied sample OTU counts. (b) Stacked bar charts depict the relative coverage of predicted coding sequences (CDS) by metagenomes from the T=0 and each substrate amendment. The taxonomy color key indicates the order-level classification or class level, if the order was unclassified. ‘Other’ refers to low-coverage or unclassified CDS.
In addition to serving as the basis for a metaproteome search database, analysis of metagenomes from pooled replicates from each treatment group revealed the underlying taxonomic composition of the initial and end-point communities (Mueller et al., 2015; Bryson et al., 2016) complementing the analysis of 16S rRNA gene amplicon libraries. Assessment of relative abundances for dominant taxonomic lineages (Figure 1b) was based on both the recruitment of metagenomic reads to the assembled protein-coding sequences (CDS) and rarefied OTU counts (Supplementary Table S5; Supplementary Figure S2). Whole-community profiles showed that the acetate and lipid treatments most closely resembled the original sampled community, whereas communities from the glucose and starch treatments and the amino acids and protein treatments were most similar to each other, respectively (Figure 1b).
Comparisons of relative abundances for OTU counts summed at the order-level taxonomy indicated significant differences for 35 orders across all treatments (Supplementary Table S5). Examination of the relative abundance shifts for specific populations within the six treatments allowed subsequent comparisons between these community dynamics (Figure 1b) and substrate assimilation, as measured by the SIP approaches. Metagenomic and 16S rRNA amplicon data showed that Rhodobacterales generally decreased in relative abundance across all treatments, except for the amino acids and acetate treatments (Supplementary Figure S2). The Alteromonadales order increased in coverage across all treatments (Supplementary Figure S2), indicating a general rapid growth response to incubation conditions as has been previously reported (Pedler et al., 2014). Notably, these observed relative abundance increases were highest in the protein and amino acids treatments, both of which contain organic nitrogen. Conversely, oligotrophic marine Gammaproteobacteria (Cho and Giovannoni, 2004), including SAR86, SAR92 and other Cellvibrionales populations generally declined in relative abundance across all treatments (Supplementary Figure S2). The Flavobacteriales order generally increased in relative abundance in all treatments and increased significantly in the starch and glucose treatments (Supplementary Figure S2).
Substrates differ in whole-community assimilation
As amplicon sequencing revealed significantly reproducible substrate-specific population shifts, we hypothesized that community structure may have been shaped by population-specific substrate utilization, and further evaluated this using SIP enrichment of peptides and rRNA. We identified an average of 22 915 peptides and 4087 proteins in each of the 21 metaproteomes (Supplementary Table S4). Detection of labeled peptides in each of the six treatments produced distinct % 13C enrichment profiles (Figure 2) with significantly different average enrichment values for the whole-community metaproteomes (analysis of variance, F=15.75, P<0.001). The average % 13C enrichment of labeled peptides detected in the amino-acid treatment was significantly higher than the other five treatments (t-test, P<0.01; Figure 2; Supplementary Table S4). Thirty-one populations (defined at the genus or broader level of taxonomy) assimilated substrate in all six treatments, suggesting a broad ability of the dominant members of the community to assimilate the classes of DOC substrates tested. This was evident by the similar richness of community members with detected substrate assimilation across all treatments (Supplementary Figure S3). Although, the number of taxa for which substrate assimilation was detected above background levels indicated widespread utilization of the added substrates, a comparison of relative substrate assimilation indicated substantial differences between individual taxa (Supplementary Figure S4). These relative substrate assimilation data are based on the combination of two factors: the % 13C enrichment of labeled peptides per taxa (average enrichment, an indicator of resource specialization) and the proportion of labeled peptides (label frequency, an indicator of de novo protein synthesis activity). For example, of all the detected label in the metaproteome for the starch treatment, nearly 70% was accounted for by Flavobacteriaceae proteins (Supplementary Figure S4). In the acetate treatments, nearly 60% of the total detected label was found in Rhodobacteraceae proteins.
Substrate specialization is taxonomically conserved
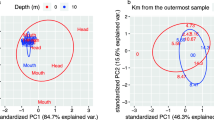
Our observation of clear biases in substrate assimilation patterns by diverse taxa across all treatments (Supplementary Figure S4) prompted us to investigate whether trends in resource partitioning were more similar among related populations than expected under a random null model. Correlation scores for pairwise comparisons of average enrichment values were significantly higher among related taxa at the class rank than would be expected for random taxonomic assignments (permutational analysis of variance, Pseudo-F=14.4, P-value ⩽0.001). This correlation analysis also demonstrated significantly different variances between classes (Levene’s test, statistic=6.2, P-value=0.003), with less variance within the Alphaproteobacteria than within the Flavobacteriales and the Gammaproteobacteria populations (Figure 3b).
(a) Clustering of taxa by relative incorporation of labeled substrate into proteins. The heatmap depicts standardized % 13C enrichment, indicating a population’s average enrichment relative to the other populations in each treatment. The color scale indicates z-scores: −2 (blue); 0 (black); and 2 (yellow). The distance scale for hierarchical clustering indicates Pearson’s correlation scores for standardized values. Taxonomy of protein bins is assigned up to genus level when possible. Class-level taxonomy is indicated by the following colors: Alphaproteobacteria (red); Gammaproteobacteria (green); Flavobacteriia (blue); and others (white). (b) Pairwise Pearson’s correlation coefficients of average enrichment values for populations within the three most abundant classes represented in a.
As previously shown in Figure 2, the six substrates were assimilated at different levels of % 13C enrichment, potentially due to existing substrate in the samples or biological differences in assimilation efficiency. Thus, to compare substrate assimilation by individual taxa, relative to other taxa, regardless of individual treatment effects, it was necessary to standardize average enrichment values within each sample (see Materials and Methods). Hierarchical clustering of these standardized enrichment values divided taxa into four groups based on average enrichment trends (Figure 3a). In addition to identifying significant trends in substrate assimilation across all treatments, the quantified average enrichment values for individual populations were compared to the expected values from random samples of labeled PSM from the whole-community metaproteomes. These analyses revealed how these substrates are partitioned in these communities and the interpopulation variation for the three dominant class level taxonomies found in these samples.
Proteins assigned to genera or higher taxonomies within the Flavobacteriales order formed a highly correlated cluster with average enrichment values among the highest of the starch treatment community (Figure 3a). These included proteins assigned to Formosa, Myroides and Flavobacterium that had significantly high average enrichment values (P<0.05; Supplementary Table S1) in at least one replicate from starch treatment. The exception to this trend was proteins assigned to Owenweeksia, which had significantly high average enrichment in one protein treatment and accounted for most of the variability seen in pairwise correlation scores for the Flavobacteriales (Figure 3b).
Gammaproteobacteria populations demonstrated less highly correlated average enrichment values than the other two abundant classes suggesting divergence of resource utilization strategies within this broad taxonomic group (Figure 3b). Oligotrophic marine Gammaproteobacteria clades, consisting of SAR92 and OM60/NOR5 clades of the Cellvibrionales order along with SAR86 and other unclassified Gammaproteobacteria, were among the populations with significantly high % 13C enrichment in at least one lipids treatments (Supplementary Table S1), averaging 3.9%, 8.0%, 5.6% and 4.8%, respectively, compared to the whole-community average enrichment (3.3±0.19%). In comparison, Glaciecola, Pseudoalteromonas and Colwellia genera from the Alteromonadales order assimilated amino acids at significantly high average enrichment (P<0.05; 35.7%, 41.3% and 23.6% respectively) in addition to lipids.
The third dominant order, comprised of Rhodobacterales populations, was among the most abundant at the time of sampling and had significantly high average enrichment in the glucose, amino-acid and acetate treatments. However, some intra-order variability was observed as proteins assigned to the Loktanella and Roseobacter had significantly high average enrichment in all three lipids treatments (Supplementary Table S1). These two genera plus Sulfitibacter, and Planktomarina genera had significantly high enrichment values in many of the treatments, with the notable exception of the starch treatments. Although raw average enrichment values were positively correlated between Pelagibacter and these Rhodobacterales populations (Figure 3b), it did not have high average enrichment values relative to the other populations in any of the six treatments.
Substrate preference and availability influence activity
In addition to measuring substrate specialization, proteomic SIP data can indicate de novo protein synthesis activity through quantification of ‘label frequency’, the relative frequency of labeled versus total PSM for a population. Pairwise Pearson’s correlation coefficients for label frequency values indicated that trends in de novo protein synthesis activity followed substrate specialization patterns and were also significantly more similar between related populations than random (permutational analysis of variance, pseudo-F=8.9, P-value ⩽0.001). Indeed, within treatment measures of label frequency and average enrichment were highly correlated (average Pearson’s r=0.58, s.d.=0.22, n=18). However, related populations did exhibit heterogeneity in terms of the magnitude of protein synthesis as quantified by label frequency values. This was evident from the hierarchical clustering of populations based on label frequency z-scores (Supplementary Figure S5A). Genera within the Rhodobacteriales, which were abundant at the time of sampling, and the Alteromonadales, which exhibited rapid relative abundance increases during the incubations, were consistently among the most active populations (Supplementary Figure S5A). Flavobacteriales populations generally had the highest label frequency in the starch treatment, and the Flavobacterium genus demonstrated much higher activity than others from this order. Populations of Gammaproteobacteria from clades within the Cellvibrionales and SAR86 along with Methylophilales (a member of the Betaproteobacteria), and Pelagibacteriales (a member of the Alphaproteobacteria) had low overall protein synthesis activity across all treatments (Supplementary Figure S5A). Although populations of Alphaproteobacteria exhibited the most correlated resource specialization patterns (Figure 3b), they also exhibited the greatest heterogeneity in terms of protein synthesis activity (Supplementary Figure S5B) among the three abundant classes.
Chip-SIP identifies substrate-assimilating populations at high taxonomic resolution
Population-level substrate assimilation patterns were also measured in rRNA pools. Out of 390 taxa represented by probe spots on the microarray, Chip-SIP identified 72 taxa with 13C enrichment in at least one treatment (Figure 4; Supplementary Figure S6). As observed in the proteomic SIP results, glucose and amino-acid treatments had the most widespread assimilation across 16S rRNA-targeted taxa, 40 and 39 out of 72, respectively. Starch was the third most widely assimilated (32 out of 72 taxa), followed by the protein treatment. Among the taxa with 13C-enriched rRNA, members of the Flavobacteriales order (Figure 4), primarily from the Cryomorphaceaea and Flavobacteriaceae families, were highly enriched with 13C from the starch treatment, similar to the trend in the proteomic SIP data. Gammaproteobacteria from the SAR86 clade, the Cellvibrionales and Alteromonadales order, plus other unclassified clades were enriched with 13C from amino acids, glucose and to a lesser extent starch. Rhodobacterales taxa were 13C-enriched in the amino-acid, glucose and protein treatments. 13C label from protein assimilation was also detected in SAR116 Alphaproteobacteria, as well as in the SAR406 clade, Deltaproteobacteria, Epsilonproteobacteria and Flavobacteria. Lipid assimilation was only detected in a SAR406 and a deep SAR11 taxa, while acetate assimilation was only detected after 32 h of incubation (see Materials and Methods) in rRNA hybridized to one Rhodobacteraceae probe set. Similar to the proteomic SIP data, class-level groupings of probe spots had significantly higher within-group Pearson’s correlation coefficients than random groupings (permutational analysis of variance, pseudo-F=3.1, P-value ⩽0.001). Although the detected trends in substrate assimilation were more similar for taxa at the class level, comparisons between closely related populations within order-level taxonomies revealed some variation in substrate assimilation. Glucose, starch and lipid assimilation was detected for a deep ecotype of SAR11, whereas as surface-associated ecotype was found to be highly enriched only in amino acids. Within the Flavobacteriales order, there were two populations that exhibited higher assimilation of glucose than starch. And finally, within the Rhodobacteriales order, there was some variation in the assimilation of amino acids and glucose.
Phylogenetic tree of rRNA sequences used for Chip-SIP microarray probes and accompanying heatmap depicting 13C enrichment values (average of two replicates) for combined probe sets. Taxonomy is indicated by the color key. Names for Chip-SIP probe sets are indicated in Supplementary Figure S6.
Discussion
This study utilized metagenomics, 16S rRNA gene sequencing, proteomic SIP and rRNA Chip-SIP in a combined assessment of resource partitioning among sympatric populations of coastal marine microbes. Metagenomics and 16S rRNA gene sequence libraries indicated distinct structural changes in the sampled community that accompanied each substrate addition. However, the relationship between substrate assimilation and population dynamics was often inconsistent. Relative abundance data from sequencing libraries and substrate assimilation data from proteomic SIP data were the most consistent for populations of Alteromonadales, which exhibited relative abundance increases and significant substrate assimilation in the amino-acid and protein treatments. Populations of Pelagibacterales, with previously reported slow growth rates (Rappé et al., 2002), were abundant at the time of sampling, but exhibited neither increased relative abundance nor high average enrichment values for peptides across all treatments, despite significant numbers of labeled peptides being detected in all treatments. Flavobacteriales abundance and assimilation were positively correlated in the starch treatment, but in the glucose treatment, an increase in their relative abundance was not accompanied by significant glucose assimilation. Conversely, Rhodobacterales populations accounted for the highest amount of detected substrate assimilation in their proteomes but exhibited negligible shifts in their relative abundance. Oligotrophic marine Gammaproteobacteria had low label frequency (the proportion of newly made 13C peptides relative to the total pool) and generally decreased in relative abundance across all treatments, yet were among the most 13C-enriched in the lipid treatment.
The differences observed between these populations may suggest different life strategies. In the case of Rhodobacterales, these populations are actively incorporating substrate, but it is not translating into rapid growth as seen for the Alteromonadales. In another example, genomic data would predict widespread glucose utilization by diverse populations in our samples. However, the combined SIP and metagenomic approaches allowed us to distinguish between the responses of Flavobacteriales populations, which rapidly increased in abundance, and Rhodobacterales populations, which allocated this resource into biomass without a similar increase in relative abundance. These observations suggest that the role of substrate utilization in population dynamics is more complex than can be predicted by genomic data or revealed by relative abundance metrics alone. This apparent lack of correlation between relative abundance shifts and substrate assimilation patterns may result from the fact that populations within a complex microbial community can vary in their growth efficiency and rates (Roller et al., 2016) and the inherent challenges of linking relative abundance changes to absolute changes (Nayfach and Pollard, 2016; Props et al., 2016). In addition, unmeasured substrate in natural samples may support growth that would not be measured by SIP approaches.
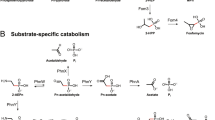
Indeed, interpretation of SIP data, while powerful, can also present challenges, as substrate assimilation may be affected by biological factors relating to the uptake (Kirchman, 2003), physical factors, including the concentration of similar substrates in the sampled sea water and the accessibility of each substrate (Nagata, 2008), or flux (Chubukov et al., 2014) of specific substrates into amino acid and RNA. To address this latter issue, we evaluated how label incorporation into protein and rRNA was differently quantified in the biomass of populations as measured by the two SIP approaches. Amino-acid and glucose assimilation were the most highly detected in both peptides and rRNA, indicating broad and efficient utilization and assimilation by diverse populations. However, detection of isotopic enrichment was less widespread for the lipid and acetate treatments with Chip-SIP compared to proteomic SIP, suggesting lower overall incorporation of the 13C label from these compounds into nucleotides and subsequent de novo-synthesized RNAs than into amino acids and de novo-synthesized proteins. This may indicate differences in the sensitivity of the two techniques; with proteomic SIP one has the ability to quantify isotope incorporation for multiple peptides independently, whereas in Chip-SIP all 16S rRNA molecules are hybridized to a single probe spot. Alternatively, the allocation of specific substrates into biomolecules may vary depending on where a substrate enters pathways of central carbon metabolism. Increased cellular concentrations of acetyl-CoA from direct utilization of acetate or beta oxidation of lipids could result in the preferential flux of these substrates into the tricarboxylic acid cycle and amino-acid biosynthesis pathways via the anaplerotic reaction sequence of the glyoxylate shunt (Sauer and Eikmanns, 2005; Gottschalk, 2012), and the subsequent redirection of other unlabeled carbon sources to the synthesis of carbohydrates and nucleotides via gluconeogenesis and the pentose phosphate pathways (Chubukov et al., 2014; Wolfe, 2015). This explanation is further supported by the opposing trend observed among Gammaproteobacteria populations, which showed higher glucose assimilation measured by Chip-SIP (Figure 4) than by proteomic SIP (Figure 3).
The substrate assimilation trends we measured revealed evolutionary cohesive ecological strategies among related populations and divided the microbial community into distinct guilds. Rhodobacterales populations exhibited a preference for low-molecular-weight monomers: acetate; glucose; and amino acids (either directly or as products from extracellular hydrolysis of whole proteins) (Nagata, 2008). This order demonstrated a more generalist lifestyle than other populations, supporting prior genomic observations for these clades (Newton et al., 2010). Pelagibacterales populations exhibited similar trends in resource assimilation as the Rhodobacterales, but were less active overall in assimilating substrate into protein biomass. Resource partitioning between these two abundant Alphaproteobacteria populations may be better described in terms of competitive advantages under diffusion-limited oligotrophic conditions given our observations for substrate assimilation. Flavobacteriales populations displayed a distinct advantage in assimilating starch. These findings confirm prior genomic observations for Flavobacteriales taxa, which have identified high numbers of enzymes for extracellular binding, hydrolysis and transport of complex carbohydrates (Mann et al., 2013; Xing et al., 2014). Specialization for lipid assimilation was observed among low-activity members of the Gammaproteobacteria, including populations from SAR86, SAR92 and OM60/NOR5 clades, as well as fast growing populations of Alteromonadales. This observation for SAR86 confirms an earlier hypothesis for lipid utilization based on genomic analysis (Dupont et al., 2012). In addition, metatranscriptomic studies have identified the importance of fatty acid metabolic genes within populations of Cellvibrionales and Alteromonadales (McCarren et al., 2010; Gifford et al., 2013).
Overall, these patterns in community resource partitioning confirmed prior observations of population dynamics made during phytoplankton blooms (Teeling et al., 2012; Buchan et al., 2014). Significantly similar substrate assimilation trends were identified at the class level, implying evolutionary conserved ecological strategies among related populations. Furthermore, these results suggest that functional guilds within marine surface waters can be defined at deep branching taxonomic lineages, confirming predictions based on genomic data (Martiny et al., 2013). Flavobacteriales are observed to increase when carbohydrate stocks increase, indicating a tightly coupled relationship with the release of this DOC type from primary producers. Gammaproteobacteria populations emerge later in bloom timelines when proteins and lipids are more likely to accumulate as unhealthy phytoplankton begin to leak or lyse. It is also evident that different lineages within the Gammaproteobacteria exhibit ecological strategies that separate oligotrophic clades with lower biosynthetic responses to nutrient availability from copiotrophic clades of Alteromonadales that rapidly respond to nutrient replete conditions. Alphaproteobacteria populations also showed distinct responses to nutrient additions. Whereas the Rhodobacterales populations generally exhibited significant protein biosynthesis with the addition of low-molecular-weight compounds, this response was not observed for Pelagibacterales populations that also assimilated these substrates. These findings fit with prior observations of relatively stable population sizes for Pelagibacterales throughout phytoplankton bloom cycles and the observed bloom responses of some Rhodobacterales populations to phytoplankton blooms (Buchan et al., 2014).
The general trends identified in this study for relatively broad taxonomic groups may reflect the choice of substrates used in these experiments. More distinct DOC molecules, such as specific amino acids, fatty acids, specific aromatic compounds or organic sulfur compounds, such as dimethylsulfoniopropionate, may be less conserved at finer taxonomic scales (Martiny et al., 2013). While the presence or absence of specific genomic-encoded metabolic pathways may account for some variability in activity between closely related taxa, the evolutionary fine tuning of individual microbial strains to substrate concentrations or environmental parameters, such as temperature and salinity, may be of greater importance. Indeed, fine scale variation in the specific growth responses of taxa within the Vibrio genus to a single class of carbon substrate has been attributed to the gene copy number for a specific enzyme, rather than its mere presence/absence (Hehemann et al., 2016). Determining how individual populations respond to substrate availability under various environmental conditions, such as resource concentration or temperature, requires utilization of techniques that quantify activity, such as the SIP approaches presented here, that cannot be assessed using more conventional genomic-based inquiry.
Prior work has demonstrated that at low resource concentrations substrate specialists are the most active (Sarmento et al., 2016), suggesting that this specialization is driven by competition for limited resources. While this study tested the assimilation of substrates that were amended all at once and at non-limiting concentrations, the sensitivity of the two SIP approaches allowed the delineation of resource partitioning within communities, even though all of the tested substrates were utilized by all of the more abundant microbial populations. We found conserved resource partitioning across populations belonging to similar taxonomic ranks, which implies widespread overlap in the utilization of the same substrates by related taxa. However, population-specific measures of activity revealed variable responses within lineages. Genomic-, transcriptomic- and proteomic-based inquiry along with population surveys have enabled hypotheses about resource partitioning, but without SIP approaches, the degree to which this partitioning occurs cannot be quantified. Our results suggest that resource preferences may rely on phylogenetically conserved physiological traits that govern the acquisition and flux of metabolites through central metabolic pathways. Transient success of discrete populations within these phylogenetically defined functional guilds may be additionally dependent upon external factors, including substrate or nutrient concentrations, grazing pressure and viral lysis, inter-species interactions or differential adaptation to distinct physical and chemical environments. Our observation of overlapping resource utilization also implies some functional redundancy within related populations, a factor that could potentiate more resilient ecosystems in the face of environmental change. But more importantly, these observations elucidate the population-level mechanisms by which whole communities partition resources, which results in rapid responses to phytoplankton blooms and consumption of labile DOM.
References
Azam F, Malfatti F . (2007). Microbial structuring of marine ecosystems. Nat Rev Microbiol 5: 782–791.
Barnett D, Garrison E, Quinlan A, Strömberg M, Marth G . (2011). BamTools: a C++ API and toolkit for analyzing and managing BAM files. Bioinformatics 27: 1691–1692.
Biddanda B, Benner R . (1997). Carbon, nitrogen, and carbohydrate fluxes during the production of particulate and dissolved organic matter by marine phytoplankton. Limnol Oceanogr 42: 506–518.
Bryson S, Li Z, Pett-Ridge J, Hettich RL, Mayali X, Pan C et al. (2016). Proteomic stable isotope probing reveals taxonomically distinct patterns in amino acid assimilation by coastal marine bacterioplankton. mSystems 1: e00027–15.
Buchan A, LeCleir GR, Gulvik CA, González JM . (2014). Master recyclers: features and functions of bacteria associated with phytoplankton blooms. Nat Rev Microbiol 12: 686–698.
Cho J-C, Giovannoni SJ . (2004). Cultivation and growth characteristics of a diverse group of oligotrophic marine gammaproteobacteria. Appl Environ Microbiol 70: 432–440.
Chubukov V, Gerosa L, Kochanowski K, Sauer U . (2014). Coordination of microbial metabolism. Nat Rev Microbiol 12: 327–340.
DeLong EF, Preston CM, Mincer T, Rich V, Hallam SJ, Frigaard N-U et al. (2006). Community genomics among stratified microbial assemblages in the ocean’s interior. Science 311: 496–503.
Dinasquet J, Kragh T, Schrøter M-L, Søndergaard M, Riemann L . (2013). Functional and compositional succession of bacterioplankton in response to a gradient in bioavailable dissolved organic carbon: bacterial community response to a gradient in bDOC. Environ Microbiol 15: 2616–2628.
Ducklow H . (2000) Bacterial production and biomass in the oceans. In: Kirchman D (ed). Microbial Ecology of the Oceans. Wiley: New York, NY, USA, pp 85–120.
Dumont MG, Murrell JC . (2005). Stable isotope probing—linking microbial identity to function. Nat Rev Microbiol 3: 499–504.
Dupont CL, Rusch DB, Yooseph S, Lombardo M-J, Alexander Richter R, Valas R et al. (2012). Genomic insights to SAR86, an abundant and uncultivated marine bacterial lineage. ISME J 6: 1186–1199.
Edgar RC . (2004). MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32: 1792–1797.
Fuhrman JA, Hewson I, Schwalbach MS, Steele JA, Brown MV, Naeem S . (2006). Annually reoccurring bacterial communities are predictable from ocean conditions. Proc Natl Acad Sci USA 103: 13104–13109.
Geider RJ, Roche JL . (2002). Redfield revisited: variability of C[ratio ]N[ratio ]P in marine microalgae and its biochemical basis. Eur J Phycol 37: 1–17.
Gifford SM, Sharma S, Booth M, Moran MA . (2013). Expression patterns reveal niche diversification in a marine microbial assemblage. ISME J 7: 281–298.
Gilbert JA, Steele JA, Caporaso JG, Steinbrück L, Reeder J, Temperton B et al. (2012). Defining seasonal marine microbial community dynamics. ISME J 6: 298–308.
Gómez-Consarnau L, Lindh MV, Gasol JM, Pinhassi J . (2012). Structuring of bacterioplankton communities by specific dissolved organic carbon compounds. Environ Microbiol 14: 2361–2378.
Gottschalk G . (2012) Bacterial Metabolism. Springer Science & Business Media: New York, NY, USA.
Hehemann J-H, Arevalo P, Datta MS, Yu X, Corzett CH, Henschel A et al. (2016). Adaptive radiation by waves of gene transfer leads to fine-scale resource partitioning in marine microbes. Nat Commun 7: 12860.
Howe E, Holton K, Nair S, Schlauch D, Sinha R, Quackenbush J . (2010) MeV: MultiExperiment Viewer. In: Ochs MF, Casagrande JT, Davuluri RV (eds). Biomedical Informatics for Cancer Research. Springer: Boston, MA, USA, pp: 267–277.
Hyatt D, Pan C . (2012). Exhaustive database searching for amino acid mutations in proteomes. Bioinformatics 28: 1895–1901.
Kirchman DL . (2003) The contribution of monomers and other low-molecular weight compounds to the flux of dissolved organic material in aquatic ecosystems. In: Sinsabaugh SEGFL (ed). Aquatic Ecosystems. Academic Press: Burlington, VT, USA, pp 217–241.
Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD . (2013). Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl Environ Microbiol 79: 5112–5120.
Langmead B, Salzberg SL . (2012). Fast gapped-read alignment with Bowtie 2. Nat Methods 9: 357–359.
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar et al. (2004). ARB: a software environment for sequence data. Nucleic Acids Res 32: 1363–1371.
Mann AJ, Hahnke RL, Huang S, Werner J, Xing P, Barbeyron T et al. (2013). The genome of the alga-associated marine flavobacterium Formosa agariphila KMM 3901 T reveals a broad potential for degradation of algal polysaccharides. Appl Environ Microbiol 79: 6813–6822.
Martiny AC, Treseder K, Pusch G . (2013). Phylogenetic conservatism of functional traits in microorganisms. ISME J 7: 830–838.
Mayali X, Weber PK, Brodie EL, Mabery S, Hoeprich PD, Pett-Ridge J . (2012). High-throughput isotopic analysis of RNA microarrays to quantify microbial resource use. ISME J 6: 1210–1221.
Mayali X, Weber PK, Mabery S, Pett-Ridge J . (2014). Phylogenetic patterns in the microbial response to resource availability: amino acid incorporation in San Francisco Bay. PLoS ONE 9: e95842.
Mayali X, Weber PK, Pett-Ridge J . (2013). Taxon-specific C/N relative use efficiency for amino acids in an estuarine community. FEMS Microbiol Ecol 83: 402–412.
McCarren J, Becker JW, Repeta DJ, Shi Y, Young CR, Malmstrom RR et al. (2010). Microbial community transcriptomes reveal microbes and metabolic pathways associated with dissolved organic matter turnover in the sea. Proc Natl Acad Sci USA 107: 16420–16427.
Meon B, Kirchman DL . (2001). Dynamics and molecular composition of dissolved organic material during experimental phytoplankton blooms. Mar Chem 75: 185–199.
Miller CS, Baker BJ, Thomas BC, Singer SW, Banfield JF . (2011). EMIRGE: reconstruction of full-length ribosomal genes from microbial community short read sequencing data. Genome Biol 12: R44.
Morris RM, Frazar CD, Carlson CA . (2012). Basin-scale patterns in the abundance of SAR11 subclades, marine Actinobacteria (OM1), members of the Roseobacter clade and OCS116 in the South Atlantic: Bacterioplankton communities in the South Atlantic. Environ Microbiol 14: 1133–1144.
Morris RM, Nunn BL, Frazar C, Goodlett DR, Ting YS, Rocap G . (2010). Comparative metaproteomics reveals ocean-scale shifts in microbial nutrient utilization and energy transduction. ISME J 4: 673–685.
Mueller RS, Bryson S, Kieft B, Li Z, Pett-Ridge J, Chavez F et al. (2015). Metagenome sequencing of a coastal marine microbial community from Monterey Bay, California. Genome Announc 3: e00341–15.
Myklestad SM . (2000) Dissolved organic carbon from phytoplankton. In: Wangersky PJ (ed). The Handbook of Environmental Chemistry. Marine Chemistry. Springer: Berlin Heidelberg, pp 111–148.
Nagata T . (2008) Organic matter–bacteria interactions in seawater. In: Kirchman DL (ed). Microbial Ecology of the Oceans. John Wiley & Sons, Inc.: New York, NY, USA, pp 207–241.
Nayfach S, Pollard KS . (2016). Toward accurate and quantitative comparative metagenomics. Cell 166: 1103–1116.
Newton RJ, Griffin LE, Bowles KM, Meile C, Gifford S, Givens CE et al. (2010). Genome characteristics of a generalist marine bacterial lineage. ISME J 4: 784–798.
Norrman B, Zwelfel UL, Hopkinson CS, Brian F . (1995). Production and utilization of dissolved organic carbon during an experimental diatom bloom. Limnol Oceanogr 40: 898–907.
Pan C, Fischer CR, Hyatt D, Bowen BP, Hettich RL, Banfield JF . (2011). Quantitative tracking of isotope flows in proteomes of microbial communities. Mol Cell Proteomics 10: M110.006049.
Pedler BE, Aluwihare LI, Azam F . (2014). Single bacterial strain capable of significant contribution to carbon cycling in the surface ocean. Proc Natl Acad Sci USA 111: 7202–7207.
Price MN, Dehal PS, Arkin AP . (2010). FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS ONE 5: e9490.
Props R, Kerckhof F-M, Rubbens P, De Vrieze J, Hernandez Sanabria E, Waegeman W et al. (2016). Absolute quantification of microbial taxon abundances. ISME J 11: 584–587.
Rappé MS, Connon SA, Vergin KL, Giovannoni SJ . (2002). Cultivation of the ubiquitous SAR11 marine bacterioplankton clade. Nature 418: 630–633.
Roller BRK, Stoddard SF, Schmidt TM . (2016). Exploiting rRNA operon copy number to investigate bacterial reproductive strategies. Nat Microbiol 1: 16160.
Sarmento H, Morana C, Gasol JM . (2016). Bacterioplankton niche partitioning in the use of phytoplankton-derived dissolved organic carbon: quantity is more important than quality. ISME J 10: 2582–2592.
Sauer U, Eikmanns BJ . (2005). The PEP–pyruvate–oxaloacetate node as the switch point for carbon flux distribution in bacteria: we dedicate this paper to Rudolf K. Thauer, Director of the Max-Planck-Institute for Terrestrial Microbiology in Marburg, Germany, on the occasion of his 65th birthday. FEMS Microbiol Rev 29: 765–794.
Talavera G, Castresana J . (2007). Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst Biol 56: 564–577.
Teeling H, Fuchs BM, Becher D, Klockow C, Gardebrecht A, Bennke CM et al. (2012). Substrate-controlled succession of marine bacterioplankton populations induced by a phytoplankton bloom. Science 336: 608–611.
Treusch AH, Vergin KL, Finlay LA, Donatz MG, Burton RM, Carlson CA et al. (2009). Seasonality and vertical structure of microbial communities in an ocean gyre. ISME J 3: 1148–1163.
Tripp HJ . (2008). Counting marine microbes with Guava Easy-Cyte 96 well plate reading flow cytometer. Protoc Exch doi:10.1038/nprot.2008.29.
Tripp HJ, Kitner JB, Schwalbach MS, Dacey JWH, Wilhelm LJ, Giovannoni SJ . (2008). SAR11 marine bacteria require exogenous reduced sulphur for growth. Nature 452: 741–744.
Wang Y, Ahn T-H, Li Z, Pan C . (2013). Sipros/ProRata: a versatile informatics system for quantitative community proteomics. Bioinformatics 29: 2064–2065.
Washburn MP, Wolters D, Yates JR . (2001). Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nat Biotechnol 19: 242–247.
Whitman WB, Coleman DC, Wiebe WJ . (1998). Prokaryotes: the unseen majority. Proc Natl Acad Sci USA 95: 6578–6583.
Whittaker RH . (1972). Evolution and measurement of species diversity. Taxon 21: 213–251.
Wolfe AJ . (2015). Glycolysis for the microbiome generation. Microbiol Spectr 3.
Xing P, Hahnke RL, Unfried F, Markert S, Huang S, Barbeyron T et al. (2014). Niches of two polysaccharide-degrading Polaribacter isolates from the North Sea during a spring diatom bloom. ISME J 9: 1410–1422.
Yokokawa T, Nagata T . (2010). Linking bacterial community structure to carbon fluxes in marine environments. J Oceanogr 66: 1–12.
Acknowledgements
This work was funded by the Gordon and Betty Moore Foundation Marine Microbiology Initiative (Grant Id: GBMF3302). This research used resources of the Oak Ridge Leadership Computing Facility. Oak Ridge National Laboratory is supported by the Office of Science of the US Department of Energy and managed by the University of Tennessee-Battelle, L.L.C. for the U.S. Department of Energy. Work at LLNL was conducted under the auspices of DOE Contract DE-AC52-07NA27344; where Chip-SIP method development and NanoSIMS lab operations are provided by award SCW1039 from the DOE OBER Genomic Science Program. We thank the crew of the RV Rachel Carson, the Monterey Bay Research Institute, the lab of Stephen Giovannoni and the Oregon State University Center for Genomic Research and Biocomputing.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Bryson, S., Li, Z., Chavez, F. et al. Phylogenetically conserved resource partitioning in the coastal microbial loop. ISME J 11, 2781–2792 (2017). https://doi.org/10.1038/ismej.2017.128
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2017.128
This article is cited by
-
Resource partitioning and amino acid assimilation in a terrestrial geothermal spring
The ISME Journal (2023)
-
Single-cell isotope tracing reveals functional guilds of bacteria associated with the diatom Phaeodactylum tricornutum
Nature Communications (2023)
-
Rapid bacterioplankton transcription cascades regulate organic matter utilization during phytoplankton bloom progression in a coastal upwelling system
The ISME Journal (2022)
-
Simulating metagenomic stable isotope probing datasets with MetaSIPSim
BMC Bioinformatics (2020)
-
In marine Bacteroidetes the bulk of glycan degradation during algae blooms is mediated by few clades using a restricted set of genes
The ISME Journal (2019)